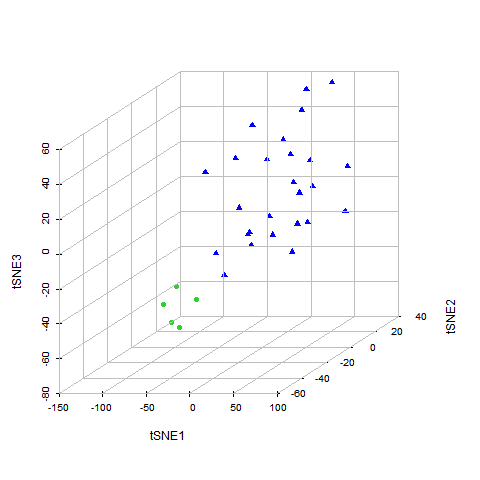
Disease specific cell death gene regulator search
Search Results:
CHOL specific autosis gene regulaors.
Different color in column1 means high , low , non-diff expressed gene in disease.
| NO. | Disease | CellDeathType | Genesymbol | Fullname | Control_mean | Disease_mean | log2FC | pvalue | padj | Weight | Detail |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | CHOL | autosis | DHODH | dihydroorotate dehydrogenase (quinone) | 11.03 | 7.84 | -0.49 | 0.0 | 0.0 | 100.0 | Detail |
| 2 | CHOL | autosis | EBP | EBP cholestenol delta-isomerase | 12.9 | 10.26 | -0.33 | 0.0 | 0.0 | 100.0 | Detail |
| 3 | CHOL | autosis | EZH2 | enhancer of zeste 2 polycomb repressive complex 2 subunit | 5.28 | 8.07 | 0.61 | 0.0 | 0.0 | 100.0 | Detail |
| 4 | CHOL | autosis | GCLC | glutamate-cysteine ligase catalytic subunit | 11.9 | 9.65 | -0.3 | 0.0 | 0.0 | 100.0 | Detail |
| 5 | CHOL | autosis | HGF | hepatocyte growth factor | 8.14 | 4.79 | -0.76 | 0.0 | 0.0 | 100.0 | Detail |
| 6 | CHOL | autosis | HGFAC | HGF activator | 13.11 | 5.41 | -1.28 | 0.0 | 0.0 | 100.0 | Detail |
| 7 | CHOL | autosis | IVD | isovaleryl-CoA dehydrogenase | 13.43 | 11.03 | -0.28 | 0.0 | 0.0 | 100.0 | Detail |
| 8 | CHOL | autosis | LGALS3 | galectin 3 | 8.73 | 11.52 | 0.4 | 0.0 | 0.0 | 100.0 | Detail |
| 9 | CHOL | autosis | PSME3 | proteasome activator subunit 3 | 10.17 | 11.2 | 0.14 | 0.0 | 0.0 | 100.0 | Detail |
| 10 | CHOL | autosis | SDC2 | syndecan 2 | 13.35 | 9.73 | -0.46 | 0.0 | 0.0 | 100.0 | Detail |
| 11 | CHOL | autosis | SLC25A20 | solute carrier family 25 member 20 | 11.68 | 8.32 | -0.49 | 0.0 | 0.0 | 100.0 | Detail |
| 12 | CHOL | autosis | SMOX | spermine oxidase | 7.18 | 9.51 | 0.41 | 0.0 | 0.0 | 100.0 | Detail |
| 13 | CHOL | autosis | SOD1 | superoxide dismutase 1 | 14.37 | 11.92 | -0.27 | 0.0 | 0.0 | 100.0 | Detail |
| 14 | CHOL | autosis | MCM2 | minichromosome maintenance complex component 2 | 7.32 | 9.99 | 0.45 | 0.0 | 0.0 | 100.0 | Detail |
| 15 | CHOL | autosis | MCM7 | minichromosome maintenance complex component 7 | 9.7 | 11.35 | 0.23 | 0.0 | 0.0 | 100.0 | Detail |
| 16 | CHOL | autosis | BIRC5 | baculoviral IAP repeat containing 5 | 3.49 | 7.89 | 1.18 | 0.0 | 0.0 | 100.0 | Detail |
| 17 | CHOL | autosis | HMGA1 | high mobility group AT-hook 1 | 8.69 | 12.04 | 0.47 | 0.0 | 0.0 | 100.0 | Detail |
| 18 | CHOL | autosis | CDC20 | cell division cycle 20 | 3.88 | 8.58 | 1.14 | 0.0 | 0.0 | 100.0 | Detail |
| 19 | CHOL | autosis | CHEK1 | checkpoint kinase 1 | 5.51 | 7.74 | 0.49 | 0.0 | 0.0 | 100.0 | Detail |
| 20 | CHOL | autosis | PLK1 | polo like kinase 1 | 4.05 | 8.39 | 1.05 | 0.0 | 0.0 | 100.0 | Detail |
| 21 | CHOL | autosis | TIMM8A | translocase of inner mitochondrial membrane 8A | 8.2 | 6.95 | -0.24 | 0.0 | 0.0 | 100.0 | Detail |
| 22 | CHOL | autosis | BNIP3 | BCL2 interacting protein 3 | 12.08 | 9.55 | -0.34 | 0.0 | 0.0 | 100.0 | Detail |
| 23 | CHOL | autosis | SPINT2 | serine peptidase inhibitor, Kunitz type 2 | 9.41 | 13.17 | 0.48 | 0.0 | 0.0 | 100.0 | Detail |
| 24 | CHOL | autosis | E2F1 | E2F transcription factor 1 | 5.09 | 8.78 | 0.79 | 0.0 | 0.0 | 100.0 | Detail |
| 25 | CHOL | autosis | KIFC1 | kinesin family member C1 | 4.48 | 8.33 | 0.9 | 0.0 | 0.0 | 100.0 | Detail |
| 26 | CHOL | autosis | NEK2 | NIMA related kinase 2 | 2.87 | 7.47 | 1.38 | 0.0 | 0.0 | 100.0 | Detail |
| 27 | CHOL | autosis | ADRB2 | adrenoceptor beta 2 | 8.02 | 3.79 | -1.08 | 0.0 | 0.0 | 100.0 | Detail |
| 28 | CHOL | autosis | ITGB4 | integrin subunit beta 4 | 7.2 | 12.45 | 0.79 | 0.0 | 0.0 | 100.0 | Detail |
| 29 | CHOL | autosis | POR | cytochrome p450 oxidoreductase | 14.05 | 11.34 | -0.31 | 0.0 | 0.0 | 100.0 | Detail |
| 30 | CHOL | autosis | TCF3 | transcription factor 3 | 8.88 | 10.97 | 0.31 | 0.0 | 0.0 | 100.0 | Detail |
| 31 | CHOL | autosis | DHRS3 | dehydrogenase/reductase 3 | 13.03 | 10.72 | -0.28 | 0.0 | 0.0 | 100.0 | Detail |
| 32 | CHOL | autosis | BAK1 | BCL2 antagonist/killer 1 | 7.76 | 9.47 | 0.29 | 0.0 | 0.0 | 100.0 | Detail |
| 33 | CHOL | autosis | SRC | SRC proto-oncogene, non-receptor tyrosine kinase | 8.83 | 11.05 | 0.32 | 0.0 | 0.0 | 100.0 | Detail |
| 34 | CHOL | autosis | CDK19 | cyclin dependent kinase 19 | 8.36 | 9.75 | 0.22 | 0.0 | 0.0 | 100.0 | Detail |
| 35 | CHOL | autosis | GGCX | gamma-glutamyl carboxylase | 12.3 | 10.14 | -0.28 | 0.0 | 0.0 | 100.0 | Detail |
| 36 | CHOL | autosis | ITCH | itchy E3 ubiquitin protein ligase | 11.08 | 10.01 | -0.15 | 0.0 | 0.0 | 100.0 | Detail |
| 37 | CHOL | autosis | KHDRBS1 | KH RNA binding domain containing, signal transduction associated 1 | 11.02 | 11.8 | 0.1 | 0.0 | 0.0 | 100.0 | Detail |
| 38 | CHOL | autosis | MMAB | metabolism of cobalamin associated B | 11.66 | 9.25 | -0.33 | 0.0 | 0.0 | 100.0 | Detail |
| 39 | CHOL | autosis | MMP11 | matrix metallopeptidase 11 | 5.6 | 11.01 | 0.97 | 0.0 | 0.0 | 100.0 | Detail |
| 40 | CHOL | autosis | NRF1 | nuclear respiratory factor 1 | 7.59 | 8.58 | 0.18 | 0.0 | 0.0 | 100.0 | Detail |
| 41 | CHOL | autosis | PLXNA1 | plexin A1 | 7.89 | 10.3 | 0.38 | 0.0 | 0.0 | 100.0 | Detail |
| 42 | CHOL | autosis | PRKD2 | protein kinase D2 | 9.39 | 10.86 | 0.21 | 0.0 | 0.0 | 100.0 | Detail |
| 43 | CHOL | autosis | ST3GAL6 | ST3 beta-galactoside alpha-2,3-sialyltransferase 6 | 10.41 | 6.31 | -0.72 | 0.0 | 0.0 | 100.0 | Detail |
| 44 | CHOL | autosis | BUB1 | BUB1 mitotic checkpoint serine/threonine kinase | 3.0 | 7.67 | 1.35 | 0.0 | 0.0 | 100.0 | Detail |
| 45 | CHOL | autosis | UBE2C | ubiquitin conjugating enzyme E2 C | 3.49 | 8.26 | 1.24 | 0.0 | 0.0 | 100.0 | Detail |
| 46 | CHOL | autosis | RCAN1 | regulator of calcineurin 1 | 12.38 | 9.08 | -0.45 | 0.0 | 0.0 | 100.0 | Detail |
| 47 | CHOL | autosis | ARF3 | ADP ribosylation factor 3 | 11.09 | 12.19 | 0.14 | 0.0 | 0.0 | 100.0 | Detail |
| 48 | CHOL | autosis | CCNA2 | cyclin A2 | 4.89 | 8.01 | 0.71 | 0.0 | 0.0 | 100.0 | Detail |
| 49 | CHOL | autosis | CDC6 | cell division cycle 6 | 4.61 | 8.31 | 0.85 | 0.0 | 0.0 | 100.0 | Detail |
| 50 | CHOL | autosis | MYBL2 | MYB proto-oncogene like 2 | 3.96 | 8.84 | 1.16 | 0.0 | 0.0 | 100.0 | Detail |
| 51 | CHOL | autosis | PAQR4 | progestin and adipoQ receptor family member 4 | 6.48 | 9.9 | 0.61 | 0.0 | 0.0 | 100.0 | Detail |
| 52 | CHOL | autosis | SLC38A1 | solute carrier family 38 member 1 | 8.27 | 12.06 | 0.54 | 0.0 | 0.0 | 100.0 | Detail |
| 53 | CHOL | autosis | FBXO31 | F-box protein 31 | 10.6 | 9.27 | -0.19 | 0.0 | 0.0 | 100.0 | Detail |
| 54 | CHOL | autosis | TM7SF2 | transmembrane 7 superfamily member 2 | 12.11 | 8.5 | -0.51 | 0.0 | 0.0 | 100.0 | Detail |
| 55 | CHOL | autosis | TRIM16 | tripartite motif containing 16 | 6.83 | 8.83 | 0.37 | 0.0 | 0.0 | 100.0 | Detail |
| 56 | CHOL | autosis | ADH7 | alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide | 2.99 | 0.13 | -4.49 | 0.0 | 0.0 | 100.0 | Detail |
| 57 | CHOL | autosis | ASS1 | argininosuccinate synthase 1 | 15.64 | 11.6 | -0.43 | 0.0 | 0.0 | 100.0 | Detail |
| 58 | CHOL | autosis | AXIN1 | axin 1 | 8.85 | 10.18 | 0.2 | 0.0 | 0.0 | 100.0 | Detail |
| 59 | CHOL | autosis | DCTPP1 | dCTP pyrophosphatase 1 | 8.56 | 10.19 | 0.25 | 0.0 | 0.0 | 100.0 | Detail |
| 60 | CHOL | autosis | EXO1 | exonuclease 1 | 2.09 | 6.78 | 1.7 | 0.0 | 0.0 | 100.0 | Detail |
| 61 | CHOL | autosis | HSP90AB1 | heat shock protein 90 alpha family class B member 1 | 13.55 | 14.77 | 0.12 | 0.0 | 0.0 | 100.0 | Detail |
| 62 | CHOL | autosis | JRK | Jrk helix-turn-helix protein | 6.65 | 8.65 | 0.38 | 0.0 | 0.0 | 100.0 | Detail |
| 63 | CHOL | autosis | RHOB | ras homolog family member B | 15.2 | 12.77 | -0.25 | 0.0 | 0.0 | 100.0 | Detail |
| 64 | CHOL | autosis | SLC6A6 | solute carrier family 6 member 6 | 7.2 | 10.15 | 0.5 | 0.0 | 0.0 | 100.0 | Detail |
| 65 | CHOL | autosis | CDC45 | cell division cycle 45 | 2.71 | 7.54 | 1.48 | 0.0 | 0.0 | 100.0 | Detail |
| 66 | CHOL | autosis | SELENBP1 | selenium binding protein 1 | 13.24 | 8.98 | -0.56 | 0.0 | 0.0 | 100.0 | Detail |
| 67 | CHOL | autosis | FTL | ferritin light chain | 17.55 | 15.35 | -0.19 | 0.0 | 0.0 | 100.0 | Detail |
| 68 | CHOL | autosis | NCOR2 | nuclear receptor corepressor 2 | 10.56 | 11.92 | 0.18 | 0.0 | 0.0 | 100.0 | Detail |
| 69 | CHOL | autosis | SHMT1 | serine hydroxymethyltransferase 1 | 14.44 | 10.06 | -0.52 | 0.0 | 0.0 | 100.0 | Detail |
| 70 | CHOL | autosis | PCSK6 | proprotein convertase subtilisin/kexin type 6 | 13.41 | 9.12 | -0.56 | 0.0 | 0.0 | 100.0 | Detail |
| 71 | CHOL | autosis | PGRMC1 | progesterone receptor membrane component 1 | 13.93 | 11.3 | -0.3 | 0.0 | 0.0 | 100.0 | Detail |
| 72 | CHOL | autosis | SPC25 | SPC25 component of NDC80 kinetochore complex | 1.66 | 6.08 | 1.88 | 0.0 | 0.0 | 100.0 | Detail |
| 73 | CHOL | autosis | RFC4 | replication factor C subunit 4 | 6.94 | 8.77 | 0.34 | 0.0 | 0.0 | 100.0 | Detail |
| 74 | CHOL | autosis | DNAJB6 | DnaJ heat shock protein family (Hsp40) member B6 | 9.62 | 10.48 | 0.12 | 0.0 | 0.0 | 100.0 | Detail |
| 75 | CHOL | autosis | MCM3 | minichromosome maintenance complex component 3 | 9.27 | 11.36 | 0.29 | 0.0 | 0.0 | 100.0 | Detail |
| 76 | CHOL | autosis | N4BP2L1 | NEDD4 binding protein 2 like 1 | 10.69 | 8.04 | -0.41 | 0.0 | 0.0 | 100.0 | Detail |
| 77 | CHOL | autosis | MT1X | metallothionein 1X | 14.59 | 8.95 | -0.71 | 0.0 | 0.0 | 100.0 | Detail |
| 78 | CHOL | autosis | TRIB1 | tribbles pseudokinase 1 | 12.91 | 10.12 | -0.35 | 0.0 | 0.0 | 100.0 | Detail |
| 79 | CHOL | autosis | NUDT6 | nudix hydrolase 6 | 8.84 | 5.6 | -0.66 | 0.0 | 0.0 | 100.0 | Detail |
| 80 | CHOL | autosis | NMB | neuromedin B | 4.63 | 7.43 | 0.68 | 0.0 | 0.0 | 100.0 | Detail |
| 81 | CHOL | autosis | GABBR1 | gamma-aminobutyric acid type B receptor subunit 1 | 7.08 | 10.19 | 0.53 | 0.0 | 0.0 | 100.0 | Detail |
| 82 | CHOL | autosis | CBFA2T2 | CBFA2/RUNX1 partner transcriptional co-repressor 2 | 8.07 | 9.51 | 0.24 | 0.0 | 0.0 | 100.0 | Detail |
| 83 | CHOL | autosis | COPS7B | COP9 signalosome subunit 7B | 8.89 | 9.87 | 0.15 | 0.0 | 0.0 | 100.0 | Detail |
| 84 | CHOL | autosis | MT1H | metallothionein 1H | 12.1 | 3.54 | -1.77 | 0.0 | 0.0 | 100.0 | Detail |
| 85 | CHOL | autosis | CDH15 | cadherin 15 | 6.06 | 1.76 | -1.78 | 0.0 | 0.0 | 100.0 | Detail |
| 86 | CHOL | autosis | MCM6 | minichromosome maintenance complex component 6 | 7.87 | 9.82 | 0.32 | 0.0 | 0.0 | 100.0 | Detail |
| 87 | CHOL | autosis | EIF5 | eukaryotic translation initiation factor 5 | 12.69 | 11.73 | -0.11 | 0.0 | 0.0 | 100.0 | Detail |
| 88 | CHOL | autosis | LYPD1 | LY6/PLAUR domain containing 1 | 4.6 | 9.36 | 1.02 | 0.0 | 0.0 | 100.0 | Detail |
| 89 | CHOL | autosis | FAM8A1 | family with sequence similarity 8 member A1 | 11.8 | 10.33 | -0.19 | 0.0 | 0.0 | 100.0 | Detail |
| 90 | CHOL | autosis | CD302 | CD302 molecule | 12.99 | 8.73 | -0.57 | 0.0 | 0.0 | 100.0 | Detail |
| 91 | CHOL | autosis | CTH | cystathionine gamma-lyase | 11.31 | 5.86 | -0.95 | 0.0 | 0.0 | 100.0 | Detail |
| 92 | CHOL | autosis | GGPS1 | geranylgeranyl diphosphate synthase 1 | 8.69 | 9.55 | 0.14 | 0.0 | 0.0 | 100.0 | Detail |
| 93 | CHOL | autosis | DEPDC1B | DEP domain containing 1B | 3.84 | 8.78 | 1.19 | 0.0 | 0.0 | 100.0 | Detail |
| 94 | CHOL | autosis | GCHFR | GTP cyclohydrolase I feedback regulator | 10.83 | 7.94 | -0.45 | 0.0 | 0.0 | 100.0 | Detail |
| 95 | CHOL | autosis | GOLPH3L | golgi phosphoprotein 3 like | 8.46 | 9.9 | 0.23 | 0.0 | 0.0 | 100.0 | Detail |
| 96 | CHOL | autosis | SLC25A15 | solute carrier family 25 member 15 | 12.44 | 8.22 | -0.6 | 0.0 | 0.0 | 100.0 | Detail |
| 97 | CHOL | autosis | CLIP2 | CAP-Gly domain containing linker protein 2 | 7.87 | 11.12 | 0.5 | 0.0 | 0.0 | 100.0 | Detail |
| 98 | CHOL | autosis | TTLL4 | tubulin tyrosine ligase like 4 | 7.83 | 9.69 | 0.31 | 0.0 | 0.0 | 100.0 | Detail |
| 99 | CHOL | autosis | ISOC1 | isochorismatase domain containing 1 | 11.33 | 9.08 | -0.32 | 0.0 | 0.0 | 100.0 | Detail |
| 100 | CHOL | autosis | BDH2 | 3-hydroxybutyrate dehydrogenase 2 | 11.09 | 9.01 | -0.3 | 0.0 | 0.0 | 100.0 | Detail |
| 101 | CHOL | autosis | ADO | 2-aminoethanethiol dioxygenase | 8.52 | 9.44 | 0.15 | 0.0 | 0.0 | 100.0 | Detail |
| 102 | CHOL | autosis | CENPI | centromere protein I | 0.82 | 4.6 | 2.49 | 0.0 | 0.0 | 100.0 | Detail |
| 103 | CHOL | autosis | POLA2 | DNA polymerase alpha 2, accessory subunit | 6.94 | 8.33 | 0.26 | 0.0 | 0.0 | 100.0 | Detail |
| 104 | CHOL | autosis | LMNB2 | lamin B2 | 8.15 | 10.55 | 0.37 | 0.0 | 0.0 | 100.0 | Detail |
| 105 | CHOL | autosis | FAM13A | family with sequence similarity 13 member A | 11.41 | 7.33 | -0.64 | 0.0 | 0.0 | 100.0 | Detail |
| 106 | CHOL | autosis | ALDH5A1 | aldehyde dehydrogenase 5 family member A1 | 12.72 | 9.61 | -0.41 | 0.0 | 0.0 | 100.0 | Detail |
| 107 | CHOL | autosis | LARP4 | La ribonucleoprotein 4 | 11.43 | 9.93 | -0.2 | 0.0 | 0.0 | 100.0 | Detail |
| 108 | CHOL | autosis | PRDM4 | PR/SET domain 4 | 8.18 | 9.61 | 0.23 | 0.0 | 0.0 | 100.0 | Detail |
| 109 | CHOL | autosis | PMPCA | peptidase, mitochondrial processing subunit alpha | 11.78 | 10.15 | -0.21 | 0.0 | 0.0 | 100.0 | Detail |
| 110 | CHOL | autosis | CNNM4 | cyclin and CBS domain divalent metal cation transport mediator 4 | 7.76 | 9.63 | 0.31 | 0.0 | 0.0 | 100.0 | Detail |
| 111 | CHOL | autosis | AMPD3 | adenosine monophosphate deaminase 3 | 6.17 | 8.09 | 0.39 | 0.0 | 0.0 | 100.0 | Detail |
| 112 | CHOL | autosis | CYB5A | cytochrome b5 type A | 13.98 | 10.39 | -0.43 | 0.0 | 0.0 | 100.0 | Detail |
| 113 | CHOL | autosis | MTM1 | myotubularin 1 | 9.26 | 7.63 | -0.28 | 0.0 | 0.0 | 100.0 | Detail |
| 114 | CHOL | autosis | CCNF | cyclin F | 5.51 | 7.72 | 0.49 | 0.0 | 0.0 | 100.0 | Detail |
| 115 | CHOL | autosis | KIF18B | kinesin family member 18B | 3.1 | 7.56 | 1.29 | 0.0 | 0.0 | 100.0 | Detail |
| 116 | CHOL | autosis | CYTH2 | cytohesin 2 | 9.42 | 11.21 | 0.25 | 0.0 | 0.0 | 100.0 | Detail |
| 117 | CHOL | autosis | FBXL18 | F-box and leucine rich repeat protein 18 | 6.7 | 8.62 | 0.36 | 0.0 | 0.0 | 100.0 | Detail |
| 118 | CHOL | autosis | LCAT | lecithin-cholesterol acyltransferase | 12.59 | 7.79 | -0.69 | 0.0 | 0.0 | 100.0 | Detail |
| 119 | CHOL | autosis | MAFG | MAF bZIP transcription factor G | 7.61 | 9.36 | 0.3 | 0.0 | 0.0 | 100.0 | Detail |
| 120 | CHOL | autosis | SLC39A14 | solute carrier family 39 member 14 | 14.36 | 11.8 | -0.28 | 0.0 | 0.0 | 100.0 | Detail |
| 121 | CHOL | autosis | UBE2A | ubiquitin conjugating enzyme E2 A | 9.56 | 10.57 | 0.14 | 0.0 | 0.0 | 100.0 | Detail |
| 122 | CHOL | autosis | IL13RA1 | interleukin 13 receptor subunit alpha 1 | 12.5 | 11.42 | -0.13 | 0.0 | 0.0 | 100.0 | Detail |
| 123 | CHOL | autosis | ATP2B2 | ATPase plasma membrane Ca2+ transporting 2 | 10.64 | 5.65 | -0.91 | 0.0 | 0.0 | 100.0 | Detail |
| 124 | CHOL | autosis | ZKSCAN5 | zinc finger with KRAB and SCAN domains 5 | 7.55 | 8.61 | 0.19 | 0.0 | 0.0 | 100.0 | Detail |
| 125 | CHOL | autosis | ATP11C | ATPase phospholipid transporting 11C | 10.98 | 8.11 | -0.44 | 0.0 | 0.0 | 100.0 | Detail |
| 126 | CHOL | autosis | CCDC93 | coiled-coil domain containing 93 | 8.13 | 9.74 | 0.26 | 0.0 | 0.0 | 100.0 | Detail |
| 127 | CHOL | autosis | TTPAL | alpha tocopherol transfer protein like | 11.15 | 8.8 | -0.34 | 0.0 | 0.0 | 100.0 | Detail |
| 128 | CHOL | autosis | ITSN1 | intersectin 1 | 10.45 | 8.54 | -0.29 | 0.0 | 0.0 | 100.0 | Detail |
| 129 | CHOL | autosis | HADH | hydroxyacyl-CoA dehydrogenase | 12.54 | 9.55 | -0.39 | 0.0 | 0.0 | 100.0 | Detail |
| 130 | CHOL | autosis | BDH1 | 3-hydroxybutyrate dehydrogenase 1 | 13.09 | 7.48 | -0.81 | 0.0 | 0.0 | 100.0 | Detail |
| 131 | CHOL | autosis | BTD | biotinidase | 10.82 | 8.31 | -0.38 | 0.0 | 0.0 | 100.0 | Detail |
| 132 | CHOL | autosis | IQCC | IQ motif containing C | 4.64 | 6.46 | 0.48 | 0.0 | 0.0 | 100.0 | Detail |
| 133 | CHOL | autosis | NRBF2 | nuclear receptor binding factor 2 | 9.85 | 8.62 | -0.19 | 0.0 | 0.0 | 100.0 | Detail |
| 134 | CHOL | autosis | RCAN3 | RCAN family member 3 | 4.38 | 6.16 | 0.49 | 0.0 | 0.0 | 100.0 | Detail |
| 135 | CHOL | autosis | ACOT13 | acyl-CoA thioesterase 13 | 11.33 | 9.47 | -0.26 | 0.0 | 0.0 | 100.0 | Detail |
| 136 | CHOL | autosis | HSD17B10 | hydroxysteroid 17-beta dehydrogenase 10 | 12.02 | 10.61 | -0.18 | 0.0 | 0.0 | 100.0 | Detail |
| 137 | CHOL | autosis | GNPNAT1 | glucosamine-phosphate N-acetyltransferase 1 | 11.57 | 9.41 | -0.3 | 0.0 | 0.0 | 100.0 | Detail |
| 138 | CHOL | autosis | AP3M2 | adaptor related protein complex 3 subunit mu 2 | 7.05 | 8.96 | 0.35 | 0.0 | 0.0 | 100.0 | Detail |
| 139 | CHOL | autosis | C11orf71 | chromosome 11 open reading frame 71 | 9.4 | 7.66 | -0.29 | 0.0 | 0.0 | 100.0 | Detail |
| 140 | CHOL | autosis | CSNK1G1 | casein kinase 1 gamma 1 | 7.39 | 8.94 | 0.27 | 0.0 | 0.0 | 100.0 | Detail |
| 141 | CHOL | autosis | DHRS1 | dehydrogenase/reductase 1 | 11.98 | 8.79 | -0.45 | 0.0 | 0.0 | 100.0 | Detail |
| 142 | CHOL | autosis | ERLIN1 | ER lipid raft associated 1 | 11.74 | 9.48 | -0.31 | 0.0 | 0.0 | 100.0 | Detail |
| 143 | CHOL | autosis | MED22 | mediator complex subunit 22 | 8.02 | 9.79 | 0.29 | 0.0 | 0.0 | 100.0 | Detail |
| 144 | CHOL | autosis | PIGC | phosphatidylinositol glycan anchor biosynthesis class C | 8.68 | 9.91 | 0.19 | 0.0 | 0.0 | 100.0 | Detail |
| 145 | CHOL | autosis | SEC24B | SEC24 homolog B, COPII coat complex component | 11.03 | 9.62 | -0.2 | 0.0 | 0.0 | 100.0 | Detail |
| 146 | CHOL | autosis | STBD1 | starch binding domain 1 | 12.02 | 9.98 | -0.27 | 0.0 | 0.0 | 100.0 | Detail |
| 147 | CHOL | autosis | ATP1B3 | ATPase Na+/K+ transporting subunit beta 3 | 8.11 | 10.77 | 0.41 | 0.0 | 0.0 | 100.0 | Detail |
| 148 | CHOL | autosis | CCDC28A | coiled-coil domain containing 28A | 10.03 | 8.64 | -0.21 | 0.0 | 0.0 | 100.0 | Detail |
| 149 | CHOL | autosis | DUSP12 | dual specificity phosphatase 12 | 7.04 | 8.44 | 0.26 | 0.0 | 0.0 | 100.0 | Detail |
| 150 | CHOL | autosis | GTPBP2 | GTP binding protein 2 | 8.75 | 10.36 | 0.24 | 0.0 | 0.0 | 100.0 | Detail |
| 151 | CHOL | autosis | PXMP2 | peroxisomal membrane protein 2 | 12.02 | 8.48 | -0.5 | 0.0 | 0.0 | 100.0 | Detail |
| 152 | CHOL | autosis | RMND5A | required for meiotic nuclear division 5 homolog A | 11.84 | 10.3 | -0.2 | 0.0 | 0.0 | 100.0 | Detail |
| 153 | CHOL | autosis | RNF44 | ring finger protein 44 | 9.08 | 10.74 | 0.24 | 0.0 | 0.0 | 100.0 | Detail |
| 154 | CHOL | autosis | MKI67 | marker of proliferation Ki-67 | 6.19 | 9.84 | 0.67 | 0.0 | 0.0 | 95.83 | Detail |
| 155 | CHOL | autosis | MCM10 | minichromosome maintenance 10 replication initiation factor | 2.69 | 6.47 | 1.26 | 0.0 | 0.0 | 95.83 | Detail |
| 156 | CHOL | autosis | KIF15 | kinesin family member 15 | 2.6 | 5.9 | 1.18 | 0.0 | 0.0 | 95.83 | Detail |
| 157 | CHOL | autosis | FA2H | fatty acid 2-hydroxylase | 3.03 | 8.42 | 1.47 | 0.0 | 0.0 | 95.83 | Detail |
| 158 | CHOL | autosis | FANCF | FA complementation group F | 7.99 | 9.04 | 0.18 | 0.0 | 0.0 | 95.83 | Detail |
| 159 | CHOL | autosis | CDC25C | cell division cycle 25C | 1.96 | 6.0 | 1.61 | 0.0 | 0.0 | 95.83 | Detail |
| 160 | CHOL | autosis | TMSB15B | thymosin beta 15B | 1.19 | 4.96 | 2.06 | 0.0 | 0.0 | 95.83 | Detail |
| 161 | CHOL | autosis | AURKB | aurora kinase B | 3.47 | 7.4 | 1.09 | 0.0 | 0.0 | 95.83 | Detail |
| 162 | CHOL | autosis | PRKCD | protein kinase C delta | 8.33 | 9.38 | 0.17 | 0.0 | 0.0 | 95.83 | Detail |
| 163 | CHOL | autosis | IGFBP1 | insulin like growth factor binding protein 1 | 14.58 | 9.11 | -0.68 | 0.0 | 0.0 | 95.83 | Detail |
| 164 | CHOL | autosis | IRS1 | insulin receptor substrate 1 | 10.78 | 8.97 | -0.26 | 0.0 | 0.0 | 95.83 | Detail |
| 165 | CHOL | autosis | DLGAP4 | DLG associated protein 4 | 9.92 | 11.39 | 0.2 | 0.0 | 0.0 | 95.83 | Detail |
| 166 | CHOL | autosis | EHMT2 | euchromatic histone lysine methyltransferase 2 | 9.16 | 10.77 | 0.23 | 0.0 | 0.0 | 95.83 | Detail |
| 167 | CHOL | autosis | KANK1 | KN motif and ankyrin repeat domains 1 | 11.7 | 10.03 | -0.22 | 0.0 | 0.0 | 95.83 | Detail |
| 168 | CHOL | autosis | KLF12 | KLF transcription factor 12 | 10.0 | 7.84 | -0.35 | 0.0 | 0.0 | 95.83 | Detail |
| 169 | CHOL | autosis | NUDT1 | nudix hydrolase 1 | 6.0 | 8.02 | 0.42 | 0.0 | 0.0 | 95.83 | Detail |
| 170 | CHOL | autosis | RHBDF1 | rhomboid 5 homolog 1 | 7.59 | 10.31 | 0.44 | 0.0 | 0.0 | 95.83 | Detail |
| 171 | CHOL | autosis | CAPN2 | calpain 2 | 10.72 | 12.58 | 0.23 | 0.0 | 0.0 | 95.83 | Detail |
| 172 | CHOL | autosis | NTHL1 | nth like DNA glycosylase 1 | 9.91 | 8.51 | -0.22 | 0.0 | 0.0 | 95.83 | Detail |
| 173 | CHOL | autosis | STX1A | syntaxin 1A | 4.15 | 7.21 | 0.8 | 0.0 | 0.0 | 95.83 | Detail |
| 174 | CHOL | autosis | PGRMC2 | progesterone receptor membrane component 2 | 11.66 | 10.6 | -0.14 | 0.0 | 0.0 | 95.83 | Detail |
| 175 | CHOL | autosis | TPM4 | tropomyosin 4 | 10.99 | 13.37 | 0.28 | 0.0 | 0.0 | 95.83 | Detail |
| 176 | CHOL | autosis | THUMPD2 | THUMP domain containing 2 | 6.77 | 8.07 | 0.25 | 0.0 | 0.0 | 95.83 | Detail |
| 177 | CHOL | autosis | RALGAPB | Ral GTPase activating protein non-catalytic subunit beta | 9.44 | 10.44 | 0.14 | 0.0 | 0.0 | 95.83 | Detail |
| 178 | CHOL | autosis | CASP2 | caspase 2 | 8.13 | 9.88 | 0.28 | 0.0 | 0.0 | 95.83 | Detail |
| 179 | CHOL | autosis | CUL7 | cullin 7 | 9.26 | 10.71 | 0.21 | 0.0 | 0.0 | 95.83 | Detail |
| 180 | CHOL | autosis | USP1 | ubiquitin specific peptidase 1 | 8.67 | 9.67 | 0.16 | 0.0 | 0.0 | 95.83 | Detail |
| 181 | CHOL | autosis | OSGIN1 | oxidative stress induced growth inhibitor 1 | 12.06 | 8.41 | -0.52 | 0.0 | 0.0 | 95.83 | Detail |
| 182 | CHOL | autosis | HJURP | Holliday junction recognition protein | 3.53 | 7.55 | 1.1 | 0.0 | 0.0 | 95.83 | Detail |
| 183 | CHOL | autosis | MCM8 | minichromosome maintenance 8 homologous recombination repair factor | 5.67 | 7.71 | 0.44 | 0.0 | 0.0 | 95.83 | Detail |
| 184 | CHOL | autosis | RGS17 | regulator of G protein signaling 17 | 0.63 | 4.39 | 2.8 | 0.0 | 0.0 | 95.83 | Detail |
| 185 | CHOL | autosis | SLC30A1 | solute carrier family 30 member 1 | 9.82 | 7.54 | -0.38 | 0.0 | 0.0 | 95.83 | Detail |
| 186 | CHOL | autosis | ELOVL6 | ELOVL fatty acid elongase 6 | 10.55 | 7.25 | -0.54 | 0.0 | 0.0 | 95.83 | Detail |
| 187 | CHOL | autosis | GLRX5 | glutaredoxin 5 | 10.95 | 9.64 | -0.18 | 0.0 | 0.0 | 95.83 | Detail |
| 188 | CHOL | autosis | ACSL5 | acyl-CoA synthetase long chain family member 5 | 12.17 | 8.56 | -0.51 | 0.0 | 0.0 | 95.83 | Detail |
| 189 | CHOL | autosis | RAD1 | RAD1 checkpoint DNA exonuclease | 8.08 | 8.82 | 0.13 | 0.0 | 0.0 | 95.83 | Detail |
| 190 | CHOL | autosis | PHTF1 | putative homeodomain transcription factor 1 | 7.34 | 8.46 | 0.2 | 0.0 | 0.0 | 95.83 | Detail |
| 191 | CHOL | autosis | TGDS | TDP-glucose 4,6-dehydratase | 9.19 | 7.62 | -0.27 | 0.0 | 0.0 | 95.83 | Detail |
| 192 | CHOL | autosis | GNPDA1 | glucosamine-6-phosphate deaminase 1 | 8.58 | 9.76 | 0.19 | 0.0 | 0.0 | 95.83 | Detail |
| 193 | CHOL | autosis | GTF3C2 | general transcription factor IIIC subunit 2 | 9.29 | 10.43 | 0.17 | 0.0 | 0.0 | 95.83 | Detail |
| 194 | CHOL | autosis | OSBPL3 | oxysterol binding protein like 3 | 6.35 | 9.43 | 0.57 | 0.0 | 0.0 | 95.83 | Detail |
| 195 | CHOL | autosis | PLEKHB2 | pleckstrin homology domain containing B2 | 9.72 | 11.3 | 0.22 | 0.0 | 0.0 | 95.83 | Detail |
| 196 | CHOL | autosis | UBXN8 | UBX domain protein 8 | 9.28 | 7.97 | -0.22 | 0.0 | 0.0 | 95.83 | Detail |
| 197 | CHOL | autosis | ABL2 | ABL proto-oncogene 2, non-receptor tyrosine kinase | 8.28 | 9.63 | 0.22 | 0.0 | 0.0 | 91.67 | Detail |
| 198 | CHOL | autosis | APEX2 | apurinic/apyrimidinic endodeoxyribonuclease 2 | 8.75 | 9.52 | 0.12 | 0.0 | 0.0 | 91.67 | Detail |
| 199 | CHOL | autosis | CSNK2A1 | casein kinase 2 alpha 1 | 9.27 | 10.5 | 0.18 | 0.0 | 0.0 | 91.67 | Detail |
| 200 | CHOL | autosis | KDM5B | lysine demethylase 5B | 8.78 | 10.54 | 0.26 | 0.0 | 0.0 | 91.67 | Detail |
| 201 | CHOL | autosis | PLIN2 | perilipin 2 | 13.87 | 10.41 | -0.41 | 0.0 | 0.0 | 91.67 | Detail |
| 202 | CHOL | autosis | RRM2 | ribonucleotide reductase regulatory subunit M2 | 5.4 | 9.07 | 0.75 | 0.0 | 0.0 | 91.67 | Detail |
| 203 | CHOL | autosis | FUT4 | fucosyltransferase 4 | 7.34 | 9.61 | 0.39 | 0.0 | 0.0 | 91.67 | Detail |
| 204 | CHOL | autosis | NF2 | NF2, moesin-ezrin-radixin like (MERLIN) tumor suppressor | 8.86 | 10.13 | 0.19 | 0.0 | 0.0 | 91.67 | Detail |
| 205 | CHOL | autosis | CDCA4 | cell division cycle associated 4 | 5.95 | 7.73 | 0.38 | 0.0 | 0.0 | 91.67 | Detail |
| 206 | CHOL | autosis | FHOD1 | formin homology 2 domain containing 1 | 7.13 | 9.33 | 0.39 | 0.0 | 0.0 | 91.67 | Detail |
| 207 | CHOL | autosis | SLC7A11 | solute carrier family 7 member 11 | 2.52 | 7.16 | 1.51 | 0.0 | 0.0 | 91.67 | Detail |
| 208 | CHOL | autosis | CDC7 | cell division cycle 7 | 4.55 | 6.83 | 0.59 | 0.0 | 0.0 | 91.67 | Detail |
| 209 | CHOL | autosis | EFNA4 | ephrin A4 | 6.41 | 8.28 | 0.37 | 0.0 | 0.0 | 91.67 | Detail |
| 210 | CHOL | autosis | XBP1 | X-box binding protein 1 | 13.88 | 12.13 | -0.19 | 0.0 | 0.0 | 91.67 | Detail |
| 211 | CHOL | autosis | INSL3 | insulin like 3 | 1.01 | 3.26 | 1.69 | 0.0 | 0.0 | 91.67 | Detail |
| 212 | CHOL | autosis | CDCA3 | cell division cycle associated 3 | 3.79 | 7.15 | 0.92 | 0.0 | 0.0 | 91.67 | Detail |
| 213 | CHOL | autosis | GINS4 | GINS complex subunit 4 | 5.22 | 7.25 | 0.47 | 0.0 | 0.0 | 91.67 | Detail |
| 214 | CHOL | autosis | ETS2 | ETS proto-oncogene 2, transcription factor | 13.24 | 11.03 | -0.26 | 0.0 | 0.0 | 91.67 | Detail |
| 215 | CHOL | autosis | MECP2 | methyl-CpG binding protein 2 | 9.21 | 10.38 | 0.17 | 0.0 | 0.0 | 91.67 | Detail |
| 216 | CHOL | autosis | LEF1 | lymphoid enhancer binding factor 1 | 3.66 | 7.34 | 1.0 | 0.0 | 0.0 | 91.67 | Detail |
| 217 | CHOL | autosis | STC2 | stanniocalcin 2 | 3.98 | 7.7 | 0.95 | 0.0 | 0.0 | 91.67 | Detail |
| 218 | CHOL | autosis | LSS | lanosterol synthase | 11.9 | 9.94 | -0.26 | 0.0 | 0.0 | 91.67 | Detail |
| 219 | CHOL | autosis | PER3 | period circadian regulator 3 | 10.66 | 7.51 | -0.51 | 0.0 | 0.0 | 91.67 | Detail |
| 220 | CHOL | autosis | RND3 | Rho family GTPase 3 | 12.42 | 9.66 | -0.36 | 0.0 | 0.0 | 91.67 | Detail |
| 221 | CHOL | autosis | YWHAH | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein eta | 10.58 | 11.85 | 0.16 | 0.0 | 0.0 | 91.67 | Detail |
| 222 | CHOL | autosis | ENDOD1 | endonuclease domain containing 1 | 8.6 | 10.56 | 0.3 | 0.0 | 0.0 | 91.67 | Detail |
| 223 | CHOL | autosis | KPNB1 | karyopherin subunit beta 1 | 11.29 | 12.15 | 0.11 | 0.0 | 0.0 | 91.67 | Detail |
| 224 | CHOL | autosis | PIP4K2B | phosphatidylinositol-5-phosphate 4-kinase type 2 beta | 9.89 | 10.74 | 0.12 | 0.0 | 0.0 | 91.67 | Detail |
| 225 | CHOL | autosis | PPIF | peptidylprolyl isomerase F | 12.69 | 10.76 | -0.24 | 0.0 | 0.0 | 91.67 | Detail |
| 226 | CHOL | autosis | CLDN14 | claudin 14 | 10.35 | 6.66 | -0.64 | 0.0 | 0.0 | 91.67 | Detail |
| 227 | CHOL | autosis | RHOQ | ras homolog family member Q | 9.29 | 10.34 | 0.15 | 0.0 | 0.0 | 91.67 | Detail |
| 228 | CHOL | autosis | DERA | deoxyribose-phosphate aldolase | 10.76 | 9.37 | -0.2 | 0.0 | 0.0 | 91.67 | Detail |
| 229 | CHOL | autosis | CLCF1 | cardiotrophin like cytokine factor 1 | 5.96 | 9.11 | 0.61 | 0.0 | 0.0 | 91.67 | Detail |
| 230 | CHOL | autosis | C3orf52 | chromosome 3 open reading frame 52 | 3.5 | 7.82 | 1.16 | 0.0 | 0.0 | 91.67 | Detail |
| 231 | CHOL | autosis | IBTK | inhibitor of Bruton tyrosine kinase | 11.26 | 10.19 | -0.14 | 0.0 | 0.0 | 91.67 | Detail |
| 232 | CHOL | autosis | CDKN3 | cyclin dependent kinase inhibitor 3 | 2.87 | 6.59 | 1.2 | 0.0 | 0.0 | 87.5 | Detail |
| 233 | CHOL | autosis | CTSC | cathepsin C | 10.36 | 12.55 | 0.28 | 0.0 | 0.0 | 87.5 | Detail |
| 234 | CHOL | autosis | IL6R | interleukin 6 receptor | 10.43 | 7.68 | -0.44 | 0.0 | 0.0 | 87.5 | Detail |
| 235 | CHOL | autosis | JAG1 | jagged canonical Notch ligand 1 | 9.14 | 11.7 | 0.36 | 0.0 | 0.0 | 87.5 | Detail |
| 236 | CHOL | autosis | NAAA | N-acylethanolamine acid amidase | 10.29 | 8.82 | -0.22 | 0.0 | 0.0 | 87.5 | Detail |
| 237 | CHOL | autosis | PYCARD | PYD and CARD domain containing | 7.84 | 10.09 | 0.36 | 0.0 | 0.0 | 87.5 | Detail |
| 238 | CHOL | autosis | RIT1 | Ras like without CAAX 1 | 8.09 | 9.07 | 0.16 | 0.0 | 0.0 | 87.5 | Detail |
| 239 | CHOL | autosis | TLR4 | toll like receptor 4 | 9.03 | 7.12 | -0.34 | 0.0 | 0.0 | 87.5 | Detail |
| 240 | CHOL | autosis | CEBPD | CCAAT enhancer binding protein delta | 12.07 | 10.36 | -0.22 | 0.0 | 0.0 | 87.5 | Detail |
| 241 | CHOL | autosis | MXI1 | MAX interactor 1, dimerization protein | 10.5 | 9.26 | -0.18 | 0.0 | 0.0 | 87.5 | Detail |
| 242 | CHOL | autosis | CDK1 | cyclin dependent kinase 1 | 4.8 | 8.44 | 0.82 | 0.0 | 0.0 | 87.5 | Detail |
| 243 | CHOL | autosis | TOP2A | DNA topoisomerase II alpha | 5.45 | 9.93 | 0.87 | 0.0 | 0.0 | 87.5 | Detail |
| 244 | CHOL | autosis | RIPK2 | receptor interacting serine/threonine kinase 2 | 7.85 | 9.31 | 0.25 | 0.0 | 0.0 | 87.5 | Detail |
| 245 | CHOL | autosis | CDKN2B | cyclin dependent kinase inhibitor 2B | 5.52 | 7.73 | 0.49 | 0.0 | 0.0 | 87.5 | Detail |
| 246 | CHOL | autosis | CLU | clusterin | 17.45 | 14.85 | -0.23 | 0.0 | 0.0 | 87.5 | Detail |
| 247 | CHOL | autosis | ZNF165 | zinc finger protein 165 | 5.14 | 7.22 | 0.49 | 0.0 | 0.0 | 87.5 | Detail |
| 248 | CHOL | autosis | TMSB10 | thymosin beta 10 | 12.08 | 15.07 | 0.32 | 0.0 | 0.0 | 87.5 | Detail |
| 249 | CHOL | autosis | HELLS | helicase, lymphoid specific | 3.77 | 6.48 | 0.78 | 0.0 | 0.0 | 87.5 | Detail |
| 250 | CHOL | autosis | KIF20A | kinesin family member 20A | 3.89 | 7.87 | 1.01 | 0.0 | 0.0 | 87.5 | Detail |
| 251 | CHOL | autosis | SPPL2A | signal peptide peptidase like 2A | 10.46 | 9.28 | -0.17 | 0.0 | 0.0 | 87.5 | Detail |
| 252 | CHOL | autosis | STK24 | serine/threonine kinase 24 | 10.37 | 11.54 | 0.15 | 0.0 | 0.0 | 87.5 | Detail |
| 253 | CHOL | autosis | ZBTB4 | zinc finger and BTB domain containing 4 | 9.48 | 10.87 | 0.2 | 0.0 | 0.0 | 87.5 | Detail |
| 254 | CHOL | autosis | CDR2 | cerebellar degeneration related protein 2 | 9.09 | 10.16 | 0.16 | 0.0 | 0.0 | 87.5 | Detail |
| 255 | CHOL | autosis | ZFHX4 | zinc finger homeobox 4 | 8.67 | 5.27 | -0.72 | 0.0 | 0.0 | 87.5 | Detail |
| 256 | CHOL | autosis | TNFSF9 | TNF superfamily member 9 | 2.4 | 5.26 | 1.13 | 0.0 | 0.0 | 87.5 | Detail |
| 257 | CHOL | autosis | VOPP1 | VOPP1 WW domain binding protein | 8.93 | 10.13 | 0.18 | 0.0 | 0.0 | 87.5 | Detail |
| 258 | CHOL | autosis | CRADD | CASP2 and RIPK1 domain containing adaptor with death domain | 9.05 | 7.74 | -0.23 | 0.0 | 0.0 | 87.5 | Detail |
| 259 | CHOL | autosis | MAP1S | microtubule associated protein 1S | 8.92 | 10.16 | 0.19 | 0.0 | 0.0 | 87.5 | Detail |
| 260 | CHOL | autosis | DTYMK | deoxythymidylate kinase | 8.08 | 9.39 | 0.22 | 0.0 | 0.0 | 87.5 | Detail |
| 261 | CHOL | autosis | MCCC2 | methylcrotonyl-CoA carboxylase subunit 2 | 12.44 | 11.11 | -0.16 | 0.0 | 0.0 | 87.5 | Detail |
| 262 | CHOL | autosis | RBM8A | RNA binding motif protein 8A | 10.05 | 10.76 | 0.1 | 0.0 | 0.0 | 87.5 | Detail |
| 263 | CHOL | autosis | PLXNA3 | plexin A3 | 7.26 | 9.87 | 0.44 | 0.0 | 0.0 | 87.5 | Detail |
| 264 | CHOL | autosis | GJC1 | gap junction protein gamma 1 | 4.63 | 7.59 | 0.71 | 0.0 | 0.0 | 87.5 | Detail |
| 265 | CHOL | autosis | RAB11FIP1 | RAB11 family interacting protein 1 | 9.42 | 11.6 | 0.3 | 0.0 | 0.0 | 87.5 | Detail |
| 266 | CHOL | autosis | KIF3C | kinesin family member 3C | 5.82 | 9.05 | 0.64 | 0.0 | 0.0 | 87.5 | Detail |
| 267 | CHOL | autosis | STX3 | syntaxin 3 | 8.12 | 9.85 | 0.28 | 0.0 | 0.0 | 87.5 | Detail |
| 268 | CHOL | autosis | AGBL5 | AGBL carboxypeptidase 5 | 8.39 | 9.5 | 0.18 | 0.0 | 0.0 | 87.5 | Detail |
| 269 | CHOL | autosis | DNAJB2 | DnaJ heat shock protein family (Hsp40) member B2 | 10.68 | 11.55 | 0.11 | 0.0 | 0.0 | 87.5 | Detail |
| 270 | CHOL | autosis | FGD6 | FYVE, RhoGEF and PH domain containing 6 | 7.78 | 10.21 | 0.39 | 0.0 | 0.0 | 87.5 | Detail |
| 271 | CHOL | autosis | TNPO2 | transportin 2 | 9.8 | 10.7 | 0.13 | 0.0 | 0.0 | 87.5 | Detail |
| 272 | CHOL | autosis | ABCA1 | ATP binding cassette subfamily A member 1 | 11.81 | 8.66 | -0.45 | 0.0 | 0.0 | 83.33 | Detail |
| 273 | CHOL | autosis | ATR | ATR serine/threonine kinase | 8.26 | 9.4 | 0.19 | 0.0 | 0.0 | 83.33 | Detail |
| 274 | CHOL | autosis | IGFBP3 | insulin like growth factor binding protein 3 | 14.09 | 10.87 | -0.37 | 0.0 | 0.0 | 83.33 | Detail |
| 275 | CHOL | autosis | PMAIP1 | phorbol-12-myristate-13-acetate-induced protein 1 | 3.88 | 6.86 | 0.82 | 0.0 | 0.0 | 83.33 | Detail |
| 276 | CHOL | autosis | POLD1 | DNA polymerase delta 1, catalytic subunit | 8.14 | 9.29 | 0.19 | 0.0 | 0.0 | 83.33 | Detail |
| 277 | CHOL | autosis | RNMT | RNA guanine-7 methyltransferase | 8.99 | 9.88 | 0.14 | 0.0 | 0.0 | 83.33 | Detail |
| 278 | CHOL | autosis | SIRT7 | sirtuin 7 | 8.13 | 9.17 | 0.17 | 0.0 | 0.0 | 83.33 | Detail |
| 279 | CHOL | autosis | SLC6A8 | solute carrier family 6 member 8 | 7.05 | 9.92 | 0.49 | 0.0 | 0.0 | 83.33 | Detail |
| 280 | CHOL | autosis | CCNE1 | cyclin E1 | 3.76 | 7.03 | 0.9 | 0.0 | 0.0 | 83.33 | Detail |
| 281 | CHOL | autosis | TUG1 | taurine up-regulated 1 | 10.91 | 11.98 | 0.13 | 0.0 | 0.0 | 83.33 | Detail |
| 282 | CHOL | autosis | NQO1 | NAD(P)H quinone dehydrogenase 1 | 6.28 | 10.83 | 0.78 | 0.0 | 0.0 | 83.33 | Detail |
| 283 | CHOL | autosis | DBI | diazepam binding inhibitor, acyl-CoA binding protein | 12.66 | 11.4 | -0.15 | 0.0 | 0.0 | 83.33 | Detail |
| 284 | CHOL | autosis | HPGD | 15-hydroxyprostaglandin dehydrogenase | 12.75 | 6.27 | -1.02 | 0.0 | 0.0 | 83.33 | Detail |
| 285 | CHOL | autosis | CDCA7 | cell division cycle associated 7 | 2.67 | 7.07 | 1.41 | 0.0 | 0.0 | 83.33 | Detail |
| 286 | CHOL | autosis | JMJD6 | jumonji domain containing 6, arginine demethylase and lysine hydroxylase | 8.41 | 9.39 | 0.16 | 0.0 | 0.0 | 83.33 | Detail |
| 287 | CHOL | autosis | BLCAP | BLCAP apoptosis inducing factor | 11.31 | 10.45 | -0.11 | 0.0 | 0.0 | 83.33 | Detail |
| 288 | CHOL | autosis | SMURF1 | SMAD specific E3 ubiquitin protein ligase 1 | 9.43 | 10.66 | 0.18 | 0.0 | 0.0 | 83.33 | Detail |
| 289 | CHOL | autosis | TICAM1 | TIR domain containing adaptor molecule 1 | 9.26 | 10.31 | 0.15 | 0.0 | 0.0 | 83.33 | Detail |
| 290 | CHOL | autosis | IL11 | interleukin 11 | 0.82 | 4.38 | 2.42 | 0.0 | 0.0 | 83.33 | Detail |
| 291 | CHOL | autosis | PRSS23 | serine protease 23 | 9.29 | 11.19 | 0.27 | 0.0 | 0.0 | 83.33 | Detail |
| 292 | CHOL | autosis | TUBA1B | tubulin alpha 1b | 12.25 | 13.61 | 0.15 | 0.0 | 0.0 | 83.33 | Detail |
| 293 | CHOL | autosis | ABCC5 | ATP binding cassette subfamily C member 5 | 7.83 | 9.5 | 0.28 | 0.0 | 0.0 | 83.33 | Detail |
| 294 | CHOL | autosis | SLC38A2 | solute carrier family 38 member 2 | 13.76 | 11.81 | -0.22 | 0.0 | 0.0 | 83.33 | Detail |
| 295 | CHOL | autosis | AADAC | arylacetamide deacetylase | 13.21 | 8.03 | -0.72 | 0.0 | 0.0 | 83.33 | Detail |
| 296 | CHOL | autosis | CAPNS1 | calpain small subunit 1 | 12.09 | 13.02 | 0.11 | 0.0 | 0.0 | 83.33 | Detail |
| 297 | CHOL | autosis | GLRX | glutaredoxin | 10.7 | 8.59 | -0.32 | 0.0 | 0.0 | 83.33 | Detail |
| 298 | CHOL | autosis | BAZ2A | bromodomain adjacent to zinc finger domain 2A | 10.05 | 11.24 | 0.16 | 0.0 | 0.0 | 83.33 | Detail |
| 299 | CHOL | autosis | FECH | ferrochelatase | 10.26 | 9.03 | -0.18 | 0.0 | 0.0 | 83.33 | Detail |
| 300 | CHOL | autosis | ZNF365 | zinc finger protein 365 | 1.46 | 4.51 | 1.63 | 0.0 | 0.0 | 83.33 | Detail |
| 301 | CHOL | autosis | TMED5 | transmembrane p24 trafficking protein 5 | 11.5 | 10.31 | -0.16 | 0.0 | 0.0 | 83.33 | Detail |
| 302 | CHOL | autosis | PPP1R3C | protein phosphatase 1 regulatory subunit 3C | 10.07 | 6.95 | -0.53 | 0.0 | 0.0 | 83.33 | Detail |
| 303 | CHOL | autosis | TP53I11 | tumor protein p53 inducible protein 11 | 8.79 | 10.61 | 0.27 | 0.0 | 0.0 | 83.33 | Detail |
| 304 | CHOL | autosis | ZNF202 | zinc finger protein 202 | 6.84 | 7.87 | 0.2 | 0.0 | 0.0 | 83.33 | Detail |
| 305 | CHOL | autosis | SLC39A8 | solute carrier family 39 member 8 | 10.28 | 7.72 | -0.41 | 0.0 | 0.0 | 83.33 | Detail |
| 306 | CHOL | autosis | TUBA1A | tubulin alpha 1a | 9.04 | 11.46 | 0.34 | 0.0 | 0.0 | 83.33 | Detail |
| 307 | CHOL | autosis | ATP8B4 | ATPase phospholipid transporting 8B4 (putative) | 6.99 | 5.38 | -0.38 | 0.0 | 0.0 | 83.33 | Detail |
| 308 | CHOL | autosis | C1orf216 | chromosome 1 open reading frame 216 | 7.41 | 8.66 | 0.23 | 0.0 | 0.0 | 83.33 | Detail |
| 309 | CHOL | autosis | CDR2L | cerebellar degeneration related protein 2 like | 6.66 | 10.07 | 0.6 | 0.0 | 0.0 | 83.33 | Detail |
| 310 | CHOL | autosis | GK | glycerol kinase | 10.05 | 8.4 | -0.26 | 0.0 | 0.0 | 83.33 | Detail |
| 311 | CHOL | autosis | GMEB2 | glucocorticoid modulatory element binding protein 2 | 8.32 | 9.1 | 0.13 | 0.0 | 0.0 | 83.33 | Detail |
| 312 | CHOL | autosis | MGST2 | microsomal glutathione S-transferase 2 | 11.75 | 9.82 | -0.26 | 0.0 | 0.0 | 83.33 | Detail |
| 313 | CHOL | autosis | NSMAF | neutral sphingomyelinase activation associated factor | 7.75 | 8.98 | 0.21 | 0.0 | 0.0 | 83.33 | Detail |
| 314 | CHOL | autosis | ARPC1B | actin related protein 2/3 complex subunit 1B | 10.69 | 12.32 | 0.2 | 0.0 | 0.0 | 79.17 | Detail |
| 315 | CHOL | autosis | CALD1 | caldesmon 1 | 13.18 | 12.33 | -0.1 | 0.0 | 0.0 | 79.17 | Detail |
| 316 | CHOL | autosis | PITX1 | paired like homeodomain 1 | 1.36 | 7.08 | 2.38 | 0.0 | 0.0 | 79.17 | Detail |
| 317 | CHOL | autosis | SERPINH1 | serpin family H member 1 | 10.32 | 11.78 | 0.19 | 0.0 | 0.0 | 79.17 | Detail |
| 318 | CHOL | autosis | TFPI | tissue factor pathway inhibitor | 12.85 | 9.87 | -0.38 | 0.0 | 0.0 | 79.17 | Detail |
| 319 | CHOL | autosis | EIF5A2 | eukaryotic translation initiation factor 5A2 | 5.85 | 7.38 | 0.34 | 0.0 | 0.0 | 79.17 | Detail |
| 320 | CHOL | autosis | FGFR1 | fibroblast growth factor receptor 1 | 8.7 | 11.1 | 0.35 | 0.0 | 0.0 | 79.17 | Detail |
| 321 | CHOL | autosis | ANXA10 | annexin A10 | 10.54 | 3.87 | -1.45 | 0.0 | 0.0 | 79.17 | Detail |
| 322 | CHOL | autosis | CCNG2 | cyclin G2 | 7.93 | 9.42 | 0.25 | 0.0 | 0.0 | 79.17 | Detail |
| 323 | CHOL | autosis | DOK1 | docking protein 1 | 6.59 | 8.36 | 0.34 | 0.0 | 0.0 | 79.17 | Detail |
| 324 | CHOL | autosis | FZD1 | frizzled class receptor 1 | 8.28 | 9.78 | 0.24 | 0.0 | 0.0 | 79.17 | Detail |
| 325 | CHOL | autosis | RASAL2 | RAS protein activator like 2 | 8.12 | 10.25 | 0.34 | 0.0 | 0.0 | 79.17 | Detail |
| 326 | CHOL | autosis | CTTN | cortactin | 11.36 | 12.46 | 0.13 | 0.0 | 0.0 | 79.17 | Detail |
| 327 | CHOL | autosis | CTNNA1 | catenin alpha 1 | 12.27 | 13.34 | 0.12 | 0.0 | 0.0 | 79.17 | Detail |
| 328 | CHOL | autosis | KLF9 | KLF transcription factor 9 | 12.47 | 10.14 | -0.3 | 0.0 | 0.0 | 79.17 | Detail |
| 329 | CHOL | autosis | SLC26A2 | solute carrier family 26 member 2 | 6.79 | 8.7 | 0.36 | 0.0 | 0.0 | 79.17 | Detail |
| 330 | CHOL | autosis | ABI1 | abl interactor 1 | 9.66 | 10.54 | 0.13 | 0.0 | 0.0 | 79.17 | Detail |
| 331 | CHOL | autosis | NME2 | NME/NM23 nucleoside diphosphate kinase 2 | 11.94 | 13.23 | 0.15 | 0.0 | 0.0 | 79.17 | Detail |
| 332 | CHOL | autosis | FJX1 | four-jointed box kinase 1 | 5.65 | 8.3 | 0.56 | 0.0 | 0.0 | 79.17 | Detail |
| 333 | CHOL | autosis | ATP1A1 | ATPase Na+/K+ transporting subunit alpha 1 | 12.82 | 15.48 | 0.27 | 0.0 | 0.0 | 79.17 | Detail |
| 334 | CHOL | autosis | CDYL | chromodomain Y like | 7.64 | 8.57 | 0.17 | 0.0 | 0.0 | 79.17 | Detail |
| 335 | CHOL | autosis | EZH1 | enhancer of zeste 1 polycomb repressive complex 2 subunit | 8.68 | 9.71 | 0.16 | 0.0 | 0.0 | 79.17 | Detail |
| 336 | CHOL | autosis | RAB38 | RAB38, member RAS oncogene family | 3.78 | 6.61 | 0.81 | 0.0 | 0.0 | 79.17 | Detail |
| 337 | CHOL | autosis | NCOA6 | nuclear receptor coactivator 6 | 9.46 | 10.37 | 0.13 | 0.0 | 0.0 | 79.17 | Detail |
| 338 | CHOL | autosis | ATP10A | ATPase phospholipid transporting 10A (putative) | 5.03 | 7.39 | 0.55 | 0.0 | 0.0 | 79.17 | Detail |
| 339 | CHOL | autosis | SLC25A36 | solute carrier family 25 member 36 | 7.36 | 9.33 | 0.34 | 0.0 | 0.0 | 79.17 | Detail |
| 340 | CHOL | autosis | DHFR | dihydrofolate reductase | 9.63 | 8.31 | -0.21 | 0.0 | 0.0 | 79.17 | Detail |
| 341 | CHOL | autosis | NAA40 | N-alpha-acetyltransferase 40, NatD catalytic subunit | 7.45 | 8.98 | 0.27 | 0.0 | 0.0 | 79.17 | Detail |
| 342 | CHOL | autosis | GNAI3 | G protein subunit alpha i3 | 10.55 | 11.26 | 0.09 | 0.0 | 0.0 | 79.17 | Detail |
| 343 | CHOL | autosis | BTN2A1 | butyrophilin subfamily 2 member A1 | 8.14 | 9.41 | 0.21 | 0.0 | 0.0 | 79.17 | Detail |
| 344 | CHOL | autosis | MEGF9 | multiple EGF like domains 9 | 11.01 | 8.71 | -0.34 | 0.0 | 0.0 | 79.17 | Detail |
| 345 | CHOL | autosis | MKRN1 | makorin ring finger protein 1 | 10.56 | 11.22 | 0.09 | 0.0 | 0.0 | 79.17 | Detail |
| 346 | CHOL | autosis | SLC25A38 | solute carrier family 25 member 38 | 10.28 | 9.14 | -0.17 | 0.0 | 0.0 | 79.17 | Detail |
| 347 | CHOL | autosis | TCTA | T cell leukemia translocation altered | 10.21 | 9.42 | -0.12 | 0.0 | 0.0 | 79.17 | Detail |
| 348 | CHOL | autosis | ZNF174 | zinc finger protein 174 | 7.18 | 7.93 | 0.14 | 0.0 | 0.0 | 79.17 | Detail |
| 349 | CHOL | autosis | ARMCX6 | armadillo repeat containing X-linked 6 | 6.06 | 7.63 | 0.33 | 0.0 | 0.0 | 79.17 | Detail |
| 350 | CHOL | autosis | MED17 | mediator complex subunit 17 | 8.18 | 9.12 | 0.16 | 0.0 | 0.0 | 79.17 | Detail |
| 351 | CHOL | autosis | TANC2 | tetratricopeptide repeat, ankyrin repeat and coiled-coil containing 2 | 6.83 | 9.42 | 0.46 | 0.0 | 0.0 | 75.0 | Detail |
| 352 | CHOL | autosis | TSPO | translocator protein | 9.94 | 11.58 | 0.22 | 0.0 | 0.0 | 75.0 | Detail |
| 353 | CHOL | autosis | DAXX | death domain associated protein | 9.73 | 10.67 | 0.13 | 0.0 | 0.0 | 75.0 | Detail |
| 354 | CHOL | autosis | HRAS | HRas proto-oncogene, GTPase | 8.31 | 9.42 | 0.18 | 0.0 | 0.0 | 75.0 | Detail |
| 355 | CHOL | autosis | TGFB2 | transforming growth factor beta 2 | 5.75 | 9.7 | 0.76 | 0.0 | 0.0 | 75.0 | Detail |
| 356 | CHOL | autosis | LRP8 | LDL receptor related protein 8 | 2.79 | 6.12 | 1.13 | 0.0 | 0.0 | 75.0 | Detail |
| 357 | CHOL | autosis | MAPK7 | mitogen-activated protein kinase 7 | 7.62 | 8.61 | 0.18 | 0.0 | 0.0 | 75.0 | Detail |
| 358 | CHOL | autosis | ARHGEF2 | Rho/Rac guanine nucleotide exchange factor 2 | 8.39 | 10.23 | 0.29 | 0.0 | 0.0 | 75.0 | Detail |
| 359 | CHOL | autosis | ITGB5 | integrin subunit beta 5 | 11.43 | 12.61 | 0.14 | 0.0 | 0.0 | 75.0 | Detail |
| 360 | CHOL | autosis | IER3 | immediate early response 3 | 9.0 | 11.06 | 0.3 | 0.0 | 0.0 | 75.0 | Detail |
| 361 | CHOL | autosis | HOPX | HOP homeobox | 5.01 | 7.76 | 0.63 | 0.0 | 0.0 | 75.0 | Detail |
| 362 | CHOL | autosis | TRAF2 | TNF receptor associated factor 2 | 8.32 | 9.53 | 0.2 | 0.0 | 0.0 | 75.0 | Detail |
| 363 | CHOL | autosis | PKNOX1 | PBX/knotted 1 homeobox 1 | 7.76 | 8.65 | 0.16 | 0.0 | 0.0 | 75.0 | Detail |
| 364 | CHOL | autosis | DYNLL1 | dynein light chain LC8-type 1 | 11.07 | 11.76 | 0.09 | 0.0 | 0.0 | 75.0 | Detail |
| 365 | CHOL | autosis | CDH3 | cadherin 3 | 2.03 | 5.36 | 1.4 | 0.0 | 0.0 | 75.0 | Detail |
| 366 | CHOL | autosis | MMP10 | matrix metallopeptidase 10 | 0.79 | 4.92 | 2.63 | 0.0 | 0.01 | 75.0 | Detail |
| 367 | CHOL | autosis | TRIM36 | tripartite motif containing 36 | 2.8 | 4.79 | 0.78 | 0.0 | 0.0 | 75.0 | Detail |
| 368 | CHOL | autosis | MCM4 | minichromosome maintenance complex component 4 | 8.14 | 9.98 | 0.29 | 0.0 | 0.0 | 75.0 | Detail |
| 369 | CHOL | autosis | ANP32A | acidic nuclear phosphoprotein 32 family member A | 10.91 | 11.58 | 0.09 | 0.0 | 0.0 | 75.0 | Detail |
| 370 | CHOL | autosis | UBE2D1 | ubiquitin conjugating enzyme E2 D1 | 7.86 | 8.59 | 0.13 | 0.0 | 0.0 | 75.0 | Detail |
| 371 | CHOL | autosis | MTHFD2 | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 2, methenyltetrahydrofolate cyclohydrolase | 6.13 | 8.74 | 0.51 | 0.0 | 0.0 | 75.0 | Detail |
| 372 | CHOL | autosis | WDR76 | WD repeat domain 76 | 4.37 | 7.07 | 0.7 | 0.0 | 0.0 | 75.0 | Detail |
| 373 | CHOL | autosis | UAP1L1 | UDP-N-acetylglucosamine pyrophosphorylase 1 like 1 | 6.44 | 8.21 | 0.35 | 0.0 | 0.0 | 75.0 | Detail |
| 374 | CHOL | autosis | ADNP2 | ADNP homeobox 2 | 8.27 | 9.04 | 0.13 | 0.0 | 0.0 | 75.0 | Detail |
| 375 | CHOL | autosis | IQCB1 | IQ motif containing B1 | 7.18 | 8.51 | 0.25 | 0.0 | 0.0 | 75.0 | Detail |
| 376 | CHOL | autosis | TMEM109 | transmembrane protein 109 | 10.72 | 11.73 | 0.13 | 0.0 | 0.0 | 75.0 | Detail |
| 377 | CHOL | autosis | UPF3B | UPF3B regulator of nonsense mediated mRNA decay | 7.39 | 8.37 | 0.18 | 0.0 | 0.01 | 75.0 | Detail |
| 378 | CHOL | autosis | WDR47 | WD repeat domain 47 | 6.63 | 7.96 | 0.27 | 0.0 | 0.0 | 75.0 | Detail |
| 379 | CHOL | autosis | ZNF282 | zinc finger protein 282 | 8.98 | 9.9 | 0.14 | 0.0 | 0.0 | 75.0 | Detail |
| 380 | CHOL | autosis | DHX34 | DExH-box helicase 34 | 8.15 | 9.33 | 0.2 | 0.0 | 0.0 | 75.0 | Detail |
| 381 | CHOL | autosis | FAM53C | family with sequence similarity 53 member C | 8.92 | 9.82 | 0.14 | 0.0 | 0.0 | 75.0 | Detail |
| 382 | CHOL | autosis | FARSA | phenylalanyl-tRNA synthetase subunit alpha | 10.01 | 10.62 | 0.09 | 0.0 | 0.0 | 75.0 | Detail |
| 383 | CHOL | autosis | FNBP4 | formin binding protein 4 | 9.27 | 10.15 | 0.13 | 0.0 | 0.0 | 75.0 | Detail |
| 384 | CHOL | autosis | HDDC2 | HD domain containing 2 | 8.12 | 9.16 | 0.17 | 0.0 | 0.0 | 75.0 | Detail |
| 385 | CHOL | autosis | RNF19B | ring finger protein 19B | 8.13 | 9.2 | 0.18 | 0.0 | 0.0 | 75.0 | Detail |
| 386 | CHOL | autosis | CADM1 | cell adhesion molecule 1 | 10.89 | 9.36 | -0.22 | 0.0 | 0.0 | 75.0 | Detail |
| 387 | CHOL | autosis | RORA | RAR related orphan receptor A | 10.66 | 8.61 | -0.31 | 0.0 | 0.0 | 75.0 | Detail |
| 388 | CHOL | autosis | IGFBP2 | insulin like growth factor binding protein 2 | 12.79 | 9.26 | -0.47 | 0.0 | 0.0 | 75.0 | Detail |
| 389 | CHOL | autosis | RAB27A | RAB27A, member RAS oncogene family | 9.56 | 8.45 | -0.18 | 0.0 | 0.0 | 75.0 | Detail |
| 390 | CHOL | autosis | TXNRD2 | thioredoxin reductase 2 | 10.95 | 9.46 | -0.21 | 0.0 | 0.0 | 75.0 | Detail |
| 391 | CHOL | autosis | GADD45A | growth arrest and DNA damage inducible alpha | 11.79 | 9.75 | -0.27 | 0.0 | 0.0 | 75.0 | Detail |
| 392 | CHOL | autosis | NNMT | nicotinamide N-methyltransferase | 14.59 | 11.57 | -0.34 | 0.0 | 0.0 | 75.0 | Detail |
| 393 | CHOL | autosis | RBL2 | RB transcriptional corepressor like 2 | 11.35 | 10.39 | -0.13 | 0.0 | 0.0 | 75.0 | Detail |
| 394 | CHOL | autosis | NR1D2 | nuclear receptor subfamily 1 group D member 2 | 10.5 | 8.93 | -0.23 | 0.0 | 0.0 | 75.0 | Detail |
| 395 | CHOL | autosis | ALDH1B1 | aldehyde dehydrogenase 1 family member B1 | 13.24 | 10.45 | -0.34 | 0.0 | 0.0 | 75.0 | Detail |
| 396 | CHOL | autosis | SH2D4A | SH2 domain containing 4A | 9.92 | 8.77 | -0.18 | 0.0 | 0.0 | 75.0 | Detail |
| 397 | CHOL | autosis | C6orf120 | chromosome 6 open reading frame 120 | 10.22 | 9.12 | -0.16 | 0.0 | 0.0 | 75.0 | Detail |
| 398 | CHOL | autosis | DNAJB9 | DnaJ heat shock protein family (Hsp40) member B9 | 11.3 | 9.78 | -0.21 | 0.0 | 0.0 | 75.0 | Detail |
| 399 | CHOL | autosis | MRPL34 | mitochondrial ribosomal protein L34 | 11.18 | 9.88 | -0.18 | 0.0 | 0.0 | 75.0 | Detail |
| 400 | CHOL | autosis | ALDH3B2 | aldehyde dehydrogenase 3 family member B2 | 0.54 | 4.98 | 3.2 | 0.0 | 0.01 | 72.92 | Detail |
| 401 | CHOL | autosis | APOL1 | apolipoprotein L1 | 13.34 | 11.36 | -0.23 | 0.0 | 0.0 | 70.83 | Detail |
| 402 | CHOL | autosis | CENPJ | centromere protein J | 5.79 | 6.96 | 0.27 | 0.0 | 0.0 | 70.83 | Detail |
| 403 | CHOL | autosis | COL4A2 | collagen type IV alpha 2 chain | 11.16 | 13.5 | 0.28 | 0.0 | 0.0 | 70.83 | Detail |
| 404 | CHOL | autosis | AURKA | aurora kinase A | 5.91 | 7.97 | 0.43 | 0.0 | 0.0 | 70.83 | Detail |
| 405 | CHOL | autosis | CEBPB | CCAAT enhancer binding protein beta | 12.24 | 10.68 | -0.2 | 0.0 | 0.0 | 70.83 | Detail |
| 406 | CHOL | autosis | ITGA6 | integrin subunit alpha 6 | 9.51 | 10.74 | 0.18 | 0.0 | 0.0 | 70.83 | Detail |
| 407 | CHOL | autosis | WWC1 | WW and C2 domain containing 1 | 9.52 | 11.47 | 0.27 | 0.0 | 0.0 | 70.83 | Detail |
| 408 | CHOL | autosis | MCM5 | minichromosome maintenance complex component 5 | 9.29 | 10.56 | 0.18 | 0.0 | 0.0 | 70.83 | Detail |
| 409 | CHOL | autosis | RRS1 | ribosome biogenesis regulator 1 homolog | 8.14 | 9.12 | 0.16 | 0.0 | 0.0 | 70.83 | Detail |
| 410 | CHOL | autosis | FUCA1 | alpha-L-fucosidase 1 | 11.37 | 9.83 | -0.21 | 0.0 | 0.0 | 70.83 | Detail |
| 411 | CHOL | autosis | IL13RA2 | interleukin 13 receptor subunit alpha 2 | 6.01 | 2.06 | -1.55 | 0.0 | 0.01 | 70.83 | Detail |
| 412 | CHOL | autosis | ADCY3 | adenylate cyclase 3 | 7.51 | 9.22 | 0.3 | 0.0 | 0.0 | 70.83 | Detail |
| 413 | CHOL | autosis | TGIF2 | TGFB induced factor homeobox 2 | 8.61 | 9.66 | 0.17 | 0.0 | 0.0 | 70.83 | Detail |
| 414 | CHOL | autosis | VANGL1 | VANGL planar cell polarity protein 1 | 7.85 | 9.28 | 0.24 | 0.0 | 0.0 | 70.83 | Detail |
| 415 | CHOL | autosis | CCNJ | cyclin J | 6.44 | 7.34 | 0.19 | 0.0 | 0.0 | 70.83 | Detail |
| 416 | CHOL | autosis | PGAP3 | post-GPI attachment to proteins phospholipase 3 | 10.96 | 9.93 | -0.14 | 0.0 | 0.0 | 70.83 | Detail |
| 417 | CHOL | autosis | ZNF143 | zinc finger protein 143 | 7.44 | 8.24 | 0.15 | 0.0 | 0.0 | 70.83 | Detail |
| 418 | CHOL | autosis | NID1 | nidogen 1 | 12.13 | 10.75 | -0.17 | 0.0 | 0.0 | 70.83 | Detail |
| 419 | CHOL | autosis | MAP4K3 | mitogen-activated protein kinase kinase kinase kinase 3 | 9.06 | 10.54 | 0.22 | 0.0 | 0.0 | 70.83 | Detail |
| 420 | CHOL | autosis | RHOF | ras homolog family member F, filopodia associated | 5.07 | 8.68 | 0.77 | 0.0 | 0.0 | 70.83 | Detail |
| 421 | CHOL | autosis | CRIM1 | cysteine rich transmembrane BMP regulator 1 | 9.61 | 11.59 | 0.27 | 0.0 | 0.0 | 70.83 | Detail |
| 422 | CHOL | autosis | RABIF | RAB interacting factor | 7.79 | 8.56 | 0.14 | 0.0 | 0.0 | 70.83 | Detail |
| 423 | CHOL | autosis | SERINC1 | serine incorporator 1 | 12.82 | 11.69 | -0.13 | 0.0 | 0.0 | 70.83 | Detail |
| 424 | CHOL | autosis | TRADD | TNFRSF1A associated via death domain | 8.94 | 9.84 | 0.14 | 0.0 | 0.0 | 70.83 | Detail |
| 425 | CHOL | autosis | DEAF1 | DEAF1 transcription factor | 8.31 | 9.4 | 0.18 | 0.0 | 0.0 | 70.83 | Detail |
| 426 | CHOL | autosis | TCF20 | transcription factor 20 | 9.31 | 10.12 | 0.12 | 0.0 | 0.0 | 70.83 | Detail |
| 427 | CHOL | autosis | BPGM | bisphosphoglycerate mutase | 8.04 | 8.9 | 0.15 | 0.0 | 0.0 | 70.83 | Detail |
| 428 | CHOL | autosis | GOLGA4 | golgin A4 | 11.39 | 10.29 | -0.15 | 0.0 | 0.0 | 70.83 | Detail |
| 429 | CHOL | autosis | GPATCH8 | G-patch domain containing 8 | 9.21 | 10.17 | 0.14 | 0.0 | 0.0 | 70.83 | Detail |
| 430 | CHOL | autosis | GPRC5B | G protein-coupled receptor class C group 5 member B | 8.29 | 10.34 | 0.32 | 0.0 | 0.0 | 70.83 | Detail |
| 431 | CHOL | autosis | IFRD2 | interferon related developmental regulator 2 | 10.66 | 9.82 | -0.12 | 0.0 | 0.0 | 70.83 | Detail |
| 432 | CHOL | autosis | RNFT2 | ring finger protein, transmembrane 2 | 2.59 | 6.02 | 1.22 | 0.0 | 0.0 | 70.83 | Detail |
| 433 | CHOL | autosis | SEC24A | SEC24 homolog A, COPII coat complex component | 10.8 | 9.31 | -0.21 | 0.0 | 0.0 | 70.83 | Detail |
| 434 | CHOL | autosis | SYNGR3 | synaptogyrin 3 | 2.51 | 6.32 | 1.33 | 0.0 | 0.0 | 70.83 | Detail |
| 435 | CHOL | autosis | CKLF | chemokine like factor | 7.4 | 8.67 | 0.23 | 0.0 | 0.0 | 66.67 | Detail |
| 436 | CHOL | autosis | HSPB1 | heat shock protein family B (small) member 1 | 12.13 | 13.61 | 0.17 | 0.0 | 0.0 | 66.67 | Detail |
| 437 | CHOL | autosis | PDGFB | platelet derived growth factor subunit B | 8.0 | 9.62 | 0.27 | 0.0 | 0.0 | 66.67 | Detail |
| 438 | CHOL | autosis | RTN2 | reticulon 2 | 6.09 | 8.11 | 0.41 | 0.0 | 0.01 | 66.67 | Detail |
| 439 | CHOL | autosis | SNRPE | small nuclear ribonucleoprotein polypeptide E | 8.85 | 10.06 | 0.18 | 0.0 | 0.0 | 66.67 | Detail |
| 440 | CHOL | autosis | SOX9 | SRY-box transcription factor 9 | 8.7 | 11.25 | 0.37 | 0.0 | 0.0 | 66.67 | Detail |
| 441 | CHOL | autosis | BAX | BCL2 associated X, apoptosis regulator | 9.41 | 10.58 | 0.17 | 0.0 | 0.0 | 66.67 | Detail |
| 442 | CHOL | autosis | DUSP1 | dual specificity phosphatase 1 | 13.88 | 11.89 | -0.22 | 0.0 | 0.0 | 66.67 | Detail |
| 443 | CHOL | autosis | PPARD | peroxisome proliferator activated receptor delta | 9.0 | 10.06 | 0.16 | 0.0 | 0.0 | 66.67 | Detail |
| 444 | CHOL | autosis | ACTB | actin beta | 15.97 | 16.6 | 0.06 | 0.0 | 0.0 | 66.67 | Detail |
| 445 | CHOL | autosis | CASK | calcium/calmodulin dependent serine protein kinase | 9.43 | 10.46 | 0.15 | 0.0 | 0.0 | 66.67 | Detail |
| 446 | CHOL | autosis | VCL | vinculin | 9.89 | 11.16 | 0.17 | 0.0 | 0.0 | 66.67 | Detail |
| 447 | CHOL | autosis | CAV2 | caveolin 2 | 10.62 | 11.53 | 0.12 | 0.0 | 0.0 | 66.67 | Detail |
| 448 | CHOL | autosis | HOOK2 | hook microtubule tethering protein 2 | 9.18 | 10.34 | 0.17 | 0.0 | 0.0 | 66.67 | Detail |
| 449 | CHOL | autosis | RBM38 | RNA binding motif protein 38 | 8.61 | 9.61 | 0.16 | 0.0 | 0.0 | 66.67 | Detail |
| 450 | CHOL | autosis | LBH | LBH regulator of WNT signaling pathway | 8.99 | 10.91 | 0.28 | 0.0 | 0.0 | 66.67 | Detail |
| 451 | CHOL | autosis | FOXJ3 | forkhead box J3 | 9.37 | 10.03 | 0.1 | 0.0 | 0.0 | 66.67 | Detail |
| 452 | CHOL | autosis | CSNK1G2 | casein kinase 1 gamma 2 | 10.02 | 10.85 | 0.11 | 0.0 | 0.0 | 66.67 | Detail |
| 453 | CHOL | autosis | RYBP | RING1 and YY1 binding protein | 10.04 | 9.11 | -0.14 | 0.0 | 0.0 | 66.67 | Detail |
| 454 | CHOL | autosis | SIK3 | SIK family kinase 3 | 10.02 | 8.95 | -0.16 | 0.0 | 0.0 | 66.67 | Detail |
| 455 | CHOL | autosis | NR2F6 | nuclear receptor subfamily 2 group F member 6 | 11.32 | 10.43 | -0.12 | 0.0 | 0.0 | 66.67 | Detail |
| 456 | CHOL | autosis | GALNT10 | polypeptide N-acetylgalactosaminyltransferase 10 | 7.58 | 9.61 | 0.34 | 0.0 | 0.0 | 66.67 | Detail |
| 457 | CHOL | autosis | NME7 | NME/NM23 family member 7 | 6.98 | 8.16 | 0.23 | 0.0 | 0.0 | 66.67 | Detail |
| 458 | CHOL | autosis | ARPP19 | cAMP regulated phosphoprotein 19 | 10.84 | 11.56 | 0.09 | 0.0 | 0.0 | 66.67 | Detail |
| 459 | CHOL | autosis | GON4L | gon-4 like | 8.95 | 10.01 | 0.16 | 0.0 | 0.0 | 66.67 | Detail |
| 460 | CHOL | autosis | GPR107 | G protein-coupled receptor 107 | 9.98 | 11.01 | 0.14 | 0.0 | 0.0 | 66.67 | Detail |
| 461 | CHOL | autosis | PORCN | porcupine O-acyltransferase | 5.45 | 7.27 | 0.41 | 0.0 | 0.0 | 66.67 | Detail |
| 462 | CHOL | autosis | ZNF83 | zinc finger protein 83 | 7.91 | 9.84 | 0.31 | 0.0 | 0.0 | 66.67 | Detail |
| 463 | CHOL | autosis | ABTB2 | ankyrin repeat and BTB domain containing 2 | 9.23 | 7.65 | -0.27 | 0.0 | 0.0 | 66.67 | Detail |
| 464 | CHOL | autosis | ETFB | electron transfer flavoprotein subunit beta | 13.29 | 11.66 | -0.19 | 0.0 | 0.0 | 66.67 | Detail |
| 465 | CHOL | autosis | PCYOX1L | prenylcysteine oxidase 1 like | 5.64 | 7.54 | 0.42 | 0.0 | 0.0 | 66.67 | Detail |
| 466 | CHOL | autosis | HNF1B | HNF1 homeobox B | 8.98 | 11.43 | 0.35 | 0.0 | 0.0 | 62.5 | Detail |
| 467 | CHOL | autosis | HNRNPC | heterogeneous nuclear ribonucleoprotein C | 12.66 | 13.02 | 0.04 | 0.0 | 0.0 | 62.5 | Detail |
| 468 | CHOL | autosis | IVNS1ABP | influenza virus NS1A binding protein | 10.52 | 11.8 | 0.17 | 0.0 | 0.0 | 62.5 | Detail |
| 469 | CHOL | autosis | PLAU | plasminogen activator, urokinase | 6.26 | 8.71 | 0.48 | 0.0 | 0.0 | 62.5 | Detail |
| 470 | CHOL | autosis | RAB9A | RAB9A, member RAS oncogene family | 8.55 | 9.24 | 0.11 | 0.0 | 0.0 | 62.5 | Detail |
| 471 | CHOL | autosis | RELB | RELB proto-oncogene, NF-kB subunit | 8.21 | 10.0 | 0.29 | 0.0 | 0.0 | 62.5 | Detail |
| 472 | CHOL | autosis | ATP7A | ATPase copper transporting alpha | 6.75 | 7.95 | 0.24 | 0.0 | 0.0 | 62.5 | Detail |
| 473 | CHOL | autosis | MAP3K1 | mitogen-activated protein kinase kinase kinase 1 | 8.68 | 10.16 | 0.23 | 0.0 | 0.0 | 62.5 | Detail |
| 474 | CHOL | autosis | CDK2 | cyclin dependent kinase 2 | 8.68 | 9.51 | 0.13 | 0.0 | 0.0 | 62.5 | Detail |
| 475 | CHOL | autosis | GINS2 | GINS complex subunit 2 | 6.15 | 7.91 | 0.36 | 0.0 | 0.0 | 62.5 | Detail |
| 476 | CHOL | autosis | SMARCD3 | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 | 6.42 | 8.56 | 0.42 | 0.0 | 0.0 | 62.5 | Detail |
| 477 | CHOL | autosis | LONP1 | lon peptidase 1, mitochondrial | 11.82 | 11.2 | -0.08 | 0.0 | 0.0 | 62.5 | Detail |
| 478 | CHOL | autosis | CTDSPL | CTD small phosphatase like | 9.86 | 10.84 | 0.14 | 0.0 | 0.0 | 62.5 | Detail |
| 479 | CHOL | autosis | PTHLH | parathyroid hormone like hormone | 3.01 | 8.19 | 1.44 | 0.0 | 0.0 | 62.5 | Detail |
| 480 | CHOL | autosis | GPAA1 | glycosylphosphatidylinositol anchor attachment 1 | 11.15 | 11.75 | 0.08 | 0.0 | 0.0 | 62.5 | Detail |
| 481 | CHOL | autosis | SUB1 | SUB1 regulator of transcription | 11.17 | 12.16 | 0.12 | 0.0 | 0.0 | 62.5 | Detail |
| 482 | CHOL | autosis | GALNT2 | polypeptide N-acetylgalactosaminyltransferase 2 | 12.3 | 11.07 | -0.15 | 0.0 | 0.0 | 62.5 | Detail |
| 483 | CHOL | autosis | GALNT6 | polypeptide N-acetylgalactosaminyltransferase 6 | 5.2 | 7.77 | 0.58 | 0.0 | 0.0 | 62.5 | Detail |
| 484 | CHOL | autosis | ITGA2 | integrin subunit alpha 2 | 5.82 | 9.47 | 0.7 | 0.0 | 0.0 | 62.5 | Detail |
| 485 | CHOL | autosis | LIF | LIF interleukin 6 family cytokine | 6.17 | 9.89 | 0.68 | 0.0 | 0.0 | 62.5 | Detail |
| 486 | CHOL | autosis | SOCS6 | suppressor of cytokine signaling 6 | 10.43 | 9.26 | -0.17 | 0.0 | 0.0 | 62.5 | Detail |
| 487 | CHOL | autosis | PRDX4 | peroxiredoxin 4 | 11.9 | 10.99 | -0.11 | 0.0 | 0.0 | 62.5 | Detail |
| 488 | CHOL | autosis | HRH1 | histamine receptor H1 | 3.92 | 6.47 | 0.72 | 0.0 | 0.0 | 62.5 | Detail |
| 489 | CHOL | autosis | SLC6A4 | solute carrier family 6 member 4 | 0.54 | 2.69 | 2.31 | 0.0 | 0.01 | 62.5 | Detail |
| 490 | CHOL | autosis | M6PR | mannose-6-phosphate receptor, cation dependent | 10.96 | 11.53 | 0.07 | 0.0 | 0.0 | 62.5 | Detail |
| 491 | CHOL | autosis | ITGB8 | integrin subunit beta 8 | 6.23 | 9.72 | 0.64 | 0.0 | 0.0 | 62.5 | Detail |
| 492 | CHOL | autosis | ABHD5 | abhydrolase domain containing 5, lysophosphatidic acid acyltransferase | 9.37 | 8.38 | -0.16 | 0.0 | 0.0 | 62.5 | Detail |
| 493 | CHOL | autosis | SWAP70 | switching B cell complex subunit SWAP70 | 8.91 | 9.96 | 0.16 | 0.0 | 0.0 | 62.5 | Detail |
| 494 | CHOL | autosis | NEIL3 | nei like DNA glycosylase 3 | 1.13 | 4.11 | 1.87 | 0.0 | 0.0 | 62.5 | Detail |
| 495 | CHOL | autosis | SLC31A2 | solute carrier family 31 member 2 | 9.26 | 7.29 | -0.34 | 0.0 | 0.0 | 62.5 | Detail |
| 496 | CHOL | autosis | LSM2 | LSM2 homolog, U6 small nuclear RNA and mRNA degradation associated | 8.41 | 9.41 | 0.16 | 0.0 | 0.0 | 62.5 | Detail |
| 497 | CHOL | autosis | DUT | deoxyuridine triphosphatase | 9.34 | 10.08 | 0.11 | 0.0 | 0.0 | 62.5 | Detail |
| 498 | CHOL | autosis | WIPF2 | WAS/WASL interacting protein family member 2 | 8.87 | 9.86 | 0.15 | 0.0 | 0.0 | 62.5 | Detail |
| 499 | CHOL | autosis | UBTF | upstream binding transcription factor | 11.06 | 11.57 | 0.06 | 0.0 | 0.0 | 62.5 | Detail |
| 500 | CHOL | autosis | CCDC28B | coiled-coil domain containing 28B | 4.62 | 6.6 | 0.52 | 0.0 | 0.0 | 62.5 | Detail |
| 501 | CHOL | autosis | COX16 | cytochrome c oxidase assembly factor COX16 | 10.04 | 9.35 | -0.1 | 0.0 | 0.0 | 62.5 | Detail |
| 502 | CHOL | autosis | HINFP | histone H4 transcription factor | 7.7 | 8.46 | 0.14 | 0.0 | 0.0 | 62.5 | Detail |
| 503 | CHOL | autosis | LHFPL2 | LHFPL tetraspan subfamily member 2 | 7.36 | 9.07 | 0.3 | 0.0 | 0.0 | 62.5 | Detail |
| 504 | CHOL | autosis | WBP4 | WW domain binding protein 4 | 8.53 | 7.88 | -0.11 | 0.0 | 0.01 | 62.5 | Detail |
| 505 | CHOL | autosis | ZNF239 | zinc finger protein 239 | 4.4 | 6.67 | 0.6 | 0.0 | 0.0 | 62.5 | Detail |
| 506 | CHOL | autosis | TRMT2B | tRNA methyltransferase 2 homolog B | 7.9 | 8.44 | 0.1 | 0.0 | 0.0 | 62.5 | Detail |
| 507 | CHOL | autosis | ZNF16 | zinc finger protein 16 | 7.21 | 8.12 | 0.17 | 0.0 | 0.0 | 62.5 | Detail |
| 508 | CHOL | autosis | BCL2L1 | BCL2 like 1 | 11.39 | 13.08 | 0.2 | 0.0 | 0.0 | 58.33 | Detail |
| 509 | CHOL | autosis | NAMPT | nicotinamide phosphoribosyltransferase | 12.84 | 10.92 | -0.23 | 0.0 | 0.0 | 58.33 | Detail |
| 510 | CHOL | autosis | GMNN | geminin DNA replication inhibitor | 8.05 | 10.65 | 0.4 | 0.0 | 0.0 | 58.33 | Detail |
| 511 | CHOL | autosis | SFPQ | splicing factor proline and glutamine rich | 11.23 | 11.92 | 0.09 | 0.0 | 0.0 | 58.33 | Detail |
| 512 | CHOL | autosis | GABARAPL1 | GABA type A receptor associated protein like 1 | 12.28 | 10.89 | -0.17 | 0.0 | 0.0 | 58.33 | Detail |
| 513 | CHOL | autosis | MYLK | myosin light chain kinase | 11.82 | 10.52 | -0.17 | 0.0 | 0.0 | 58.33 | Detail |
| 514 | CHOL | autosis | IKBKB | inhibitor of nuclear factor kappa B kinase subunit beta | 9.38 | 10.04 | 0.1 | 0.0 | 0.0 | 58.33 | Detail |
| 515 | CHOL | autosis | RELN | reelin | 10.67 | 6.3 | -0.76 | 0.0 | 0.0 | 58.33 | Detail |
| 516 | CHOL | autosis | CDC25A | cell division cycle 25A | 4.32 | 5.59 | 0.37 | 0.0 | 0.0 | 58.33 | Detail |
| 517 | CHOL | autosis | ING1 | inhibitor of growth family member 1 | 8.98 | 8.09 | -0.15 | 0.0 | 0.0 | 58.33 | Detail |
| 518 | CHOL | autosis | PTP4A1 | protein tyrosine phosphatase 4A1 | 14.42 | 12.76 | -0.18 | 0.0 | 0.0 | 58.33 | Detail |
| 519 | CHOL | autosis | SYNJ2BP | synaptojanin 2 binding protein | 10.72 | 9.93 | -0.11 | 0.0 | 0.0 | 58.33 | Detail |
| 520 | CHOL | autosis | ATP1B1 | ATPase Na+/K+ transporting subunit beta 1 | 12.44 | 13.53 | 0.12 | 0.0 | 0.0 | 58.33 | Detail |
| 521 | CHOL | autosis | LMNB1 | lamin B1 | 7.71 | 9.31 | 0.27 | 0.0 | 0.0 | 58.33 | Detail |
| 522 | CHOL | autosis | TRIAP1 | TP53 regulated inhibitor of apoptosis 1 | 9.88 | 9.19 | -0.1 | 0.0 | 0.0 | 58.33 | Detail |
| 523 | CHOL | autosis | PPAN | peter pan homolog | 8.74 | 9.55 | 0.13 | 0.0 | 0.0 | 58.33 | Detail |
| 524 | CHOL | autosis | WDR7 | WD repeat domain 7 | 9.16 | 8.17 | -0.16 | 0.0 | 0.0 | 58.33 | Detail |
| 525 | CHOL | autosis | DMTF1 | cyclin D binding myb like transcription factor 1 | 8.76 | 9.71 | 0.15 | 0.0 | 0.0 | 58.33 | Detail |
| 526 | CHOL | autosis | PPM1F | protein phosphatase, Mg2+/Mn2+ dependent 1F | 8.77 | 9.77 | 0.16 | 0.0 | 0.0 | 58.33 | Detail |
| 527 | CHOL | autosis | DHX8 | DEAH-box helicase 8 | 9.21 | 9.87 | 0.1 | 0.0 | 0.0 | 58.33 | Detail |
| 528 | CHOL | autosis | CIR1 | corepressor interacting with RBPJ, CIR1 | 9.38 | 9.79 | 0.06 | 0.0 | 0.0 | 58.33 | Detail |
| 529 | CHOL | autosis | BSDC1 | BSD domain containing 1 | 11.87 | 11.13 | -0.09 | 0.0 | 0.0 | 58.33 | Detail |
| 530 | CHOL | autosis | INTS9 | integrator complex subunit 9 | 7.34 | 8.39 | 0.19 | 0.0 | 0.0 | 58.33 | Detail |
| 531 | CHOL | autosis | ISOC2 | isochorismatase domain containing 2 | 11.94 | 10.43 | -0.2 | 0.0 | 0.0 | 58.33 | Detail |
| 532 | CHOL | autosis | TJAP1 | tight junction associated protein 1 | 8.94 | 9.91 | 0.15 | 0.0 | 0.0 | 58.33 | Detail |
| 533 | CHOL | autosis | TMEM80 | transmembrane protein 80 | 8.08 | 8.83 | 0.13 | 0.0 | 0.0 | 58.33 | Detail |
| 534 | CHOL | autosis | AKAP8L | A-kinase anchoring protein 8 like | 9.46 | 10.33 | 0.13 | 0.0 | 0.0 | 58.33 | Detail |
| 535 | CHOL | autosis | POLR1A | RNA polymerase I subunit A | 8.01 | 8.91 | 0.15 | 0.0 | 0.0 | 58.33 | Detail |
| 536 | CHOL | autosis | TMCO6 | transmembrane and coiled-coil domains 6 | 8.97 | 8.08 | -0.15 | 0.0 | 0.0 | 58.33 | Detail |
| 537 | CHOL | autosis | USP36 | ubiquitin specific peptidase 36 | 9.55 | 10.34 | 0.11 | 0.0 | 0.0 | 58.33 | Detail |
| 538 | CHOL | autosis | PDE4DIP | phosphodiesterase 4D interacting protein | 11.4 | 10.23 | -0.16 | 0.0 | 0.01 | 54.17 | Detail |
| 539 | CHOL | autosis | SIRT1 | sirtuin 1 | 9.31 | 8.57 | -0.12 | 0.0 | 0.01 | 54.17 | Detail |
| 540 | CHOL | autosis | GPR161 | G protein-coupled receptor 161 | 3.82 | 5.59 | 0.55 | 0.0 | 0.01 | 54.17 | Detail |
| 541 | CHOL | autosis | MTMR3 | myotubularin related protein 3 | 9.29 | 10.07 | 0.12 | 0.0 | 0.01 | 54.17 | Detail |
| 542 | CHOL | autosis | RAD51C | RAD51 paralog C | 7.25 | 7.88 | 0.12 | 0.0 | 0.01 | 54.17 | Detail |
| 543 | CHOL | autosis | CARD10 | caspase recruitment domain family member 10 | 9.4 | 10.51 | 0.16 | 0.0 | 0.01 | 54.17 | Detail |
| 544 | CHOL | autosis | COL4A1 | collagen type IV alpha 1 chain | 10.76 | 12.99 | 0.27 | 0.0 | 0.01 | 54.17 | Detail |
| 545 | CHOL | autosis | RAB22A | RAB22A, member RAS oncogene family | 9.72 | 10.23 | 0.07 | 0.0 | 0.01 | 54.17 | Detail |
| 546 | CHOL | autosis | EIF3L | eukaryotic translation initiation factor 3 subunit L | 12.28 | 12.99 | 0.08 | 0.0 | 0.01 | 54.17 | Detail |
| 547 | CHOL | autosis | LDOC1 | LDOC1 regulator of NFKB signaling | 6.98 | 9.58 | 0.46 | 0.0 | 0.01 | 54.17 | Detail |
| 548 | CHOL | autosis | MIA2 | MIA SH3 domain ER export factor 2 | 8.78 | 6.35 | -0.47 | 0.0 | 0.01 | 54.17 | Detail |
| 549 | CHOL | autosis | HSPE1 | heat shock protein family E (Hsp10) member 1 | 11.85 | 11.15 | -0.09 | 0.0 | 0.01 | 54.17 | Detail |
| 550 | CHOL | autosis | ICAM3 | intercellular adhesion molecule 3 | 9.98 | 8.84 | -0.17 | 0.0 | 0.01 | 54.17 | Detail |
| 551 | CHOL | autosis | LARP6 | La ribonucleoprotein 6, translational regulator | 6.57 | 8.76 | 0.41 | 0.0 | 0.01 | 54.17 | Detail |
| 552 | CHOL | autosis | ACD | ACD shelterin complex subunit and telomerase recruitment factor | 7.77 | 8.94 | 0.2 | 0.0 | 0.01 | 54.17 | Detail |
| 553 | CHOL | autosis | RAD9A | RAD9 checkpoint clamp component A | 7.67 | 8.68 | 0.18 | 0.0 | 0.01 | 54.17 | Detail |
| 554 | CHOL | autosis | ATP2C1 | ATPase secretory pathway Ca2+ transporting 1 | 10.19 | 11.08 | 0.12 | 0.0 | 0.01 | 54.17 | Detail |
| 555 | CHOL | autosis | TXN2 | thioredoxin 2 | 11.55 | 10.87 | -0.09 | 0.0 | 0.01 | 54.17 | Detail |
| 556 | CHOL | autosis | RSL1D1 | ribosomal L1 domain containing 1 | 11.2 | 11.96 | 0.09 | 0.0 | 0.01 | 54.17 | Detail |
| 557 | CHOL | autosis | ARID3B | AT-rich interaction domain 3B | 6.23 | 7.31 | 0.23 | 0.0 | 0.01 | 54.17 | Detail |
| 558 | CHOL | autosis | ZNF354A | zinc finger protein 354A | 6.71 | 7.73 | 0.2 | 0.0 | 0.01 | 54.17 | Detail |
| 559 | CHOL | autosis | ZFP36L1 | ZFP36 ring finger protein like 1 | 13.34 | 12.27 | -0.12 | 0.0 | 0.01 | 54.17 | Detail |
| 560 | CHOL | autosis | ERMP1 | endoplasmic reticulum metallopeptidase 1 | 9.09 | 10.46 | 0.2 | 0.0 | 0.01 | 54.17 | Detail |
| 561 | CHOL | autosis | PTBP2 | polypyrimidine tract binding protein 2 | 7.45 | 8.44 | 0.18 | 0.0 | 0.01 | 54.17 | Detail |
| 562 | CHOL | autosis | TSC22D2 | TSC22 domain family member 2 | 9.29 | 8.28 | -0.17 | 0.0 | 0.01 | 54.17 | Detail |
| 563 | CHOL | autosis | BTN2A2 | butyrophilin subfamily 2 member A2 | 7.6 | 8.6 | 0.18 | 0.0 | 0.01 | 54.17 | Detail |
| 564 | CHOL | autosis | CNIH3 | cornichon family AMPA receptor auxiliary protein 3 | 2.62 | 4.42 | 0.75 | 0.0 | 0.01 | 54.17 | Detail |
| 565 | CHOL | autosis | FNTA | farnesyltransferase, CAAX box, subunit alpha | 9.94 | 10.5 | 0.08 | 0.0 | 0.01 | 52.08 | Detail |
| 566 | CHOL | autosis | NPC1 | NPC intracellular cholesterol transporter 1 | 8.93 | 10.18 | 0.19 | 0.0 | 0.01 | 50.0 | Detail |
| 567 | CHOL | autosis | PPARG | peroxisome proliferator activated receptor gamma | 8.39 | 7.16 | -0.23 | 0.0 | 0.01 | 50.0 | Detail |
| 568 | CHOL | autosis | SYK | spleen associated tyrosine kinase | 7.63 | 9.41 | 0.3 | 0.0 | 0.01 | 50.0 | Detail |
| 569 | CHOL | autosis | TP53 | tumor protein p53 | 9.47 | 10.68 | 0.17 | 0.0 | 0.01 | 50.0 | Detail |
| 570 | CHOL | autosis | KIN | Kin17 DNA and RNA binding protein | 7.52 | 8.05 | 0.1 | 0.0 | 0.01 | 50.0 | Detail |
| 571 | CHOL | autosis | FKBP4 | FKBP prolyl isomerase 4 | 11.28 | 11.92 | 0.08 | 0.0 | 0.01 | 50.0 | Detail |
| 572 | CHOL | autosis | LOXL2 | lysyl oxidase like 2 | 6.77 | 9.34 | 0.47 | 0.0 | 0.01 | 50.0 | Detail |
| 573 | CHOL | autosis | ENDOG | endonuclease G | 9.7 | 7.92 | -0.29 | 0.0 | 0.01 | 50.0 | Detail |
| 574 | CHOL | autosis | LOXL1 | lysyl oxidase like 1 | 6.17 | 8.41 | 0.45 | 0.0 | 0.01 | 50.0 | Detail |
| 575 | CHOL | autosis | FYN | FYN proto-oncogene, Src family tyrosine kinase | 9.75 | 8.66 | -0.17 | 0.0 | 0.01 | 50.0 | Detail |
| 576 | CHOL | autosis | GOLGA5 | golgin A5 | 10.63 | 9.89 | -0.1 | 0.0 | 0.01 | 50.0 | Detail |
| 577 | CHOL | autosis | RAC2 | Rac family small GTPase 2 | 8.61 | 10.4 | 0.27 | 0.0 | 0.01 | 50.0 | Detail |
| 578 | CHOL | autosis | VCAN | versican | 7.6 | 10.63 | 0.48 | 0.0 | 0.01 | 50.0 | Detail |
| 579 | CHOL | autosis | SPTLC2 | serine palmitoyltransferase long chain base subunit 2 | 8.48 | 9.54 | 0.17 | 0.0 | 0.01 | 50.0 | Detail |
| 580 | CHOL | autosis | KIF2A | kinesin family member 2A | 6.62 | 8.23 | 0.31 | 0.0 | 0.01 | 50.0 | Detail |
| 581 | CHOL | autosis | ZMIZ1 | zinc finger MIZ-type containing 1 | 10.12 | 11.15 | 0.14 | 0.0 | 0.01 | 50.0 | Detail |
| 582 | CHOL | autosis | P4HA2 | prolyl 4-hydroxylase subunit alpha 2 | 8.58 | 9.95 | 0.21 | 0.0 | 0.01 | 50.0 | Detail |
| 583 | CHOL | autosis | RFC2 | replication factor C subunit 2 | 9.06 | 9.73 | 0.1 | 0.0 | 0.01 | 50.0 | Detail |
| 584 | CHOL | autosis | IKZF5 | IKAROS family zinc finger 5 | 9.09 | 8.36 | -0.12 | 0.0 | 0.01 | 50.0 | Detail |
| 585 | CHOL | autosis | MTF1 | metal regulatory transcription factor 1 | 7.71 | 8.52 | 0.14 | 0.0 | 0.01 | 50.0 | Detail |
| 586 | CHOL | autosis | ERLIN2 | ER lipid raft associated 2 | 11.06 | 10.23 | -0.11 | 0.0 | 0.01 | 50.0 | Detail |
| 587 | CHOL | autosis | PKP4 | plakophilin 4 | 10.6 | 11.43 | 0.11 | 0.0 | 0.01 | 50.0 | Detail |
| 588 | CHOL | autosis | CAMTA2 | calmodulin binding transcription activator 2 | 9.6 | 10.3 | 0.1 | 0.0 | 0.01 | 50.0 | Detail |
| 589 | CHOL | autosis | VAT1 | vesicle amine transport 1 | 11.25 | 12.2 | 0.12 | 0.0 | 0.01 | 50.0 | Detail |
| 590 | CHOL | autosis | NUBPL | NUBP iron-sulfur cluster assembly factor, mitochondrial | 8.37 | 7.61 | -0.14 | 0.0 | 0.01 | 50.0 | Detail |
| 591 | CHOL | autosis | TOR3A | torsin family 3 member A | 9.5 | 10.53 | 0.15 | 0.0 | 0.01 | 50.0 | Detail |
| 592 | CHOL | autosis | ABHD14A | abhydrolase domain containing 14A | 8.76 | 7.7 | -0.19 | 0.0 | 0.01 | 50.0 | Detail |
| 593 | CHOL | autosis | MTMR1 | myotubularin related protein 1 | 8.61 | 9.44 | 0.13 | 0.0 | 0.01 | 50.0 | Detail |
| 594 | CHOL | autosis | NHP2 | NHP2 ribonucleoprotein | 9.88 | 10.57 | 0.1 | 0.0 | 0.01 | 50.0 | Detail |
| 595 | CHOL | autosis | SEC24D | SEC24 homolog D, COPII coat complex component | 10.98 | 9.63 | -0.19 | 0.0 | 0.01 | 50.0 | Detail |
| 596 | CHOL | autosis | SH3D19 | SH3 domain containing 19 | 11.21 | 10.3 | -0.12 | 0.0 | 0.01 | 50.0 | Detail |
| 597 | CHOL | autosis | UBL3 | ubiquitin like 3 | 10.98 | 9.75 | -0.17 | 0.0 | 0.01 | 50.0 | Detail |
| 598 | CHOL | autosis | ADAM17 | ADAM metallopeptidase domain 17 | 7.51 | 8.68 | 0.21 | 0.0 | 0.01 | 45.83 | Detail |
| 599 | CHOL | autosis | CSNK2A2 | casein kinase 2 alpha 2 | 8.51 | 9.05 | 0.09 | 0.0 | 0.01 | 45.83 | Detail |
| 600 | CHOL | autosis | IGF1R | insulin like growth factor 1 receptor | 6.75 | 9.11 | 0.43 | 0.0 | 0.01 | 45.83 | Detail |
| 601 | CHOL | autosis | VIM | vimentin | 12.4 | 14.31 | 0.21 | 0.0 | 0.01 | 45.83 | Detail |
| 602 | CHOL | autosis | PAX8 | paired box 8 | 4.86 | 7.33 | 0.59 | 0.0 | 0.01 | 45.83 | Detail |
| 603 | CHOL | autosis | NRAS | NRAS proto-oncogene, GTPase | 10.06 | 10.64 | 0.08 | 0.0 | 0.01 | 45.83 | Detail |
| 604 | CHOL | autosis | ATP7B | ATPase copper transporting beta | 9.91 | 7.94 | -0.32 | 0.0 | 0.01 | 45.83 | Detail |
| 605 | CHOL | autosis | RPA1 | replication protein A1 | 10.08 | 10.68 | 0.08 | 0.0 | 0.01 | 45.83 | Detail |
| 606 | CHOL | autosis | RAC1 | Rac family small GTPase 1 | 11.85 | 12.59 | 0.09 | 0.0 | 0.01 | 45.83 | Detail |
| 607 | CHOL | autosis | TXNIP | thioredoxin interacting protein | 13.43 | 11.86 | -0.18 | 0.0 | 0.01 | 45.83 | Detail |
| 608 | CHOL | autosis | CDK5 | cyclin dependent kinase 5 | 7.87 | 8.68 | 0.14 | 0.0 | 0.01 | 45.83 | Detail |
| 609 | CHOL | autosis | TPR | translocated promoter region, nuclear basket protein | 10.69 | 11.48 | 0.1 | 0.0 | 0.01 | 45.83 | Detail |
| 610 | CHOL | autosis | CITED2 | Cbp/p300 interacting transactivator with Glu/Asp rich carboxy-terminal domain 2 | 10.36 | 8.81 | -0.23 | 0.0 | 0.01 | 45.83 | Detail |
| 611 | CHOL | autosis | TOB1 | transducer of ERBB2, 1 | 12.07 | 10.75 | -0.17 | 0.0 | 0.01 | 45.83 | Detail |
| 612 | CHOL | autosis | PIK3R2 | phosphoinositide-3-kinase regulatory subunit 2 | 10.05 | 10.78 | 0.1 | 0.0 | 0.01 | 45.83 | Detail |
| 613 | CHOL | autosis | MYL9 | myosin light chain 9 | 10.3 | 11.86 | 0.2 | 0.0 | 0.01 | 45.83 | Detail |
| 614 | CHOL | autosis | WDFY3 | WD repeat and FYVE domain containing 3 | 9.16 | 10.15 | 0.15 | 0.0 | 0.01 | 45.83 | Detail |
| 615 | CHOL | autosis | DNAJA1 | DnaJ heat shock protein family (Hsp40) member A1 | 12.12 | 10.99 | -0.14 | 0.0 | 0.01 | 45.83 | Detail |
| 616 | CHOL | autosis | SLC38A6 | solute carrier family 38 member 6 | 6.17 | 6.85 | 0.15 | 0.0 | 0.01 | 45.83 | Detail |
| 617 | CHOL | autosis | PRR16 | proline rich 16 | 3.31 | 4.92 | 0.57 | 0.0 | 0.01 | 45.83 | Detail |
| 618 | CHOL | autosis | STX10 | syntaxin 10 | 8.07 | 8.92 | 0.14 | 0.0 | 0.01 | 45.83 | Detail |
| 619 | CHOL | autosis | ZCCHC14 | zinc finger CCHC-type containing 14 | 10.6 | 9.83 | -0.11 | 0.0 | 0.01 | 45.83 | Detail |
| 620 | CHOL | autosis | ZMYM3 | zinc finger MYM-type containing 3 | 9.13 | 9.7 | 0.09 | 0.0 | 0.01 | 45.83 | Detail |
| 621 | CHOL | autosis | APP | amyloid beta precursor protein | 13.73 | 14.42 | 0.07 | 0.0 | 0.01 | 41.67 | Detail |
| 622 | CHOL | autosis | GSTP1 | glutathione S-transferase pi 1 | 9.56 | 12.51 | 0.39 | 0.0 | 0.01 | 41.67 | Detail |
| 623 | CHOL | autosis | SPARC | secreted protein acidic and cysteine rich | 12.61 | 13.84 | 0.13 | 0.0 | 0.01 | 41.67 | Detail |
| 624 | CHOL | autosis | FANCA | FA complementation group A | 6.97 | 8.26 | 0.24 | 0.0 | 0.01 | 41.67 | Detail |
| 625 | CHOL | autosis | SMURF2 | SMAD specific E3 ubiquitin protein ligase 2 | 7.0 | 8.39 | 0.26 | 0.0 | 0.01 | 41.67 | Detail |
| 626 | CHOL | autosis | GTF2IRD1 | GTF2I repeat domain containing 1 | 7.22 | 8.85 | 0.29 | 0.0 | 0.01 | 41.67 | Detail |
| 627 | CHOL | autosis | HNRNPA2B1 | heterogeneous nuclear ribonucleoprotein A2/B1 | 13.44 | 13.75 | 0.03 | 0.0 | 0.01 | 41.67 | Detail |
| 628 | CHOL | autosis | KDM3A | lysine demethylase 3A | 9.17 | 10.0 | 0.12 | 0.0 | 0.01 | 41.67 | Detail |
| 629 | CHOL | autosis | RALGDS | ral guanine nucleotide dissociation stimulator | 9.06 | 10.24 | 0.18 | 0.0 | 0.01 | 41.67 | Detail |
| 630 | CHOL | autosis | ATP8A2 | ATPase phospholipid transporting 8A2 | 1.18 | 4.2 | 1.83 | 0.0 | 0.01 | 41.67 | Detail |
| 631 | CHOL | autosis | ZNF84 | zinc finger protein 84 | 8.27 | 9.3 | 0.17 | 0.0 | 0.01 | 41.67 | Detail |
| 632 | CHOL | autosis | HIC2 | HIC ZBTB transcriptional repressor 2 | 6.33 | 7.16 | 0.18 | 0.0 | 0.01 | 41.67 | Detail |
| 633 | CHOL | autosis | PRIM1 | DNA primase subunit 1 | 5.77 | 6.65 | 0.21 | 0.0 | 0.01 | 41.67 | Detail |
| 634 | CHOL | autosis | DBF4 | DBF4-CDC7 kinase regulatory subunit | 6.23 | 7.09 | 0.19 | 0.0 | 0.01 | 41.67 | Detail |
| 635 | CHOL | autosis | AP3B2 | adaptor related protein complex 3 subunit beta 2 | 1.22 | 5.01 | 2.03 | 0.0 | 0.01 | 41.67 | Detail |
| 636 | CHOL | autosis | C1orf112 | FIGNL1 interacting regulator of recombination and mitosis | 5.83 | 7.29 | 0.32 | 0.0 | 0.01 | 41.67 | Detail |
| 637 | CHOL | autosis | LUC7L | LUC7 like | 8.83 | 9.95 | 0.17 | 0.0 | 0.01 | 41.67 | Detail |
| 638 | CHOL | autosis | ADCY7 | adenylate cyclase 7 | 7.47 | 8.91 | 0.25 | 0.0 | 0.01 | 41.67 | Detail |
| 639 | CHOL | autosis | PPIC | peptidylprolyl isomerase C | 9.7 | 10.32 | 0.09 | 0.0 | 0.01 | 41.67 | Detail |
| 640 | CHOL | autosis | CXCL2 | C-X-C motif chemokine ligand 2 | 11.22 | 9.13 | -0.3 | 0.0 | 0.01 | 41.67 | Detail |
| 641 | CHOL | autosis | FAS | Fas cell surface death receptor | 9.23 | 7.95 | -0.22 | 0.0 | 0.01 | 41.67 | Detail |
| 642 | CHOL | autosis | FASN | fatty acid synthase | 13.75 | 11.74 | -0.23 | 0.0 | 0.01 | 41.67 | Detail |
| 643 | CHOL | autosis | JUN | Jun proto-oncogene, AP-1 transcription factor subunit | 12.82 | 11.72 | -0.13 | 0.0 | 0.01 | 41.67 | Detail |
| 644 | CHOL | autosis | SERPINE1 | serpin family E member 1 | 13.76 | 11.39 | -0.27 | 0.0 | 0.01 | 41.67 | Detail |
| 645 | CHOL | autosis | SMAD1 | SMAD family member 1 | 9.39 | 8.57 | -0.13 | 0.0 | 0.01 | 41.67 | Detail |
| 646 | CHOL | autosis | ATP1B2 | ATPase Na+/K+ transporting subunit beta 2 | 6.87 | 5.29 | -0.38 | 0.0 | 0.01 | 41.67 | Detail |
| 647 | CHOL | autosis | TTC17 | tetratricopeptide repeat domain 17 | 10.46 | 9.78 | -0.1 | 0.0 | 0.01 | 41.67 | Detail |
| 648 | CHOL | autosis | TMEM14A | transmembrane protein 14A | 9.75 | 8.95 | -0.12 | 0.0 | 0.01 | 41.67 | Detail |
| 649 | CHOL | autosis | DUSP10 | dual specificity phosphatase 10 | 10.12 | 8.95 | -0.18 | 0.0 | 0.01 | 41.67 | Detail |
| 650 | CHOL | autosis | EHD3 | EH domain containing 3 | 8.94 | 7.49 | -0.26 | 0.0 | 0.01 | 41.67 | Detail |
| 651 | CHOL | autosis | SOAT2 | sterol O-acyltransferase 2 | 5.61 | 2.38 | -1.24 | 0.0 | 0.01 | 41.67 | Detail |
| 652 | CHOL | autosis | DPM3 | dolichyl-phosphate mannosyltransferase subunit 3, regulatory | 9.33 | 8.32 | -0.17 | 0.0 | 0.01 | 41.67 | Detail |
| 653 | CHOL | autosis | RSAD1 | radical S-adenosyl methionine domain containing 1 | 10.54 | 9.88 | -0.09 | 0.0 | 0.01 | 41.67 | Detail |
| 654 | CHOL | autosis | ATF4 | activating transcription factor 4 | 12.47 | 13.05 | 0.07 | 0.0 | 0.01 | 37.5 | Detail |
| 655 | CHOL | autosis | PIGF | phosphatidylinositol glycan anchor biosynthesis class F | 7.53 | 7.9 | 0.07 | 0.0 | 0.01 | 37.5 | Detail |
| 656 | CHOL | autosis | HSF1 | heat shock transcription factor 1 | 10.43 | 11.0 | 0.08 | 0.0 | 0.01 | 37.5 | Detail |
| 657 | CHOL | autosis | CCND2 | cyclin D2 | 7.15 | 9.34 | 0.38 | 0.0 | 0.01 | 37.5 | Detail |
| 658 | CHOL | autosis | ADAM12 | ADAM metallopeptidase domain 12 | 4.54 | 6.72 | 0.57 | 0.0 | 0.01 | 37.5 | Detail |
| 659 | CHOL | autosis | CDH11 | cadherin 11 | 5.81 | 8.48 | 0.54 | 0.0 | 0.01 | 37.5 | Detail |
| 660 | CHOL | autosis | KCNH4 | potassium voltage-gated channel subfamily H member 4 | 0.87 | 2.93 | 1.75 | 0.01 | 0.02 | 37.5 | Detail |
| 661 | CHOL | autosis | PLAGL2 | PLAG1 like zinc finger 2 | 7.87 | 8.73 | 0.15 | 0.0 | 0.01 | 37.5 | Detail |
| 662 | CHOL | autosis | SOX7 | SRY-box transcription factor 7 | 7.89 | 6.21 | -0.35 | 0.0 | 0.01 | 37.5 | Detail |
| 663 | CHOL | autosis | CLTB | clathrin light chain B | 10.38 | 11.24 | 0.11 | 0.0 | 0.01 | 37.5 | Detail |
| 664 | CHOL | autosis | KLF10 | KLF transcription factor 10 | 11.44 | 10.07 | -0.18 | 0.0 | 0.01 | 37.5 | Detail |
| 665 | CHOL | autosis | ZYX | zyxin | 11.11 | 11.9 | 0.1 | 0.0 | 0.01 | 37.5 | Detail |
| 666 | CHOL | autosis | TOPORS | TOP1 binding arginine/serine rich protein, E3 ubiquitin ligase | 9.2 | 8.5 | -0.11 | 0.0 | 0.01 | 37.5 | Detail |
| 667 | CHOL | autosis | AHNAK2 | AHNAK nucleoprotein 2 | 5.61 | 9.34 | 0.74 | 0.0 | 0.01 | 37.5 | Detail |
| 668 | CHOL | autosis | DNA2 | DNA replication helicase/nuclease 2 | 5.57 | 6.85 | 0.3 | 0.0 | 0.01 | 37.5 | Detail |
| 669 | CHOL | autosis | SNX6 | sorting nexin 6 | 9.4 | 9.93 | 0.08 | 0.0 | 0.01 | 37.5 | Detail |
| 670 | CHOL | autosis | THAP11 | THAP domain containing 11 | 8.79 | 9.41 | 0.1 | 0.0 | 0.01 | 37.5 | Detail |
| 671 | CHOL | autosis | TASP1 | taspase 1 | 6.96 | 7.8 | 0.16 | 0.0 | 0.01 | 37.5 | Detail |
| 672 | CHOL | autosis | POLR3K | RNA polymerase III subunit K | 7.43 | 8.15 | 0.13 | 0.0 | 0.01 | 37.5 | Detail |
| 673 | CHOL | autosis | PDSS1 | decaprenyl diphosphate synthase subunit 1 | 6.13 | 7.03 | 0.2 | 0.0 | 0.01 | 37.5 | Detail |
| 674 | CHOL | autosis | SECISBP2L | SECIS binding protein 2 like | 10.11 | 9.49 | -0.09 | 0.01 | 0.02 | 37.5 | Detail |
| 675 | CHOL | autosis | SLC12A7 | solute carrier family 12 member 7 | 11.22 | 11.94 | 0.09 | 0.0 | 0.01 | 37.5 | Detail |
| 676 | CHOL | autosis | GPR3 | G protein-coupled receptor 3 | 2.6 | 4.25 | 0.71 | 0.0 | 0.01 | 37.5 | Detail |
| 677 | CHOL | autosis | CEPT1 | choline/ethanolamine phosphotransferase 1 | 9.99 | 9.4 | -0.09 | 0.0 | 0.01 | 37.5 | Detail |
| 678 | CHOL | autosis | VPS37B | VPS37B subunit of ESCRT-I | 8.74 | 9.9 | 0.18 | 0.0 | 0.01 | 37.5 | Detail |
| 679 | CHOL | autosis | ITGA5 | integrin subunit alpha 5 | 11.05 | 12.21 | 0.14 | 0.01 | 0.01 | 33.33 | Detail |
| 680 | CHOL | autosis | JUND | JunD proto-oncogene, AP-1 transcription factor subunit | 12.8 | 11.73 | -0.13 | 0.01 | 0.01 | 33.33 | Detail |
| 681 | CHOL | autosis | MARCKSL1 | MARCKS like 1 | 9.74 | 11.61 | 0.25 | 0.01 | 0.01 | 33.33 | Detail |
| 682 | CHOL | autosis | MMP1 | matrix metallopeptidase 1 | 1.8 | 5.42 | 1.59 | 0.01 | 0.02 | 33.33 | Detail |
| 683 | CHOL | autosis | OSTF1 | osteoclast stimulating factor 1 | 9.73 | 8.95 | -0.12 | 0.01 | 0.01 | 33.33 | Detail |
| 684 | CHOL | autosis | BRCC3 | BRCA1/BRCA2-containing complex subunit 3 | 8.29 | 8.82 | 0.09 | 0.01 | 0.01 | 33.33 | Detail |
| 685 | CHOL | autosis | KLF5 | KLF transcription factor 5 | 6.93 | 9.73 | 0.49 | 0.01 | 0.01 | 33.33 | Detail |
| 686 | CHOL | autosis | SLFN12 | schlafen family member 12 | 4.95 | 6.6 | 0.42 | 0.01 | 0.01 | 33.33 | Detail |
| 687 | CHOL | autosis | ACVR1 | activin A receptor type 1 | 9.5 | 10.34 | 0.12 | 0.01 | 0.01 | 33.33 | Detail |
| 688 | CHOL | autosis | LRP6 | LDL receptor related protein 6 | 10.64 | 9.45 | -0.17 | 0.01 | 0.01 | 33.33 | Detail |
| 689 | CHOL | autosis | TRIM13 | tripartite motif containing 13 | 8.46 | 9.14 | 0.11 | 0.01 | 0.01 | 33.33 | Detail |
| 690 | CHOL | autosis | FLOT1 | flotillin 1 | 11.46 | 12.28 | 0.1 | 0.01 | 0.01 | 33.33 | Detail |
| 691 | CHOL | autosis | CDKN2C | cyclin dependent kinase inhibitor 2C | 6.38 | 7.82 | 0.29 | 0.01 | 0.01 | 33.33 | Detail |
| 692 | CHOL | autosis | PER2 | period circadian regulator 2 | 9.67 | 8.67 | -0.16 | 0.01 | 0.01 | 33.33 | Detail |
| 693 | CHOL | autosis | KLF11 | KLF transcription factor 11 | 10.59 | 9.12 | -0.21 | 0.01 | 0.01 | 33.33 | Detail |
| 694 | CHOL | autosis | RPL10 | ribosomal protein L10 | 13.41 | 14.23 | 0.09 | 0.01 | 0.01 | 33.33 | Detail |
| 695 | CHOL | autosis | ARAP2 | ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 2 | 8.28 | 9.33 | 0.17 | 0.01 | 0.01 | 33.33 | Detail |
| 696 | CHOL | autosis | BRF2 | BRF2 RNA polymerase III transcription initiation factor subunit | 7.74 | 8.39 | 0.12 | 0.01 | 0.01 | 33.33 | Detail |
| 697 | CHOL | autosis | PER1 | period circadian regulator 1 | 12.31 | 10.89 | -0.18 | 0.01 | 0.01 | 33.33 | Detail |
| 698 | CHOL | autosis | SP2 | Sp2 transcription factor | 8.71 | 9.15 | 0.07 | 0.01 | 0.01 | 33.33 | Detail |
| 699 | CHOL | autosis | ARID4B | AT-rich interaction domain 4B | 9.31 | 10.06 | 0.11 | 0.01 | 0.01 | 33.33 | Detail |
| 700 | CHOL | autosis | CBLB | Cbl proto-oncogene B | 7.81 | 8.81 | 0.17 | 0.01 | 0.01 | 33.33 | Detail |
| 701 | CHOL | autosis | CNKSR1 | connector enhancer of kinase suppressor of Ras 1 | 6.18 | 8.18 | 0.4 | 0.01 | 0.01 | 33.33 | Detail |
| 702 | CHOL | autosis | STAM | signal transducing adaptor molecule | 8.02 | 8.81 | 0.14 | 0.01 | 0.01 | 33.33 | Detail |
| 703 | CHOL | autosis | ATP6V1C1 | ATPase H+ transporting V1 subunit C1 | 10.0 | 10.67 | 0.09 | 0.01 | 0.01 | 33.33 | Detail |
| 704 | CHOL | autosis | MSRB2 | methionine sulfoxide reductase B2 | 10.39 | 9.42 | -0.14 | 0.01 | 0.01 | 33.33 | Detail |
| 705 | CHOL | autosis | ARHGEF3 | Rho guanine nucleotide exchange factor 3 | 7.32 | 8.71 | 0.25 | 0.01 | 0.01 | 33.33 | Detail |
| 706 | CHOL | autosis | ATP2A1 | ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 | 2.64 | 5.15 | 0.96 | 0.01 | 0.01 | 33.33 | Detail |
| 707 | CHOL | autosis | ZBTB40 | zinc finger and BTB domain containing 40 | 7.99 | 8.82 | 0.14 | 0.01 | 0.01 | 33.33 | Detail |
| 708 | CHOL | autosis | ATP1A3 | ATPase Na+/K+ transporting subunit alpha 3 | 1.99 | 4.97 | 1.32 | 0.01 | 0.02 | 29.17 | Detail |
| 709 | CHOL | autosis | DNMT3B | DNA methyltransferase 3 beta | 4.94 | 6.42 | 0.38 | 0.01 | 0.02 | 29.17 | Detail |
| 710 | CHOL | autosis | NEU1 | neuraminidase 1 | 10.4 | 10.96 | 0.08 | 0.01 | 0.02 | 29.17 | Detail |
| 711 | CHOL | autosis | PPP1CA | protein phosphatase 1 catalytic subunit alpha | 11.58 | 12.14 | 0.07 | 0.01 | 0.02 | 29.17 | Detail |
| 712 | CHOL | autosis | SAT1 | spermidine/spermine N1-acetyltransferase 1 | 13.43 | 12.45 | -0.11 | 0.01 | 0.02 | 29.17 | Detail |
| 713 | CHOL | autosis | BARD1 | BRCA1 associated RING domain 1 | 3.12 | 5.14 | 0.72 | 0.01 | 0.02 | 29.17 | Detail |
| 714 | CHOL | autosis | PPL | periplakin | 10.22 | 10.99 | 0.1 | 0.01 | 0.02 | 29.17 | Detail |
| 715 | CHOL | autosis | EGR1 | early growth response 1 | 13.41 | 11.38 | -0.24 | 0.01 | 0.02 | 29.17 | Detail |
| 716 | CHOL | autosis | MYCBP | MYC binding protein | 8.28 | 9.11 | 0.14 | 0.01 | 0.02 | 29.17 | Detail |
| 717 | CHOL | autosis | CELF1 | CUGBP Elav-like family member 1 | 11.25 | 10.69 | -0.07 | 0.01 | 0.02 | 29.17 | Detail |
| 718 | CHOL | autosis | MAP3K5 | mitogen-activated protein kinase kinase kinase 5 | 9.0 | 7.44 | -0.27 | 0.01 | 0.02 | 29.17 | Detail |
| 719 | CHOL | autosis | MAP3K9 | mitogen-activated protein kinase kinase kinase 9 | 4.33 | 5.79 | 0.42 | 0.01 | 0.02 | 29.17 | Detail |
| 720 | CHOL | autosis | LRIG2 | leucine rich repeats and immunoglobulin like domains 2 | 6.07 | 7.07 | 0.22 | 0.01 | 0.02 | 29.17 | Detail |
| 721 | CHOL | autosis | MLLT11 | MLLT11 transcription factor 7 cofactor | 5.68 | 7.14 | 0.33 | 0.01 | 0.02 | 29.17 | Detail |
| 722 | CHOL | autosis | GNB2 | G protein subunit beta 2 | 11.63 | 12.2 | 0.07 | 0.01 | 0.02 | 29.17 | Detail |
| 723 | CHOL | autosis | HAUS6 | HAUS augmin like complex subunit 6 | 7.72 | 8.26 | 0.1 | 0.01 | 0.02 | 29.17 | Detail |
| 724 | CHOL | autosis | CTNNBIP1 | catenin beta interacting protein 1 | 9.26 | 8.63 | -0.1 | 0.01 | 0.02 | 29.17 | Detail |
| 725 | CHOL | autosis | MAP4K5 | mitogen-activated protein kinase kinase kinase kinase 5 | 8.7 | 9.52 | 0.13 | 0.01 | 0.02 | 29.17 | Detail |
| 726 | CHOL | autosis | RPL13A | ribosomal protein L13a | 13.34 | 14.36 | 0.11 | 0.01 | 0.02 | 29.17 | Detail |
| 727 | CHOL | autosis | AZI2 | 5-azacytidine induced 2 | 9.75 | 9.02 | -0.11 | 0.01 | 0.02 | 29.17 | Detail |
| 728 | CHOL | autosis | PSG9 | pregnancy specific beta-1-glycoprotein 9 | 0.21 | 2.29 | 3.45 | 0.01 | 0.02 | 29.17 | Detail |
| 729 | CHOL | autosis | MRPL40 | mitochondrial ribosomal protein L40 | 10.34 | 9.55 | -0.11 | 0.01 | 0.02 | 29.17 | Detail |
| 730 | CHOL | autosis | THAP10 | THAP domain containing 10 | 5.19 | 6.02 | 0.21 | 0.01 | 0.02 | 29.17 | Detail |
| 731 | CHOL | autosis | AK1 | adenylate kinase 1 | 9.93 | 8.81 | -0.17 | 0.01 | 0.02 | 29.17 | Detail |
| 732 | CHOL | autosis | ARMCX2 | armadillo repeat containing X-linked 2 | 6.75 | 8.75 | 0.37 | 0.01 | 0.02 | 29.17 | Detail |
| 733 | CHOL | autosis | MDFIC | MyoD family inhibitor domain containing | 9.13 | 10.02 | 0.13 | 0.01 | 0.02 | 29.17 | Detail |
| 734 | CHOL | autosis | NDEL1 | nudE neurodevelopment protein 1 like 1 | 10.07 | 9.43 | -0.1 | 0.01 | 0.02 | 29.17 | Detail |
| 735 | CHOL | autosis | RNF103 | ring finger protein 103 | 11.32 | 10.69 | -0.08 | 0.01 | 0.02 | 29.17 | Detail |
| 736 | CHOL | autosis | BMP2 | bone morphogenetic protein 2 | 7.99 | 9.33 | 0.22 | 0.01 | 0.02 | 25.0 | Detail |
| 737 | CHOL | autosis | CXCL3 | C-X-C motif chemokine ligand 3 | 2.55 | 5.21 | 1.03 | 0.01 | 0.02 | 25.0 | Detail |
| 738 | CHOL | autosis | DDIT3 | DNA damage inducible transcript 3 | 8.52 | 9.41 | 0.14 | 0.01 | 0.02 | 25.0 | Detail |
| 739 | CHOL | autosis | IL15 | interleukin 15 | 6.01 | 6.96 | 0.21 | 0.01 | 0.02 | 25.0 | Detail |
| 740 | CHOL | autosis | IL1A | interleukin 1 alpha | 0.38 | 2.55 | 2.74 | 0.01 | 0.03 | 25.0 | Detail |
| 741 | CHOL | autosis | RIPK1 | receptor interacting serine/threonine kinase 1 | 10.23 | 9.87 | -0.05 | 0.01 | 0.02 | 25.0 | Detail |
| 742 | CHOL | autosis | SIK1 | salt inducible kinase 1 | 12.08 | 10.67 | -0.18 | 0.01 | 0.02 | 25.0 | Detail |
| 743 | CHOL | autosis | CCND3 | cyclin D3 | 9.16 | 10.24 | 0.16 | 0.01 | 0.02 | 25.0 | Detail |
| 744 | CHOL | autosis | MLF1 | myeloid leukemia factor 1 | 4.78 | 7.0 | 0.55 | 0.01 | 0.02 | 25.0 | Detail |
| 745 | CHOL | autosis | CTCF | CCCTC-binding factor | 9.84 | 10.34 | 0.07 | 0.01 | 0.02 | 25.0 | Detail |
| 746 | CHOL | autosis | KDM4B | lysine demethylase 4B | 10.06 | 9.66 | -0.06 | 0.01 | 0.02 | 25.0 | Detail |
| 747 | CHOL | autosis | LTA | lymphotoxin alpha | 1.87 | 3.7 | 0.99 | 0.01 | 0.02 | 25.0 | Detail |
| 748 | CHOL | autosis | PAIP1 | poly(A) binding protein interacting protein 1 | 9.89 | 10.22 | 0.05 | 0.01 | 0.02 | 25.0 | Detail |
| 749 | CHOL | autosis | XPA | XPA, DNA damage recognition and repair factor | 8.6 | 7.87 | -0.13 | 0.01 | 0.02 | 25.0 | Detail |
| 750 | CHOL | autosis | MEIS2 | Meis homeobox 2 | 8.05 | 9.57 | 0.25 | 0.01 | 0.02 | 25.0 | Detail |
| 751 | CHOL | autosis | TUBB | tubulin beta class I | 13.27 | 14.04 | 0.08 | 0.01 | 0.02 | 25.0 | Detail |
| 752 | CHOL | autosis | RAD54B | RAD54 homolog B | 5.14 | 6.34 | 0.3 | 0.01 | 0.02 | 25.0 | Detail |
| 753 | CHOL | autosis | SLC37A4 | solute carrier family 37 member 4 | 12.77 | 11.25 | -0.18 | 0.01 | 0.02 | 25.0 | Detail |
| 754 | CHOL | autosis | CBR3 | carbonyl reductase 3 | 3.68 | 5.23 | 0.51 | 0.01 | 0.02 | 25.0 | Detail |
| 755 | CHOL | autosis | PRRG4 | proline rich and Gla domain 4 | 8.94 | 7.55 | -0.24 | 0.01 | 0.02 | 25.0 | Detail |
| 756 | CHOL | autosis | MKLN1 | muskelin 1 | 10.71 | 10.15 | -0.08 | 0.01 | 0.02 | 25.0 | Detail |
| 757 | CHOL | autosis | ATP6V0E2 | ATPase H+ transporting V0 subunit e2 | 11.15 | 10.14 | -0.14 | 0.01 | 0.02 | 25.0 | Detail |
| 758 | CHOL | autosis | RHOBTB3 | Rho related BTB domain containing 3 | 10.59 | 8.87 | -0.25 | 0.01 | 0.02 | 25.0 | Detail |
| 759 | CHOL | autosis | TIMM8B | translocase of inner mitochondrial membrane 8 homolog B | 9.49 | 8.79 | -0.11 | 0.01 | 0.02 | 25.0 | Detail |
| 760 | CHOL | autosis | YLPM1 | YLP motif containing 1 | 9.63 | 10.12 | 0.07 | 0.01 | 0.02 | 25.0 | Detail |
| 761 | CHOL | autosis | BRAP | BRCA1 associated protein | 8.63 | 9.05 | 0.07 | 0.01 | 0.02 | 20.83 | Detail |
| 762 | CHOL | autosis | CD47 | CD47 molecule | 10.1 | 10.97 | 0.12 | 0.01 | 0.02 | 20.83 | Detail |
| 763 | CHOL | autosis | COMT | catechol-O-methyltransferase | 12.64 | 11.84 | -0.09 | 0.01 | 0.02 | 20.83 | Detail |
| 764 | CHOL | autosis | MAT2A | methionine adenosyltransferase 2A | 11.52 | 12.16 | 0.08 | 0.01 | 0.02 | 20.83 | Detail |
| 765 | CHOL | autosis | PMP22 | peripheral myelin protein 22 | 8.0 | 9.24 | 0.21 | 0.01 | 0.02 | 20.83 | Detail |
| 766 | CHOL | autosis | PSPH | phosphoserine phosphatase | 7.62 | 8.77 | 0.2 | 0.01 | 0.02 | 20.83 | Detail |
| 767 | CHOL | autosis | TNFRSF1A | TNF receptor superfamily member 1A | 12.19 | 11.53 | -0.08 | 0.01 | 0.02 | 20.83 | Detail |
| 768 | CHOL | autosis | RSRC2 | arginine and serine rich coiled-coil 2 | 9.98 | 10.43 | 0.06 | 0.01 | 0.02 | 20.83 | Detail |
| 769 | CHOL | autosis | ROR1 | receptor tyrosine kinase like orphan receptor 1 | 2.86 | 5.13 | 0.84 | 0.01 | 0.02 | 20.83 | Detail |
| 770 | CHOL | autosis | OAZ1 | ornithine decarboxylase antizyme 1 | 13.01 | 13.48 | 0.05 | 0.01 | 0.02 | 20.83 | Detail |
| 771 | CHOL | autosis | QSOX1 | quiescin sulfhydryl oxidase 1 | 10.9 | 12.42 | 0.19 | 0.01 | 0.02 | 20.83 | Detail |
| 772 | CHOL | autosis | AGPAT1 | 1-acylglycerol-3-phosphate O-acyltransferase 1 | 10.61 | 11.24 | 0.08 | 0.01 | 0.02 | 20.83 | Detail |
| 773 | CHOL | autosis | MAP2K3 | mitogen-activated protein kinase kinase 3 | 11.44 | 10.79 | -0.08 | 0.01 | 0.02 | 20.83 | Detail |
| 774 | CHOL | autosis | LRRC8E | leucine rich repeat containing 8 VRAC subunit E | 7.66 | 8.32 | 0.12 | 0.01 | 0.02 | 20.83 | Detail |
| 775 | CHOL | autosis | FAM86B1 | family with sequence similarity 86 member B1 | 5.1 | 6.34 | 0.31 | 0.01 | 0.02 | 20.83 | Detail |
| 776 | CHOL | autosis | ZNF473 | zinc finger protein 473 | 7.18 | 7.87 | 0.13 | 0.01 | 0.02 | 20.83 | Detail |
| 777 | CHOL | autosis | MPHOSPH9 | M-phase phosphoprotein 9 | 7.74 | 6.88 | -0.17 | 0.01 | 0.02 | 20.83 | Detail |
| 778 | CHOL | autosis | PWP2 | PWP2 small subunit processome component | 8.86 | 9.41 | 0.09 | 0.01 | 0.02 | 20.83 | Detail |
| 779 | CHOL | autosis | SLC43A3 | solute carrier family 43 member 3 | 11.6 | 10.47 | -0.15 | 0.01 | 0.02 | 20.83 | Detail |
| 780 | CHOL | autosis | SNN | stannin | 8.34 | 9.26 | 0.15 | 0.01 | 0.02 | 20.83 | Detail |
| 781 | CHOL | autosis | CSF2 | colony stimulating factor 2 | 0.0 | 1.34 | inf | 0.01 | 0.03 | 16.67 | Detail |
| 782 | CHOL | autosis | PLEKHM1 | pleckstrin homology and RUN domain containing M1 | 8.87 | 9.5 | 0.1 | 0.01 | 0.03 | 16.67 | Detail |
| 783 | CHOL | autosis | USP18 | ubiquitin specific peptidase 18 | 8.98 | 7.79 | -0.2 | 0.01 | 0.03 | 16.67 | Detail |
| 784 | CHOL | autosis | HDAC1 | histone deacetylase 1 | 10.45 | 11.24 | 0.11 | 0.01 | 0.03 | 16.67 | Detail |
| 785 | CHOL | autosis | SYP | synaptophysin | 4.06 | 5.45 | 0.43 | 0.01 | 0.03 | 16.67 | Detail |
| 786 | CHOL | autosis | APBA2 | amyloid beta precursor protein binding family A member 2 | 4.91 | 7.14 | 0.54 | 0.01 | 0.03 | 16.67 | Detail |
| 787 | CHOL | autosis | FOSL2 | FOS like 2, AP-1 transcription factor subunit | 9.04 | 10.37 | 0.2 | 0.01 | 0.03 | 16.67 | Detail |
| 788 | CHOL | autosis | GRB10 | growth factor receptor bound protein 10 | 10.64 | 11.22 | 0.08 | 0.01 | 0.03 | 16.67 | Detail |
| 789 | CHOL | autosis | GNAI1 | G protein subunit alpha i1 | 9.54 | 8.04 | -0.25 | 0.01 | 0.03 | 16.67 | Detail |
| 790 | CHOL | autosis | STXBP3 | syntaxin binding protein 3 | 9.4 | 10.09 | 0.1 | 0.01 | 0.03 | 16.67 | Detail |
| 791 | CHOL | autosis | WSB1 | WD repeat and SOCS box containing 1 | 10.21 | 10.98 | 0.1 | 0.01 | 0.03 | 16.67 | Detail |
| 792 | CHOL | autosis | SERP1 | stress associated endoplasmic reticulum protein 1 | 12.18 | 11.59 | -0.07 | 0.01 | 0.03 | 16.67 | Detail |
| 793 | CHOL | autosis | HOXD1 | homeobox D1 | 0.67 | 2.21 | 1.72 | 0.02 | 0.03 | 16.67 | Detail |
| 794 | CHOL | autosis | KLF3 | KLF transcription factor 3 | 11.19 | 10.5 | -0.09 | 0.01 | 0.03 | 16.67 | Detail |
| 795 | CHOL | autosis | WAC | WW domain containing adaptor with coiled-coil | 10.92 | 11.35 | 0.06 | 0.01 | 0.03 | 16.67 | Detail |
| 796 | CHOL | autosis | ATP2C2 | ATPase secretory pathway Ca2+ transporting 2 | 3.43 | 6.22 | 0.86 | 0.01 | 0.03 | 16.67 | Detail |
| 797 | CHOL | autosis | BRWD1 | bromodomain and WD repeat domain containing 1 | 10.48 | 9.81 | -0.09 | 0.01 | 0.03 | 16.67 | Detail |
| 798 | CHOL | autosis | RABGAP1 | RAB GTPase activating protein 1 | 9.23 | 9.71 | 0.07 | 0.01 | 0.03 | 16.67 | Detail |
| 799 | CHOL | autosis | COQ10B | coenzyme Q10B | 9.92 | 9.3 | -0.09 | 0.01 | 0.03 | 16.67 | Detail |
| 800 | CHOL | autosis | PRDM10 | PR/SET domain 10 | 7.1 | 7.65 | 0.11 | 0.01 | 0.03 | 16.67 | Detail |
| 801 | CHOL | autosis | ZNF557 | zinc finger protein 557 | 6.7 | 7.33 | 0.13 | 0.01 | 0.03 | 16.67 | Detail |
| 802 | CHOL | autosis | AHR | aryl hydrocarbon receptor | 11.16 | 10.07 | -0.15 | 0.02 | 0.03 | 12.5 | Detail |
| 803 | CHOL | autosis | H19 | H19 imprinted maternally expressed transcript | 13.55 | 10.53 | -0.36 | 0.02 | 0.03 | 12.5 | Detail |
| 804 | CHOL | autosis | MAPK8 | mitogen-activated protein kinase 8 | 8.18 | 7.21 | -0.18 | 0.02 | 0.03 | 12.5 | Detail |
| 805 | CHOL | autosis | NFKB1 | nuclear factor kappa B subunit 1 | 9.85 | 10.37 | 0.07 | 0.02 | 0.03 | 12.5 | Detail |
| 806 | CHOL | autosis | OSBP | oxysterol binding protein | 11.54 | 10.93 | -0.08 | 0.02 | 0.03 | 12.5 | Detail |
| 807 | CHOL | autosis | INSR | insulin receptor | 12.07 | 11.4 | -0.08 | 0.02 | 0.03 | 12.5 | Detail |
| 808 | CHOL | autosis | ANTXR1 | ANTXR cell adhesion molecule 1 | 8.78 | 10.22 | 0.22 | 0.02 | 0.03 | 12.5 | Detail |
| 809 | CHOL | autosis | BAG3 | BAG cochaperone 3 | 9.19 | 9.89 | 0.11 | 0.02 | 0.03 | 12.5 | Detail |
| 810 | CHOL | autosis | NEDD4L | NEDD4 like E3 ubiquitin protein ligase | 9.31 | 10.03 | 0.11 | 0.02 | 0.03 | 12.5 | Detail |
| 811 | CHOL | autosis | POU2F1 | POU class 2 homeobox 1 | 4.16 | 5.06 | 0.28 | 0.02 | 0.03 | 12.5 | Detail |
| 812 | CHOL | autosis | ELK4 | ETS transcription factor ELK4 | 6.68 | 7.96 | 0.25 | 0.02 | 0.03 | 12.5 | Detail |
| 813 | CHOL | autosis | STAT6 | signal transducer and activator of transcription 6 | 11.73 | 12.26 | 0.06 | 0.02 | 0.03 | 12.5 | Detail |
| 814 | CHOL | autosis | CSDE1 | cold shock domain containing E1 | 13.11 | 13.46 | 0.04 | 0.02 | 0.04 | 12.5 | Detail |
| 815 | CHOL | autosis | HCFC1R1 | host cell factor C1 regulator 1 | 9.12 | 10.19 | 0.16 | 0.02 | 0.03 | 12.5 | Detail |
| 816 | CHOL | autosis | ATRN | attractin | 12.18 | 11.09 | -0.14 | 0.02 | 0.03 | 12.5 | Detail |
| 817 | CHOL | autosis | ARMCX3 | armadillo repeat containing X-linked 3 | 9.8 | 10.39 | 0.08 | 0.02 | 0.03 | 12.5 | Detail |
| 818 | CHOL | autosis | ZNF22 | zinc finger protein 22 | 9.83 | 8.98 | -0.13 | 0.02 | 0.03 | 12.5 | Detail |
| 819 | CHOL | autosis | ATXN3 | ataxin 3 | 8.64 | 7.98 | -0.11 | 0.02 | 0.03 | 12.5 | Detail |
| 820 | CHOL | autosis | ZNF277 | zinc finger protein 277 | 9.22 | 8.74 | -0.08 | 0.02 | 0.03 | 12.5 | Detail |
| 821 | CHOL | autosis | TIPARP | TCDD inducible poly(ADP-ribose) polymerase | 9.93 | 8.91 | -0.16 | 0.02 | 0.03 | 12.5 | Detail |
| 822 | CHOL | autosis | PSG4 | pregnancy specific beta-1-glycoprotein 4 | 0.21 | 3.28 | 3.97 | 0.02 | 0.04 | 12.5 | Detail |
| 823 | CHOL | autosis | TPP1 | tripeptidyl peptidase 1 | 12.31 | 11.8 | -0.06 | 0.02 | 0.03 | 12.5 | Detail |
| 824 | CHOL | autosis | EPS8L2 | EPS8 signaling adaptor L2 | 11.61 | 12.16 | 0.07 | 0.02 | 0.03 | 12.5 | Detail |
| 825 | CHOL | autosis | ZHX3 | zinc fingers and homeoboxes 3 | 9.84 | 9.06 | -0.12 | 0.02 | 0.03 | 12.5 | Detail |
| 826 | CHOL | autosis | ZNF93 | zinc finger protein 93 | 3.79 | 5.09 | 0.43 | 0.02 | 0.03 | 12.5 | Detail |
| 827 | CHOL | autosis | ZNF507 | zinc finger protein 507 | 8.68 | 9.02 | 0.06 | 0.02 | 0.03 | 12.5 | Detail |
| 828 | CHOL | autosis | FXYD2 | FXYD domain containing ion transport regulator 2 | 8.96 | 11.1 | 0.31 | 0.02 | 0.03 | 12.5 | Detail |
| 829 | CHOL | autosis | AP1S1 | adaptor related protein complex 1 subunit sigma 1 | 9.64 | 10.25 | 0.09 | 0.02 | 0.03 | 12.5 | Detail |
| 830 | CHOL | autosis | TBC1D17 | TBC1 domain family member 17 | 10.32 | 9.9 | -0.06 | 0.02 | 0.03 | 12.5 | Detail |
| 831 | CHOL | autosis | BCL2 | BCL2 apoptosis regulator | 7.15 | 8.5 | 0.25 | 0.02 | 0.04 | 8.33 | Detail |
| 832 | CHOL | autosis | CREB1 | cAMP responsive element binding protein 1 | 8.98 | 9.46 | 0.07 | 0.02 | 0.04 | 8.33 | Detail |
| 833 | CHOL | autosis | IL18 | interleukin 18 | 7.1 | 9.72 | 0.45 | 0.02 | 0.04 | 8.33 | Detail |
| 834 | CHOL | autosis | RELA | RELA proto-oncogene, NF-kB subunit | 10.72 | 11.09 | 0.05 | 0.02 | 0.04 | 8.33 | Detail |
| 835 | CHOL | autosis | PPP1R15A | protein phosphatase 1 regulatory subunit 15A | 10.26 | 11.08 | 0.11 | 0.02 | 0.04 | 8.33 | Detail |
| 836 | CHOL | autosis | AMD1 | adenosylmethionine decarboxylase 1 | 9.91 | 10.37 | 0.07 | 0.02 | 0.04 | 8.33 | Detail |
| 837 | CHOL | autosis | ADORA1 | adenosine A1 receptor | 4.61 | 6.8 | 0.56 | 0.02 | 0.04 | 8.33 | Detail |
| 838 | CHOL | autosis | FXR1 | FMR1 autosomal homolog 1 | 10.55 | 10.91 | 0.05 | 0.02 | 0.04 | 8.33 | Detail |
| 839 | CHOL | autosis | NFYA | nuclear transcription factor Y subunit alpha | 8.84 | 9.53 | 0.11 | 0.02 | 0.04 | 8.33 | Detail |
| 840 | CHOL | autosis | SMTN | smoothelin | 9.17 | 10.55 | 0.2 | 0.02 | 0.04 | 8.33 | Detail |
| 841 | CHOL | autosis | BTN3A1 | butyrophilin subfamily 3 member A1 | 9.08 | 10.11 | 0.16 | 0.02 | 0.04 | 8.33 | Detail |
| 842 | CHOL | autosis | CALU | calumenin | 10.68 | 11.54 | 0.11 | 0.02 | 0.04 | 8.33 | Detail |
| 843 | CHOL | autosis | ATP11A | ATPase phospholipid transporting 11A | 9.81 | 10.71 | 0.13 | 0.02 | 0.04 | 8.33 | Detail |
| 844 | CHOL | autosis | TRMT12 | tRNA methyltransferase 12 homolog | 7.49 | 7.93 | 0.08 | 0.02 | 0.04 | 8.33 | Detail |
| 845 | CHOL | autosis | GMFB | glia maturation factor beta | 9.54 | 10.01 | 0.07 | 0.02 | 0.04 | 8.33 | Detail |
| 846 | CHOL | autosis | ARHGAP12 | Rho GTPase activating protein 12 | 9.38 | 10.01 | 0.09 | 0.02 | 0.04 | 8.33 | Detail |
| 847 | CHOL | autosis | BCL7C | BAF chromatin remodeling complex subunit BCL7C | 9.05 | 9.92 | 0.13 | 0.02 | 0.04 | 8.33 | Detail |
| 848 | CHOL | autosis | FOS | Fos proto-oncogene, AP-1 transcription factor subunit | 12.91 | 10.8 | -0.26 | 0.02 | 0.04 | 8.33 | Detail |
| 849 | CHOL | autosis | ITGAL | integrin subunit alpha L | 9.51 | 8.19 | -0.22 | 0.02 | 0.04 | 8.33 | Detail |
| 850 | CHOL | autosis | OASL | 2'-5'-oligoadenylate synthetase like | 7.96 | 6.94 | -0.2 | 0.02 | 0.04 | 8.33 | Detail |
| 851 | CHOL | autosis | SCG5 | secretogranin V | 6.94 | 5.51 | -0.33 | 0.02 | 0.04 | 8.33 | Detail |
| 852 | CHOL | autosis | AK2 | adenylate kinase 2 | 11.77 | 11.16 | -0.08 | 0.02 | 0.04 | 8.33 | Detail |
| 853 | CHOL | autosis | MRPL18 | mitochondrial ribosomal protein L18 | 9.89 | 9.54 | -0.05 | 0.02 | 0.04 | 8.33 | Detail |
| 854 | CHOL | autosis | ADAM10 | ADAM metallopeptidase domain 10 | 9.67 | 10.51 | 0.12 | 0.02 | 0.04 | 4.17 | Detail |
| 855 | CHOL | autosis | ICAM1 | intercellular adhesion molecule 1 | 10.29 | 11.54 | 0.16 | 0.02 | 0.04 | 4.17 | Detail |
| 856 | CHOL | autosis | PFKFB3 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 | 10.42 | 11.98 | 0.2 | 0.02 | 0.04 | 4.17 | Detail |
| 857 | CHOL | autosis | NFATC1 | nuclear factor of activated T cells 1 | 7.39 | 8.47 | 0.2 | 0.02 | 0.04 | 4.17 | Detail |
| 858 | CHOL | autosis | RFC3 | replication factor C subunit 3 | 6.96 | 7.57 | 0.12 | 0.02 | 0.04 | 4.17 | Detail |
| 859 | CHOL | autosis | TAGLN | transgelin | 10.86 | 12.16 | 0.16 | 0.02 | 0.04 | 4.17 | Detail |
| 860 | CHOL | autosis | HK2 | hexokinase 2 | 5.92 | 7.84 | 0.4 | 0.02 | 0.04 | 4.17 | Detail |
| 861 | CHOL | autosis | PDLIM5 | PDZ and LIM domain 5 | 11.77 | 11.02 | -0.09 | 0.02 | 0.04 | 4.17 | Detail |
| 862 | CHOL | autosis | HERPUD1 | homocysteine inducible ER protein with ubiquitin like domain 1 | 12.04 | 11.43 | -0.08 | 0.02 | 0.04 | 4.17 | Detail |
| 863 | CHOL | autosis | HEXIM1 | HEXIM P-TEFb complex subunit 1 | 10.2 | 10.71 | 0.07 | 0.02 | 0.04 | 4.17 | Detail |
| 864 | CHOL | autosis | PRG2 | proteoglycan 2, pro eosinophil major basic protein | 6.52 | 5.35 | -0.29 | 0.02 | 0.04 | 4.17 | Detail |
| 865 | CHOL | autosis | MORC3 | MORC family CW-type zinc finger 3 | 9.4 | 8.89 | -0.08 | 0.02 | 0.04 | 4.17 | Detail |
| 866 | CHOL | autosis | SLC25A12 | solute carrier family 25 member 12 | 5.73 | 7.17 | 0.32 | 0.02 | 0.04 | 4.17 | Detail |
| 867 | CHOL | autosis | ZSCAN5A | zinc finger and SCAN domain containing 5A | 5.41 | 6.08 | 0.17 | 0.02 | 0.04 | 4.17 | Detail |
| 868 | CHOL | autosis | TAF1C | TATA-box binding protein associated factor, RNA polymerase I subunit C | 9.52 | 10.21 | 0.1 | 0.02 | 0.04 | 4.17 | Detail |
| 869 | CHOL | autosis | ACBD3 | acyl-CoA binding domain containing 3 | 9.74 | 10.36 | 0.09 | 0.02 | 0.04 | 4.17 | Detail |
| 870 | CHOL | autosis | ANKRD10 | ankyrin repeat domain 10 | 10.42 | 11.22 | 0.11 | 0.02 | 0.04 | 4.17 | Detail |
| 871 | CHOL | autosis | CLN5 | CLN5 intracellular trafficking protein | 9.82 | 9.11 | -0.11 | 0.02 | 0.04 | 4.17 | Detail |
| 872 | CHOL | autosis | OSBPL2 | oxysterol binding protein like 2 | 9.87 | 10.36 | 0.07 | 0.02 | 0.04 | 4.17 | Detail |
| 873 | CHOL | autosis | AHI1 | Abelson helper integration site 1 | 7.2 | 7.72 | 0.1 | 0.05 | 0.09 | 0.0 | Detail |
| 874 | CHOL | autosis | AKR1C1 | aldo-keto reductase family 1 member C1 | 13.36 | 12.02 | -0.15 | 0.39 | 0.47 | 0.0 | Detail |
| 875 | CHOL | autosis | ANXA1 | annexin A1 | 8.82 | 9.54 | 0.11 | 0.39 | 0.47 | 0.0 | Detail |
| 876 | CHOL | autosis | AP5Z1 | adaptor related protein complex 5 subunit zeta 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 877 | CHOL | autosis | APC | APC regulator of WNT signaling pathway | 9.12 | 9.27 | 0.02 | 0.58 | 0.66 | 0.0 | Detail |
| 878 | CHOL | autosis | ARIH1 | ariadne RBR E3 ubiquitin protein ligase 1 | 10.12 | 10.24 | 0.02 | 0.33 | 0.41 | 0.0 | Detail |
| 879 | CHOL | autosis | ATF2 | activating transcription factor 2 | 8.28 | 8.15 | -0.02 | 0.66 | 0.72 | 0.0 | Detail |
| 880 | CHOL | autosis | ATP12A | ATPase H+/K+ transporting non-gastric alpha2 subunit | 1.77 | 1.05 | -0.76 | 0.4 | 0.49 | 0.0 | Detail |
| 881 | CHOL | autosis | ATP4A | ATPase H+/K+ transporting subunit alpha | 0.17 | 0.48 | 1.48 | 0.85 | 0.89 | 0.0 | Detail |
| 882 | CHOL | autosis | BCS1L | BCS1 homolog, ubiquinol-cytochrome c reductase complex chaperone | 8.99 | 9.13 | 0.02 | 0.42 | 0.5 | 0.0 | Detail |
| 883 | CHOL | autosis | BDNF | brain derived neurotrophic factor | 1.48 | 3.14 | 1.09 | 0.03 | 0.06 | 0.0 | Detail |
| 884 | CHOL | autosis | BRAF | B-Raf proto-oncogene, serine/threonine kinase | 6.58 | 6.8 | 0.05 | 0.48 | 0.56 | 0.0 | Detail |
| 885 | CHOL | autosis | BRD2 | bromodomain containing 2 | 11.85 | 12.13 | 0.03 | 0.19 | 0.26 | 0.0 | Detail |
| 886 | CHOL | autosis | BRD4 | bromodomain containing 4 | 10.81 | 10.64 | -0.02 | 0.44 | 0.52 | 0.0 | Detail |
| 887 | CHOL | autosis | CCL2 | C-C motif chemokine ligand 2 | 8.71 | 9.7 | 0.16 | 0.28 | 0.36 | 0.0 | Detail |
| 888 | CHOL | autosis | CCN2 | cellular communication network factor 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 889 | CHOL | autosis | CCR1 | C-C motif chemokine receptor 1 | 7.25 | 6.34 | -0.19 | 0.18 | 0.24 | 0.0 | Detail |
| 890 | CHOL | autosis | CD40 | CD40 molecule | 8.67 | 8.88 | 0.03 | 0.42 | 0.5 | 0.0 | Detail |
| 891 | CHOL | autosis | CDC42 | cell division cycle 42 | 11.87 | 11.84 | -0.0 | 0.48 | 0.56 | 0.0 | Detail |
| 892 | CHOL | autosis | CDK9 | cyclin dependent kinase 9 | 10.8 | 10.36 | -0.06 | 0.1 | 0.16 | 0.0 | Detail |
| 893 | CHOL | autosis | CHUK | component of inhibitor of nuclear factor kappa B kinase complex | 9.19 | 8.8 | -0.06 | 0.05 | 0.09 | 0.0 | Detail |
| 894 | CHOL | autosis | CPE | carboxypeptidase E | 9.75 | 8.45 | -0.21 | 0.04 | 0.06 | 0.0 | Detail |
| 895 | CHOL | autosis | CTNNB1 | catenin beta 1 | 12.35 | 11.97 | -0.04 | 0.06 | 0.1 | 0.0 | Detail |
| 896 | CHOL | autosis | CTSL | cathepsin L | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 897 | CHOL | autosis | CUX1 | cut like homeobox 1 | 10.63 | 10.92 | 0.04 | 0.7 | 0.76 | 0.0 | Detail |
| 898 | CHOL | autosis | CXCL8 | C-X-C motif chemokine ligand 8 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 899 | CHOL | autosis | CYCS | cytochrome c, somatic | 11.43 | 11.31 | -0.01 | 0.66 | 0.72 | 0.0 | Detail |
| 900 | CHOL | autosis | DAAM1 | dishevelled associated activator of morphogenesis 1 | 8.65 | 7.82 | -0.14 | 0.09 | 0.14 | 0.0 | Detail |
| 901 | CHOL | autosis | DUSP5 | dual specificity phosphatase 5 | 9.14 | 9.53 | 0.06 | 0.33 | 0.41 | 0.0 | Detail |
| 902 | CHOL | autosis | DYRK3 | dual specificity tyrosine phosphorylation regulated kinase 3 | 6.68 | 6.97 | 0.06 | 0.48 | 0.56 | 0.0 | Detail |
| 903 | CHOL | autosis | EDN1 | endothelin 1 | 6.76 | 7.45 | 0.14 | 0.16 | 0.23 | 0.0 | Detail |
| 904 | CHOL | autosis | EIF2AK3 | eukaryotic translation initiation factor 2 alpha kinase 3 | 8.81 | 9.06 | 0.04 | 0.33 | 0.41 | 0.0 | Detail |
| 905 | CHOL | autosis | EPHA2 | EPH receptor A2 | 10.8 | 10.1 | -0.1 | 0.08 | 0.13 | 0.0 | Detail |
| 906 | CHOL | autosis | ERBB2 | erb-b2 receptor tyrosine kinase 2 | 11.96 | 12.16 | 0.02 | 0.26 | 0.34 | 0.0 | Detail |
| 907 | CHOL | autosis | ERVW-1 | endogenous retrovirus group W member 1, envelope | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 908 | CHOL | autosis | EZR | ezrin | 11.38 | 12.16 | 0.1 | 0.04 | 0.06 | 0.0 | Detail |
| 909 | CHOL | autosis | F2R | coagulation factor II thrombin receptor | 9.4 | 9.73 | 0.05 | 0.42 | 0.5 | 0.0 | Detail |
| 910 | CHOL | autosis | FASTK | Fas activated serine/threonine kinase | 10.76 | 11.09 | 0.04 | 0.24 | 0.31 | 0.0 | Detail |
| 911 | CHOL | autosis | FBN1 | fibrillin 1 | 9.0 | 9.93 | 0.14 | 0.06 | 0.1 | 0.0 | Detail |
| 912 | CHOL | autosis | FDXR | ferredoxin reductase | 9.57 | 9.28 | -0.04 | 0.33 | 0.41 | 0.0 | Detail |
| 913 | CHOL | autosis | FGF2 | fibroblast growth factor 2 | 7.52 | 6.13 | -0.3 | 0.06 | 0.1 | 0.0 | Detail |
| 914 | CHOL | autosis | FN1 | fibronectin 1 | 17.15 | 15.52 | -0.14 | 0.06 | 0.1 | 0.0 | Detail |
| 915 | CHOL | autosis | GAK | cyclin G associated kinase | 10.94 | 11.21 | 0.03 | 0.07 | 0.12 | 0.0 | Detail |
| 916 | CHOL | autosis | GBP1 | guanylate binding protein 1 | 10.58 | 9.47 | -0.16 | 0.04 | 0.07 | 0.0 | Detail |
| 917 | CHOL | autosis | GDF15 | growth differentiation factor 15 | 9.79 | 11.22 | 0.2 | 0.05 | 0.08 | 0.0 | Detail |
| 918 | CHOL | autosis | GLB1 | galactosidase beta 1 | 10.7 | 10.27 | -0.06 | 0.07 | 0.12 | 0.0 | Detail |
| 919 | CHOL | autosis | GNAS | GNAS complex locus | 13.93 | 13.67 | -0.03 | 0.66 | 0.72 | 0.0 | Detail |
| 920 | CHOL | autosis | GOLGB1 | golgin B1 | 11.35 | 11.72 | 0.05 | 0.07 | 0.12 | 0.0 | Detail |
| 921 | CHOL | autosis | GOLPH3 | golgi phosphoprotein 3 | 11.83 | 11.4 | -0.05 | 0.12 | 0.17 | 0.0 | Detail |
| 922 | CHOL | autosis | GPNMB | glycoprotein nmb | 9.24 | 10.25 | 0.15 | 0.13 | 0.19 | 0.0 | Detail |
| 923 | CHOL | autosis | GSK3B | glycogen synthase kinase 3 beta | 9.71 | 10.07 | 0.05 | 0.16 | 0.23 | 0.0 | Detail |
| 924 | CHOL | autosis | GTF2B | general transcription factor IIB | 9.38 | 9.08 | -0.05 | 0.05 | 0.09 | 0.0 | Detail |
| 925 | CHOL | autosis | HHEX | hematopoietically expressed homeobox | 10.57 | 10.3 | -0.04 | 0.51 | 0.59 | 0.0 | Detail |
| 926 | CHOL | autosis | HLA-C | major histocompatibility complex, class I, C | 14.05 | 14.48 | 0.04 | 0.28 | 0.36 | 0.0 | Detail |
| 927 | CHOL | autosis | HLA-DPA1 | major histocompatibility complex, class II, DP alpha 1 | 11.73 | 11.67 | -0.01 | 0.86 | 0.89 | 0.0 | Detail |
| 928 | CHOL | autosis | HLA-DQA1 | major histocompatibility complex, class II, DQ alpha 1 | 9.54 | 9.4 | -0.02 | 0.94 | 0.95 | 0.0 | Detail |
| 929 | CHOL | autosis | HMOX1 | heme oxygenase 1 | 10.76 | 9.64 | -0.16 | 0.07 | 0.12 | 0.0 | Detail |
| 930 | CHOL | autosis | HMOX2 | heme oxygenase 2 | 10.8 | 10.26 | -0.07 | 0.06 | 0.1 | 0.0 | Detail |
| 931 | CHOL | autosis | HSP90AA1 | heat shock protein 90 alpha family class A member 1 | 13.54 | 13.7 | 0.02 | 0.58 | 0.66 | 0.0 | Detail |
| 932 | CHOL | autosis | HSPD1 | heat shock protein family D (Hsp60) member 1 | 13.55 | 13.01 | -0.06 | 0.05 | 0.08 | 0.0 | Detail |
| 933 | CHOL | autosis | IKBKG | inhibitor of nuclear factor kappa B kinase regulatory subunit gamma | 9.35 | 9.4 | 0.01 | 0.74 | 0.79 | 0.0 | Detail |
| 934 | CHOL | autosis | IL6 | interleukin 6 | 2.39 | 4.32 | 0.86 | 0.16 | 0.23 | 0.0 | Detail |
| 935 | CHOL | autosis | IL6ST | interleukin 6 cytokine family signal transducer | 10.5 | 8.73 | -0.27 | 0.09 | 0.14 | 0.0 | Detail |
| 936 | CHOL | autosis | IL9 | interleukin 9 | 0.0 | 0.02 | inf | 0.73 | 0.79 | 0.0 | Detail |
| 937 | CHOL | autosis | IP6K1 | inositol hexakisphosphate kinase 1 | 9.35 | 9.78 | 0.06 | 0.04 | 0.06 | 0.0 | Detail |
| 938 | CHOL | autosis | IP6K2 | inositol hexakisphosphate kinase 2 | 9.62 | 9.99 | 0.06 | 0.13 | 0.19 | 0.0 | Detail |
| 939 | CHOL | autosis | IRF1 | interferon regulatory factor 1 | 10.2 | 10.61 | 0.06 | 0.24 | 0.31 | 0.0 | Detail |
| 940 | CHOL | autosis | IRF7 | interferon regulatory factor 7 | 9.57 | 9.67 | 0.01 | 0.94 | 0.95 | 0.0 | Detail |
| 941 | CHOL | autosis | IRF9 | interferon regulatory factor 9 | 10.21 | 10.27 | 0.01 | 0.7 | 0.76 | 0.0 | Detail |
| 942 | CHOL | autosis | ISG15 | ISG15 ubiquitin like modifier | 9.71 | 9.4 | -0.05 | 0.51 | 0.59 | 0.0 | Detail |
| 943 | CHOL | autosis | ITGB3 | integrin subunit beta 3 | 6.61 | 5.9 | -0.16 | 0.18 | 0.24 | 0.0 | Detail |
| 944 | CHOL | autosis | KITLG | KIT ligand | 6.64 | 7.27 | 0.13 | 0.36 | 0.44 | 0.0 | Detail |
| 945 | CHOL | autosis | MAPK14 | mitogen-activated protein kinase 14 | 10.71 | 10.3 | -0.06 | 0.05 | 0.08 | 0.0 | Detail |
| 946 | CHOL | autosis | MARK3 | microtubule affinity regulating kinase 3 | 9.73 | 9.82 | 0.01 | 1.0 | 1.0 | 0.0 | Detail |
| 947 | CHOL | autosis | MBTPS2 | membrane bound transcription factor peptidase, site 2 | 9.54 | 9.18 | -0.05 | 0.24 | 0.31 | 0.0 | Detail |
| 948 | CHOL | autosis | MCAT | malonyl-CoA-acyl carrier protein transacylase | 8.94 | 8.9 | -0.01 | 0.48 | 0.56 | 0.0 | Detail |
| 949 | CHOL | autosis | MCL1 | MCL1 apoptosis regulator, BCL2 family member | 13.68 | 13.36 | -0.03 | 0.45 | 0.53 | 0.0 | Detail |
| 950 | CHOL | autosis | MLXIP | MLX interacting protein | 8.89 | 9.0 | 0.02 | 0.62 | 0.69 | 0.0 | Detail |
| 951 | CHOL | autosis | MTUS1 | microtubule associated scaffold protein 1 | 11.45 | 11.05 | -0.05 | 0.24 | 0.31 | 0.0 | Detail |
| 952 | CHOL | autosis | NDUFB7 | NADH:ubiquinone oxidoreductase subunit B7 | 11.31 | 10.9 | -0.05 | 0.1 | 0.16 | 0.0 | Detail |
| 953 | CHOL | autosis | NFE2L2 | NFE2 like bZIP transcription factor 2 | 11.66 | 11.06 | -0.08 | 0.07 | 0.12 | 0.0 | Detail |
| 954 | CHOL | autosis | NFKBIA | NFKB inhibitor alpha | 12.08 | 11.71 | -0.04 | 0.12 | 0.17 | 0.0 | Detail |
| 955 | CHOL | autosis | NGF | nerve growth factor | 4.71 | 3.42 | -0.46 | 0.03 | 0.05 | 0.0 | Detail |
| 956 | CHOL | autosis | NLRP3 | NLR family pyrin domain containing 3 | 5.61 | 5.63 | 0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 957 | CHOL | autosis | NR5A1 | nuclear receptor subfamily 5 group A member 1 | 0.65 | 0.54 | -0.27 | 0.98 | 0.98 | 0.0 | Detail |
| 958 | CHOL | autosis | NXF1 | nuclear RNA export factor 1 | 10.61 | 10.95 | 0.05 | 0.05 | 0.09 | 0.0 | Detail |
| 959 | CHOL | autosis | PAFAH1B1 | platelet activating factor acetylhydrolase 1b regulatory subunit 1 | 10.95 | 11.27 | 0.04 | 0.07 | 0.11 | 0.0 | Detail |
| 960 | CHOL | autosis | PAGR1 | PAXIP1 associated glutamate rich protein 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 961 | CHOL | autosis | PGF | placental growth factor | 7.06 | 8.02 | 0.18 | 0.08 | 0.13 | 0.0 | Detail |
| 962 | CHOL | autosis | PHB | prohibitin 1 | 11.85 | 11.98 | 0.02 | 0.81 | 0.86 | 0.0 | Detail |
| 963 | CHOL | autosis | PIK3CA | phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha | 8.11 | 8.23 | 0.02 | 0.42 | 0.5 | 0.0 | Detail |
| 964 | CHOL | autosis | PLA2G15 | phospholipase A2 group XV | 9.09 | 9.4 | 0.05 | 0.42 | 0.5 | 0.0 | Detail |
| 965 | CHOL | autosis | PLA2G4A | phospholipase A2 group IVA | 4.52 | 4.57 | 0.02 | 0.9 | 0.92 | 0.0 | Detail |
| 966 | CHOL | autosis | PLK3 | polo like kinase 3 | 7.68 | 8.81 | 0.2 | 0.18 | 0.24 | 0.0 | Detail |
| 967 | CHOL | autosis | PLXNA2 | plexin A2 | 8.4 | 9.07 | 0.11 | 0.18 | 0.24 | 0.0 | Detail |
| 968 | CHOL | autosis | PSEN1 | presenilin 1 | 10.33 | 10.37 | 0.01 | 0.66 | 0.72 | 0.0 | Detail |
| 969 | CHOL | autosis | PTGS1 | prostaglandin-endoperoxide synthase 1 | 7.14 | 8.02 | 0.17 | 0.05 | 0.09 | 0.0 | Detail |
| 970 | CHOL | autosis | PTGS2 | prostaglandin-endoperoxide synthase 2 | 5.82 | 5.73 | -0.02 | 0.81 | 0.86 | 0.0 | Detail |
| 971 | CHOL | autosis | RAB40B | RAB40B, member RAS oncogene family | 8.57 | 8.78 | 0.04 | 0.45 | 0.53 | 0.0 | Detail |
| 972 | CHOL | autosis | RB1 | RB transcriptional corepressor 1 | 9.22 | 9.67 | 0.07 | 0.06 | 0.11 | 0.0 | Detail |
| 973 | CHOL | autosis | REL | REL proto-oncogene, NF-kB subunit | 5.47 | 4.86 | -0.17 | 0.51 | 0.59 | 0.0 | Detail |
| 974 | CHOL | autosis | RHOD | ras homolog family member D | 10.36 | 9.67 | -0.1 | 0.04 | 0.06 | 0.0 | Detail |
| 975 | CHOL | autosis | RNASE1 | ribonuclease A family member 1, pancreatic | 9.09 | 9.72 | 0.1 | 0.14 | 0.21 | 0.0 | Detail |
| 976 | CHOL | autosis | RRAD | RRAD, Ras related glycolysis inhibitor and calcium channel regulator | 5.32 | 6.75 | 0.35 | 0.12 | 0.17 | 0.0 | Detail |
| 977 | CHOL | autosis | RSAD2 | radical S-adenosyl methionine domain containing 2 | 6.46 | 7.09 | 0.13 | 0.19 | 0.26 | 0.0 | Detail |
| 978 | CHOL | autosis | S100A1 | S100 calcium binding protein A1 | 5.19 | 7.23 | 0.48 | 0.18 | 0.24 | 0.0 | Detail |
| 979 | CHOL | autosis | SETD2 | SET domain containing 2, histone lysine methyltransferase | 10.54 | 10.2 | -0.05 | 0.04 | 0.06 | 0.0 | Detail |
| 980 | CHOL | autosis | SGPL1 | sphingosine-1-phosphate lyase 1 | 10.85 | 10.99 | 0.02 | 0.31 | 0.39 | 0.0 | Detail |
| 981 | CHOL | autosis | SIM2 | SIM bHLH transcription factor 2 | 5.02 | 6.4 | 0.35 | 0.04 | 0.07 | 0.0 | Detail |
| 982 | CHOL | autosis | SLC33A1 | solute carrier family 33 member 1 | 9.36 | 9.31 | -0.01 | 0.98 | 0.98 | 0.0 | Detail |
| 983 | CHOL | autosis | SLC38A7 | solute carrier family 38 member 7 | 8.26 | 7.81 | -0.08 | 0.36 | 0.44 | 0.0 | Detail |
| 984 | CHOL | autosis | SLC3A2 | solute carrier family 3 member 2 | 10.92 | 11.48 | 0.07 | 0.05 | 0.08 | 0.0 | Detail |
| 985 | CHOL | autosis | SOCS1 | suppressor of cytokine signaling 1 | 5.83 | 6.35 | 0.12 | 0.58 | 0.66 | 0.0 | Detail |
| 986 | CHOL | autosis | SOD2 | superoxide dismutase 2 | 13.83 | 12.99 | -0.09 | 0.09 | 0.14 | 0.0 | Detail |
| 987 | CHOL | autosis | SOS1 | SOS Ras/Rac guanine nucleotide exchange factor 1 | 9.74 | 9.81 | 0.01 | 0.31 | 0.39 | 0.0 | Detail |
| 988 | CHOL | autosis | SPAG9 | sperm associated antigen 9 | 10.65 | 10.37 | -0.04 | 0.62 | 0.69 | 0.0 | Detail |
| 989 | CHOL | autosis | SRF | serum response factor | 9.71 | 10.15 | 0.06 | 0.06 | 0.1 | 0.0 | Detail |
| 990 | CHOL | autosis | ST3GAL4 | ST3 beta-galactoside alpha-2,3-sialyltransferase 4 | 8.87 | 8.97 | 0.02 | 0.98 | 0.98 | 0.0 | Detail |
| 991 | CHOL | autosis | TBCA | tubulin folding cofactor A | 10.91 | 10.56 | -0.05 | 0.13 | 0.19 | 0.0 | Detail |
| 992 | CHOL | autosis | TFDP2 | transcription factor Dp-2 | 7.82 | 8.04 | 0.04 | 0.58 | 0.66 | 0.0 | Detail |
| 993 | CHOL | autosis | THBD | thrombomodulin | 9.16 | 8.71 | -0.07 | 0.42 | 0.5 | 0.0 | Detail |
| 994 | CHOL | autosis | TJP1 | tight junction protein 1 | 10.59 | 11.0 | 0.05 | 0.07 | 0.12 | 0.0 | Detail |
| 995 | CHOL | autosis | TLE1 | TLE family member 1, transcriptional corepressor | 11.01 | 10.49 | -0.07 | 0.1 | 0.16 | 0.0 | Detail |
| 996 | CHOL | autosis | TLE4 | TLE family member 4, transcriptional corepressor | 7.83 | 7.31 | -0.1 | 0.7 | 0.76 | 0.0 | Detail |
| 997 | CHOL | autosis | TLR2 | toll like receptor 2 | 7.81 | 7.79 | -0.0 | 0.81 | 0.86 | 0.0 | Detail |
| 998 | CHOL | autosis | TNF | tumor necrosis factor | 3.06 | 4.01 | 0.39 | 0.42 | 0.5 | 0.0 | Detail |
| 999 | CHOL | autosis | TRIB3 | tribbles pseudokinase 3 | 10.38 | 10.68 | 0.04 | 0.36 | 0.44 | 0.0 | Detail |
| 1000 | CHOL | autosis | TXNRD1 | thioredoxin reductase 1 | 10.6 | 11.07 | 0.06 | 0.36 | 0.44 | 0.0 | Detail |
| 1001 | CHOL | autosis | UGCG | UDP-glucose ceramide glucosyltransferase | 7.51 | 7.37 | -0.03 | 0.94 | 0.95 | 0.0 | Detail |
| 1002 | CHOL | autosis | UNG | uracil DNA glycosylase | 10.21 | 9.88 | -0.05 | 0.07 | 0.12 | 0.0 | Detail |
| 1003 | CHOL | autosis | USP13 | ubiquitin specific peptidase 13 | 8.63 | 8.37 | -0.04 | 0.18 | 0.24 | 0.0 | Detail |
| 1004 | CHOL | autosis | UTP25 | UTP25 small subunit processome component | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1005 | CHOL | autosis | VCAM1 | vascular cell adhesion molecule 1 | 9.25 | 8.72 | -0.09 | 0.55 | 0.63 | 0.0 | Detail |
| 1006 | CHOL | autosis | VEGFA | vascular endothelial growth factor A | 11.75 | 12.3 | 0.07 | 0.1 | 0.16 | 0.0 | Detail |
| 1007 | CHOL | autosis | WNT5A | Wnt family member 5A | 7.63 | 6.99 | -0.13 | 0.28 | 0.36 | 0.0 | Detail |
| 1008 | CHOL | autosis | ZC3HAV1 | zinc finger CCCH-type containing, antiviral 1 | 9.51 | 9.62 | 0.02 | 0.42 | 0.5 | 0.0 | Detail |
| 1009 | CHOL | autosis | SPRY2 | sprouty RTK signaling antagonist 2 | 9.52 | 8.77 | -0.12 | 0.09 | 0.14 | 0.0 | Detail |
| 1010 | CHOL | autosis | MYC | MYC proto-oncogene, bHLH transcription factor | 10.3 | 10.68 | 0.05 | 0.28 | 0.36 | 0.0 | Detail |
| 1011 | CHOL | autosis | CDKN1C | cyclin dependent kinase inhibitor 1C | 7.37 | 7.25 | -0.02 | 0.51 | 0.59 | 0.0 | Detail |
| 1012 | CHOL | autosis | FGFR4 | fibroblast growth factor receptor 4 | 11.78 | 12.01 | 0.03 | 0.06 | 0.1 | 0.0 | Detail |
| 1013 | CHOL | autosis | TSC1 | TSC complex subunit 1 | 8.74 | 9.2 | 0.07 | 0.04 | 0.07 | 0.0 | Detail |
| 1014 | CHOL | autosis | CCND1 | cyclin D1 | 12.29 | 12.59 | 0.04 | 0.66 | 0.72 | 0.0 | Detail |
| 1015 | CHOL | autosis | RASSF1 | Ras association domain family member 1 | 8.52 | 8.94 | 0.07 | 0.08 | 0.13 | 0.0 | Detail |
| 1016 | CHOL | autosis | RNH1 | ribonuclease/angiogenin inhibitor 1 | 11.68 | 11.87 | 0.02 | 0.16 | 0.23 | 0.0 | Detail |
| 1017 | CHOL | autosis | PRDX2 | peroxiredoxin 2 | 12.27 | 11.83 | -0.05 | 0.13 | 0.19 | 0.0 | Detail |
| 1018 | CHOL | autosis | FOXL2 | forkhead box L2 | 0.8 | 0.83 | 0.04 | 0.51 | 0.59 | 0.0 | Detail |
| 1019 | CHOL | autosis | NCOA1 | nuclear receptor coactivator 1 | 9.98 | 9.82 | -0.02 | 0.58 | 0.66 | 0.0 | Detail |
| 1020 | CHOL | autosis | INHBA | inhibin subunit beta A | 6.26 | 5.94 | -0.07 | 0.74 | 0.79 | 0.0 | Detail |
| 1021 | CHOL | autosis | MDH2 | malate dehydrogenase 2 | 12.46 | 12.27 | -0.02 | 0.51 | 0.59 | 0.0 | Detail |
| 1022 | CHOL | autosis | CREBBP | CREB binding protein | 10.78 | 10.95 | 0.02 | 0.39 | 0.47 | 0.0 | Detail |
| 1023 | CHOL | autosis | CYP1B1 | cytochrome P450 family 1 subfamily B member 1 | 8.06 | 8.2 | 0.03 | 0.51 | 0.59 | 0.0 | Detail |
| 1024 | CHOL | autosis | DAPK1 | death associated protein kinase 1 | 11.05 | 10.65 | -0.05 | 0.36 | 0.44 | 0.0 | Detail |
| 1025 | CHOL | autosis | EIF4E | eukaryotic translation initiation factor 4E | 8.32 | 8.58 | 0.04 | 0.33 | 0.41 | 0.0 | Detail |
| 1026 | CHOL | autosis | GABPA | GA binding protein transcription factor subunit alpha | 9.59 | 9.25 | -0.05 | 0.05 | 0.09 | 0.0 | Detail |
| 1027 | CHOL | autosis | HDAC5 | histone deacetylase 5 | 9.88 | 10.37 | 0.07 | 0.1 | 0.16 | 0.0 | Detail |
| 1028 | CHOL | autosis | HSPA2 | heat shock protein family A (Hsp70) member 2 | 7.26 | 7.45 | 0.04 | 0.7 | 0.76 | 0.0 | Detail |
| 1029 | CHOL | autosis | IMP3 | IMP U3 small nucleolar ribonucleoprotein 3 | 10.57 | 10.24 | -0.05 | 0.03 | 0.06 | 0.0 | Detail |
| 1030 | CHOL | autosis | KDM6A | lysine demethylase 6A | 8.63 | 9.46 | 0.13 | 0.04 | 0.07 | 0.0 | Detail |
| 1031 | CHOL | autosis | NCOA3 | nuclear receptor coactivator 3 | 9.54 | 9.96 | 0.06 | 0.09 | 0.14 | 0.0 | Detail |
| 1032 | CHOL | autosis | NOTCH1 | notch receptor 1 | 9.47 | 10.0 | 0.08 | 0.09 | 0.14 | 0.0 | Detail |
| 1033 | CHOL | autosis | RSF1 | remodeling and spacing factor 1 | 8.44 | 8.59 | 0.03 | 0.48 | 0.56 | 0.0 | Detail |
| 1034 | CHOL | autosis | SKP2 | S-phase kinase associated protein 2 | 7.0 | 6.31 | -0.15 | 0.08 | 0.13 | 0.0 | Detail |
| 1035 | CHOL | autosis | STAG2 | STAG2 cohesin complex component | 10.41 | 10.66 | 0.04 | 0.13 | 0.19 | 0.0 | Detail |
| 1036 | CHOL | autosis | CDKN1A | cyclin dependent kinase inhibitor 1A | 11.9 | 11.79 | -0.01 | 0.77 | 0.83 | 0.0 | Detail |
| 1037 | CHOL | autosis | ARHGEF7 | Rho guanine nucleotide exchange factor 7 | 10.0 | 10.04 | 0.01 | 0.58 | 0.66 | 0.0 | Detail |
| 1038 | CHOL | autosis | CDC42EP3 | CDC42 effector protein 3 | 7.52 | 7.96 | 0.08 | 0.39 | 0.47 | 0.0 | Detail |
| 1039 | CHOL | autosis | DDR2 | discoidin domain receptor tyrosine kinase 2 | 4.94 | 4.44 | -0.15 | 0.81 | 0.86 | 0.0 | Detail |
| 1040 | CHOL | autosis | ELK1 | ETS transcription factor ELK1 | 9.75 | 9.58 | -0.03 | 0.33 | 0.41 | 0.0 | Detail |
| 1041 | CHOL | autosis | GEM | GTP binding protein overexpressed in skeletal muscle | 7.84 | 8.46 | 0.11 | 0.28 | 0.36 | 0.0 | Detail |
| 1042 | CHOL | autosis | GPX2 | glutathione peroxidase 2 | 11.32 | 11.29 | -0.0 | 0.77 | 0.83 | 0.0 | Detail |
| 1043 | CHOL | autosis | HSPA1B | heat shock protein family A (Hsp70) member 1B | 9.23 | 10.29 | 0.16 | 0.1 | 0.16 | 0.0 | Detail |
| 1044 | CHOL | autosis | IGFBP5 | insulin like growth factor binding protein 5 | 10.67 | 11.49 | 0.11 | 0.06 | 0.1 | 0.0 | Detail |
| 1045 | CHOL | autosis | MLH1 | mutL homolog 1 | 9.23 | 8.98 | -0.04 | 0.1 | 0.16 | 0.0 | Detail |
| 1046 | CHOL | autosis | PSG2 | pregnancy specific beta-1-glycoprotein 2 | 0.0 | 0.41 | inf | 0.17 | 0.24 | 0.0 | Detail |
| 1047 | CHOL | autosis | TGFBI | transforming growth factor beta induced | 12.02 | 12.36 | 0.04 | 0.42 | 0.5 | 0.0 | Detail |
| 1048 | CHOL | autosis | THBS1 | thrombospondin 1 | 12.36 | 11.96 | -0.05 | 0.55 | 0.63 | 0.0 | Detail |
| 1049 | CHOL | autosis | TMX1 | thioredoxin related transmembrane protein 1 | 9.55 | 9.36 | -0.03 | 0.81 | 0.86 | 0.0 | Detail |
| 1050 | CHOL | autosis | TSC2 | TSC complex subunit 2 | 10.6 | 10.84 | 0.03 | 0.08 | 0.13 | 0.0 | Detail |
| 1051 | CHOL | autosis | UBQLN2 | ubiquilin 2 | 10.09 | 9.98 | -0.02 | 0.81 | 0.86 | 0.0 | Detail |
| 1052 | CHOL | autosis | ZHX2 | zinc fingers and homeoboxes 2 | 9.81 | 10.23 | 0.06 | 0.12 | 0.17 | 0.0 | Detail |
| 1053 | CHOL | autosis | GALNT14 | polypeptide N-acetylgalactosaminyltransferase 14 | 4.83 | 6.83 | 0.5 | 0.05 | 0.08 | 0.0 | Detail |
| 1054 | CHOL | autosis | PSMB9 | proteasome 20S subunit beta 9 | 8.96 | 9.79 | 0.13 | 0.13 | 0.19 | 0.0 | Detail |
| 1055 | CHOL | autosis | CORO1A | coronin 1A | 9.19 | 9.65 | 0.07 | 0.48 | 0.56 | 0.0 | Detail |
| 1056 | CHOL | autosis | FABP4 | fatty acid binding protein 4 | 5.67 | 3.89 | -0.54 | 0.05 | 0.08 | 0.0 | Detail |
| 1057 | CHOL | autosis | ID3 | inhibitor of DNA binding 3 | 9.41 | 9.92 | 0.08 | 0.33 | 0.41 | 0.0 | Detail |
| 1058 | CHOL | autosis | MAP1LC3B | microtubule associated protein 1 light chain 3 beta | 10.46 | 10.78 | 0.04 | 0.21 | 0.29 | 0.0 | Detail |
| 1059 | CHOL | autosis | TGFBR2 | transforming growth factor beta receptor 2 | 11.18 | 11.04 | -0.02 | 0.66 | 0.72 | 0.0 | Detail |
| 1060 | CHOL | autosis | TSC22D1 | TSC22 domain family member 1 | 11.69 | 11.33 | -0.05 | 0.28 | 0.36 | 0.0 | Detail |
| 1061 | CHOL | autosis | UCP2 | uncoupling protein 2 | 9.36 | 9.4 | 0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1062 | CHOL | autosis | PLK2 | polo like kinase 2 | 9.84 | 10.24 | 0.06 | 0.24 | 0.31 | 0.0 | Detail |
| 1063 | CHOL | autosis | EFEMP1 | EGF containing fibulin extracellular matrix protein 1 | 8.42 | 9.59 | 0.19 | 0.16 | 0.23 | 0.0 | Detail |
| 1064 | CHOL | autosis | NR2C2 | nuclear receptor subfamily 2 group C member 2 | 8.71 | 8.95 | 0.04 | 0.13 | 0.19 | 0.0 | Detail |
| 1065 | CHOL | autosis | ATF3 | activating transcription factor 3 | 10.44 | 9.25 | -0.18 | 0.03 | 0.06 | 0.0 | Detail |
| 1066 | CHOL | autosis | CCN1 | cellular communication network factor 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1067 | CHOL | autosis | DFFB | DNA fragmentation factor subunit beta | 6.41 | 6.81 | 0.09 | 0.18 | 0.24 | 0.0 | Detail |
| 1068 | CHOL | autosis | FGF13 | fibroblast growth factor 13 | 5.55 | 6.25 | 0.17 | 0.31 | 0.39 | 0.0 | Detail |
| 1069 | CHOL | autosis | ATF7IP | activating transcription factor 7 interacting protein | 8.89 | 9.37 | 0.08 | 0.13 | 0.19 | 0.0 | Detail |
| 1070 | CHOL | autosis | ERCC4 | ERCC excision repair 4, endonuclease catalytic subunit | 7.58 | 7.14 | -0.09 | 0.16 | 0.23 | 0.0 | Detail |
| 1071 | CHOL | autosis | PSMD12 | proteasome 26S subunit, non-ATPase 12 | 9.84 | 9.38 | -0.07 | 0.04 | 0.07 | 0.0 | Detail |
| 1072 | CHOL | autosis | FBXO5 | F-box protein 5 | 6.32 | 6.56 | 0.05 | 0.55 | 0.63 | 0.0 | Detail |
| 1073 | CHOL | autosis | AHSA1 | activator of HSP90 ATPase activity 1 | 10.39 | 10.67 | 0.04 | 0.08 | 0.13 | 0.0 | Detail |
| 1074 | CHOL | autosis | BCL2A1 | BCL2 related protein A1 | 4.33 | 5.86 | 0.44 | 0.13 | 0.19 | 0.0 | Detail |
| 1075 | CHOL | autosis | BHLHE40 | basic helix-loop-helix family member e40 | 12.66 | 11.41 | -0.15 | 0.03 | 0.06 | 0.0 | Detail |
| 1076 | CHOL | autosis | BRIP1 | BRCA1 interacting helicase 1 | 4.73 | 5.51 | 0.22 | 0.1 | 0.16 | 0.0 | Detail |
| 1077 | CHOL | autosis | CDK8 | cyclin dependent kinase 8 | 7.06 | 6.73 | -0.07 | 0.51 | 0.59 | 0.0 | Detail |
| 1078 | CHOL | autosis | DKK1 | dickkopf WNT signaling pathway inhibitor 1 | 2.02 | 5.81 | 1.53 | 0.03 | 0.06 | 0.0 | Detail |
| 1079 | CHOL | autosis | EGLN2 | egl-9 family hypoxia inducible factor 2 | 10.5 | 10.75 | 0.03 | 0.36 | 0.44 | 0.0 | Detail |
| 1080 | CHOL | autosis | EP300 | E1A binding protein p300 | 10.49 | 10.64 | 0.02 | 0.58 | 0.66 | 0.0 | Detail |
| 1081 | CHOL | autosis | ETS1 | ETS proto-oncogene 1, transcription factor | 10.44 | 10.59 | 0.02 | 0.62 | 0.69 | 0.0 | Detail |
| 1082 | CHOL | autosis | F2RL1 | F2R like trypsin receptor 1 | 9.05 | 10.04 | 0.15 | 0.06 | 0.1 | 0.0 | Detail |
| 1083 | CHOL | autosis | FGFBP1 | fibroblast growth factor binding protein 1 | 0.75 | 1.38 | 0.88 | 0.45 | 0.53 | 0.0 | Detail |
| 1084 | CHOL | autosis | GCLM | glutamate-cysteine ligase modifier subunit | 9.88 | 9.03 | -0.13 | 0.03 | 0.05 | 0.0 | Detail |
| 1085 | CHOL | autosis | GTF2H1 | general transcription factor IIH subunit 1 | 9.15 | 9.34 | 0.03 | 0.18 | 0.24 | 0.0 | Detail |
| 1086 | CHOL | autosis | HBEGF | heparin binding EGF like growth factor | 7.12 | 8.05 | 0.18 | 0.19 | 0.26 | 0.0 | Detail |
| 1087 | CHOL | autosis | HES1 | hes family bHLH transcription factor 1 | 10.25 | 10.41 | 0.02 | 0.45 | 0.53 | 0.0 | Detail |
| 1088 | CHOL | autosis | HSPA4 | heat shock protein family A (Hsp70) member 4 | 10.7 | 11.0 | 0.04 | 0.12 | 0.17 | 0.0 | Detail |
| 1089 | CHOL | autosis | HYAL2 | hyaluronidase 2 | 10.12 | 10.23 | 0.02 | 0.94 | 0.95 | 0.0 | Detail |
| 1090 | CHOL | autosis | KDM2A | lysine demethylase 2A | 11.12 | 11.33 | 0.03 | 0.28 | 0.36 | 0.0 | Detail |
| 1091 | CHOL | autosis | PDCD4 | programmed cell death 4 | 10.33 | 10.74 | 0.06 | 0.28 | 0.36 | 0.0 | Detail |
| 1092 | CHOL | autosis | PIM1 | Pim-1 proto-oncogene, serine/threonine kinase | 9.61 | 9.36 | -0.04 | 0.74 | 0.79 | 0.0 | Detail |
| 1093 | CHOL | autosis | PIMREG | PICALM interacting mitotic regulator | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1094 | CHOL | autosis | PKD1 | polycystin 1, transient receptor potential channel interacting | 10.12 | 10.53 | 0.06 | 0.05 | 0.09 | 0.0 | Detail |
| 1095 | CHOL | autosis | RAB7A | RAB7A, member RAS oncogene family | 12.48 | 12.15 | -0.04 | 0.04 | 0.07 | 0.0 | Detail |
| 1096 | CHOL | autosis | SLC2A3 | solute carrier family 2 member 3 | 7.56 | 8.94 | 0.24 | 0.07 | 0.12 | 0.0 | Detail |
| 1097 | CHOL | autosis | SMAD4 | SMAD family member 4 | 10.64 | 10.5 | -0.02 | 0.28 | 0.36 | 0.0 | Detail |
| 1098 | CHOL | autosis | TARDBP | TAR DNA binding protein | 11.52 | 11.44 | -0.01 | 0.58 | 0.66 | 0.0 | Detail |
| 1099 | CHOL | autosis | TNFRSF11B | TNF receptor superfamily member 11b | 8.33 | 8.27 | -0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1100 | CHOL | autosis | TUSC2 | tumor suppressor 2, mitochondrial calcium regulator | 9.11 | 8.88 | -0.04 | 0.08 | 0.13 | 0.0 | Detail |
| 1101 | CHOL | autosis | ZEB2 | zinc finger E-box binding homeobox 2 | 9.06 | 8.27 | -0.13 | 0.1 | 0.16 | 0.0 | Detail |
| 1102 | CHOL | autosis | ERO1A | endoplasmic reticulum oxidoreductase 1 alpha | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1103 | CHOL | autosis | BAD | BCL2 associated agonist of cell death | 10.13 | 10.3 | 0.02 | 0.62 | 0.69 | 0.0 | Detail |
| 1104 | CHOL | autosis | XIAP | X-linked inhibitor of apoptosis | 10.21 | 10.26 | 0.01 | 0.58 | 0.66 | 0.0 | Detail |
| 1105 | CHOL | autosis | ACVR1B | activin A receptor type 1B | 10.36 | 10.27 | -0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1106 | CHOL | autosis | AKR1C3 | aldo-keto reductase family 1 member C3 | 11.86 | 12.02 | 0.02 | 0.51 | 0.59 | 0.0 | Detail |
| 1107 | CHOL | autosis | BNIP3L | BCL2 interacting protein 3 like | 10.36 | 10.18 | -0.03 | 0.45 | 0.53 | 0.0 | Detail |
| 1108 | CHOL | autosis | CDH2 | cadherin 2 | 11.4 | 9.94 | -0.2 | 0.13 | 0.19 | 0.0 | Detail |
| 1109 | CHOL | autosis | CLK1 | CDC like kinase 1 | 10.76 | 10.61 | -0.02 | 0.51 | 0.59 | 0.0 | Detail |
| 1110 | CHOL | autosis | CTNND1 | catenin delta 1 | 12.34 | 12.81 | 0.05 | 0.03 | 0.06 | 0.0 | Detail |
| 1111 | CHOL | autosis | CYP24A1 | cytochrome P450 family 24 subfamily A member 1 | 0.62 | 1.6 | 1.36 | 0.14 | 0.2 | 0.0 | Detail |
| 1112 | CHOL | autosis | DCTN4 | dynactin subunit 4 | 10.23 | 10.16 | -0.01 | 0.77 | 0.83 | 0.0 | Detail |
| 1113 | CHOL | autosis | ERBB4 | erb-b2 receptor tyrosine kinase 4 | 2.03 | 3.45 | 0.76 | 0.06 | 0.1 | 0.0 | Detail |
| 1114 | CHOL | autosis | EREG | epiregulin | 3.03 | 3.34 | 0.14 | 0.65 | 0.72 | 0.0 | Detail |
| 1115 | CHOL | autosis | F11R | F11 receptor | 12.35 | 12.66 | 0.04 | 0.08 | 0.13 | 0.0 | Detail |
| 1116 | CHOL | autosis | FTH1 | ferritin heavy chain 1 | 14.86 | 14.95 | 0.01 | 0.62 | 0.69 | 0.0 | Detail |
| 1117 | CHOL | autosis | FZD7 | frizzled class receptor 7 | 6.36 | 7.41 | 0.22 | 0.12 | 0.17 | 0.0 | Detail |
| 1118 | CHOL | autosis | GFPT1 | glutamine--fructose-6-phosphate transaminase 1 | 10.74 | 11.12 | 0.05 | 0.16 | 0.23 | 0.0 | Detail |
| 1119 | CHOL | autosis | H2AX | H2A.X variant histone | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1120 | CHOL | autosis | IGF2BP3 | insulin like growth factor 2 mRNA binding protein 3 | 4.33 | 6.63 | 0.61 | 0.03 | 0.06 | 0.0 | Detail |
| 1121 | CHOL | autosis | ILRUN | inflammation and lipid regulator with UBA-like and NBR1-like domains | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1122 | CHOL | autosis | JPT2 | Jupiter microtubule associated homolog 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1123 | CHOL | autosis | KLF4 | KLF transcription factor 4 | 8.37 | 8.03 | -0.06 | 0.62 | 0.69 | 0.0 | Detail |
| 1124 | CHOL | autosis | MAP2K5 | mitogen-activated protein kinase kinase 5 | 8.14 | 8.52 | 0.07 | 0.08 | 0.12 | 0.0 | Detail |
| 1125 | CHOL | autosis | MAP3K2 | mitogen-activated protein kinase kinase kinase 2 | 8.93 | 8.46 | -0.08 | 0.36 | 0.44 | 0.0 | Detail |
| 1126 | CHOL | autosis | PLCD1 | phospholipase C delta 1 | 8.39 | 7.82 | -0.1 | 0.05 | 0.08 | 0.0 | Detail |
| 1127 | CHOL | autosis | RAB5A | RAB5A, member RAS oncogene family | 10.28 | 9.65 | -0.09 | 0.03 | 0.05 | 0.0 | Detail |
| 1128 | CHOL | autosis | RAB6A | RAB6A, member RAS oncogene family | 11.23 | 11.2 | -0.0 | 1.0 | 1.0 | 0.0 | Detail |
| 1129 | CHOL | autosis | RARB | retinoic acid receptor beta | 7.14 | 6.73 | -0.08 | 0.36 | 0.44 | 0.0 | Detail |
| 1130 | CHOL | autosis | SERPINB2 | serpin family B member 2 | 1.2 | 1.91 | 0.67 | 0.44 | 0.52 | 0.0 | Detail |
| 1131 | CHOL | autosis | SLC4A7 | solute carrier family 4 member 7 | 7.24 | 7.87 | 0.12 | 0.28 | 0.36 | 0.0 | Detail |
| 1132 | CHOL | autosis | SQSTM1 | sequestosome 1 | 13.82 | 13.79 | -0.0 | 0.74 | 0.79 | 0.0 | Detail |
| 1133 | CHOL | autosis | SRGN | serglycin | 9.78 | 9.64 | -0.02 | 0.81 | 0.86 | 0.0 | Detail |
| 1134 | CHOL | autosis | TACSTD2 | tumor associated calcium signal transducer 2 | 7.88 | 9.63 | 0.29 | 0.28 | 0.36 | 0.0 | Detail |
| 1135 | CHOL | autosis | TFPI2 | tissue factor pathway inhibitor 2 | 6.72 | 7.91 | 0.23 | 0.39 | 0.47 | 0.0 | Detail |
| 1136 | CHOL | autosis | TNS3 | tensin 3 | 11.76 | 11.51 | -0.03 | 0.24 | 0.31 | 0.0 | Detail |
| 1137 | CHOL | autosis | UBE2B | ubiquitin conjugating enzyme E2 B | 10.39 | 10.47 | 0.01 | 0.71 | 0.77 | 0.0 | Detail |
| 1138 | CHOL | autosis | UBR7 | ubiquitin protein ligase E3 component n-recognin 7 | 9.42 | 9.28 | -0.02 | 0.28 | 0.36 | 0.0 | Detail |
| 1139 | CHOL | autosis | ULK1 | unc-51 like autophagy activating kinase 1 | 10.78 | 10.81 | 0.0 | 0.58 | 0.66 | 0.0 | Detail |
| 1140 | CHOL | autosis | UTRN | utrophin | 10.16 | 10.47 | 0.04 | 0.45 | 0.53 | 0.0 | Detail |
| 1141 | CHOL | autosis | ZFX | zinc finger protein X-linked | 8.69 | 8.95 | 0.04 | 0.26 | 0.34 | 0.0 | Detail |
| 1142 | CHOL | autosis | DDIT4 | DNA damage inducible transcript 4 | 10.9 | 11.39 | 0.06 | 0.51 | 0.59 | 0.0 | Detail |
| 1143 | CHOL | autosis | MMP13 | matrix metallopeptidase 13 | 0.0 | 1.04 | inf | 0.04 | 0.08 | 0.0 | Detail |
| 1144 | CHOL | autosis | NR2F1 | nuclear receptor subfamily 2 group F member 1 | 8.6 | 7.58 | -0.18 | 0.1 | 0.16 | 0.0 | Detail |
| 1145 | CHOL | autosis | MAU2 | MAU2 sister chromatid cohesion factor | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1146 | CHOL | autosis | DAG1 | dystroglycan 1 | 11.81 | 11.57 | -0.03 | 0.18 | 0.24 | 0.0 | Detail |
| 1147 | CHOL | autosis | TSPAN31 | tetraspanin 31 | 10.11 | 10.16 | 0.01 | 0.62 | 0.69 | 0.0 | Detail |
| 1148 | CHOL | autosis | BMP4 | bone morphogenetic protein 4 | 6.32 | 7.25 | 0.2 | 0.19 | 0.26 | 0.0 | Detail |
| 1149 | CHOL | autosis | HSF2 | heat shock transcription factor 2 | 6.82 | 7.39 | 0.12 | 0.09 | 0.14 | 0.0 | Detail |
| 1150 | CHOL | autosis | ID1 | inhibitor of DNA binding 1 | 10.88 | 10.48 | -0.05 | 0.7 | 0.76 | 0.0 | Detail |
| 1151 | CHOL | autosis | LOX | lysyl oxidase | 5.61 | 6.99 | 0.32 | 0.03 | 0.06 | 0.0 | Detail |
| 1152 | CHOL | autosis | NDRG1 | N-myc downstream regulated 1 | 11.21 | 11.71 | 0.06 | 0.21 | 0.29 | 0.0 | Detail |
| 1153 | CHOL | autosis | NRG1 | neuregulin 1 | 7.62 | 5.56 | -0.45 | 0.04 | 0.07 | 0.0 | Detail |
| 1154 | CHOL | autosis | PJA2 | praja ring finger ubiquitin ligase 2 | 11.61 | 10.97 | -0.08 | 0.07 | 0.12 | 0.0 | Detail |
| 1155 | CHOL | autosis | SLC7A5 | solute carrier family 7 member 5 | 8.23 | 9.09 | 0.14 | 0.16 | 0.23 | 0.0 | Detail |
| 1156 | CHOL | autosis | STK3 | serine/threonine kinase 3 | 8.66 | 8.82 | 0.03 | 0.42 | 0.5 | 0.0 | Detail |
| 1157 | CHOL | autosis | ZNF395 | zinc finger protein 395 | 10.17 | 9.93 | -0.03 | 0.35 | 0.43 | 0.0 | Detail |
| 1158 | CHOL | autosis | AKAP9 | A-kinase anchoring protein 9 | 11.24 | 10.82 | -0.05 | 0.06 | 0.1 | 0.0 | Detail |
| 1159 | CHOL | autosis | ATF1 | activating transcription factor 1 | 8.42 | 8.52 | 0.02 | 0.28 | 0.36 | 0.0 | Detail |
| 1160 | CHOL | autosis | BMP1 | bone morphogenetic protein 1 | 10.31 | 9.98 | -0.05 | 0.21 | 0.29 | 0.0 | Detail |
| 1161 | CHOL | autosis | BTG1 | BTG anti-proliferation factor 1 | 12.24 | 12.94 | 0.08 | 0.06 | 0.1 | 0.0 | Detail |
| 1162 | CHOL | autosis | ERCC5 | ERCC excision repair 5, endonuclease | 10.65 | 10.54 | -0.02 | 0.74 | 0.79 | 0.0 | Detail |
| 1163 | CHOL | autosis | GDNF | glial cell derived neurotrophic factor | 0.66 | 1.5 | 1.18 | 0.49 | 0.57 | 0.0 | Detail |
| 1164 | CHOL | autosis | MREG | melanoregulin | 7.76 | 8.33 | 0.1 | 0.12 | 0.17 | 0.0 | Detail |
| 1165 | CHOL | autosis | NFIL3 | nuclear factor, interleukin 3 regulated | 10.41 | 9.44 | -0.14 | 0.09 | 0.14 | 0.0 | Detail |
| 1166 | CHOL | autosis | PLA2R1 | phospholipase A2 receptor 1 | 4.92 | 5.93 | 0.27 | 0.09 | 0.14 | 0.0 | Detail |
| 1167 | CHOL | autosis | RGS4 | regulator of G protein signaling 4 | 6.85 | 8.98 | 0.39 | 0.13 | 0.19 | 0.0 | Detail |
| 1168 | CHOL | autosis | RXRB | retinoid X receptor beta | 10.0 | 10.15 | 0.02 | 0.26 | 0.34 | 0.0 | Detail |
| 1169 | CHOL | autosis | STK17B | serine/threonine kinase 17b | 7.78 | 9.07 | 0.22 | 0.03 | 0.05 | 0.0 | Detail |
| 1170 | CHOL | autosis | TCF7L1 | transcription factor 7 like 1 | 8.38 | 8.44 | 0.01 | 0.81 | 0.86 | 0.0 | Detail |
| 1171 | CHOL | autosis | TSG101 | tumor susceptibility 101 | 10.33 | 10.51 | 0.02 | 0.21 | 0.29 | 0.0 | Detail |
| 1172 | CHOL | autosis | UPP1 | uridine phosphorylase 1 | 8.22 | 8.81 | 0.1 | 0.21 | 0.29 | 0.0 | Detail |
| 1173 | CHOL | autosis | WRN | WRN RecQ like helicase | 7.28 | 7.73 | 0.09 | 0.16 | 0.23 | 0.0 | Detail |
| 1174 | CHOL | autosis | CFLAR | CASP8 and FADD like apoptosis regulator | 9.89 | 10.36 | 0.07 | 0.1 | 0.16 | 0.0 | Detail |
| 1175 | CHOL | autosis | SORBS1 | sorbin and SH3 domain containing 1 | 10.94 | 9.85 | -0.15 | 0.03 | 0.05 | 0.0 | Detail |
| 1176 | CHOL | autosis | HSPA4L | heat shock protein family A (Hsp70) member 4 like | 7.57 | 7.23 | -0.07 | 0.94 | 0.95 | 0.0 | Detail |
| 1177 | CHOL | autosis | RESF1 | retroelement silencing factor 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1178 | CHOL | autosis | STK10 | serine/threonine kinase 10 | 8.78 | 9.51 | 0.11 | 0.06 | 0.1 | 0.0 | Detail |
| 1179 | CHOL | autosis | HOXA1 | homeobox A1 | 2.78 | 3.32 | 0.25 | 0.42 | 0.5 | 0.0 | Detail |
| 1180 | CHOL | autosis | LANCL1 | LanC like glutathione S-transferase 1 | 9.85 | 10.14 | 0.04 | 0.21 | 0.29 | 0.0 | Detail |
| 1181 | CHOL | autosis | CDKN2D | cyclin dependent kinase inhibitor 2D | 5.92 | 7.22 | 0.29 | 0.03 | 0.06 | 0.0 | Detail |
| 1182 | CHOL | autosis | RPA3 | replication protein A3 | 8.43 | 8.24 | -0.03 | 0.36 | 0.44 | 0.0 | Detail |
| 1183 | CHOL | autosis | TRIM25 | tripartite motif containing 25 | 9.22 | 9.87 | 0.1 | 0.07 | 0.12 | 0.0 | Detail |
| 1184 | CHOL | autosis | CD55 | CD55 molecule (Cromer blood group) | 9.2 | 9.39 | 0.03 | 0.77 | 0.83 | 0.0 | Detail |
| 1185 | CHOL | autosis | CDK10 | cyclin dependent kinase 10 | 10.92 | 10.98 | 0.01 | 0.77 | 0.83 | 0.0 | Detail |
| 1186 | CHOL | autosis | DCP1A | decapping mRNA 1A | 8.47 | 8.52 | 0.01 | 0.77 | 0.83 | 0.0 | Detail |
| 1187 | CHOL | autosis | EBAG9 | estrogen receptor binding site associated antigen 9 | 9.58 | 9.24 | -0.05 | 0.33 | 0.41 | 0.0 | Detail |
| 1188 | CHOL | autosis | GOLGA2 | golgin A2 | 10.7 | 10.78 | 0.01 | 0.86 | 0.89 | 0.0 | Detail |
| 1189 | CHOL | autosis | GSE1 | Gse1 coiled-coil protein | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1190 | CHOL | autosis | HOXA10 | homeobox A10 | 1.65 | 4.34 | 1.4 | 0.12 | 0.18 | 0.0 | Detail |
| 1191 | CHOL | autosis | HOXA9 | homeobox A9 | 0.85 | 1.82 | 1.09 | 0.19 | 0.26 | 0.0 | Detail |
| 1192 | CHOL | autosis | HSPH1 | heat shock protein family H (Hsp110) member 1 | 9.72 | 10.52 | 0.11 | 0.04 | 0.07 | 0.0 | Detail |
| 1193 | CHOL | autosis | IL24 | interleukin 24 | 1.5 | 1.78 | 0.25 | 0.96 | 0.97 | 0.0 | Detail |
| 1194 | CHOL | autosis | KANK2 | KN motif and ankyrin repeat domains 2 | 11.0 | 11.11 | 0.02 | 0.94 | 0.95 | 0.0 | Detail |
| 1195 | CHOL | autosis | KDM4A | lysine demethylase 4A | 9.93 | 10.27 | 0.05 | 0.24 | 0.31 | 0.0 | Detail |
| 1196 | CHOL | autosis | KDM6B | lysine demethylase 6B | 10.15 | 10.1 | -0.01 | 0.81 | 0.86 | 0.0 | Detail |
| 1197 | CHOL | autosis | MEF2D | myocyte enhancer factor 2D | 10.0 | 10.55 | 0.08 | 0.03 | 0.05 | 0.0 | Detail |
| 1198 | CHOL | autosis | MSX1 | msh homeobox 1 | 5.01 | 5.58 | 0.16 | 0.24 | 0.31 | 0.0 | Detail |
| 1199 | CHOL | autosis | NASP | nuclear autoantigenic sperm protein | 10.42 | 10.87 | 0.06 | 0.07 | 0.12 | 0.0 | Detail |
| 1200 | CHOL | autosis | PDCD5 | programmed cell death 5 | 9.5 | 9.83 | 0.05 | 0.1 | 0.16 | 0.0 | Detail |
| 1201 | CHOL | autosis | PDYN | prodynorphin | 0.6 | 0.17 | -1.82 | 0.85 | 0.89 | 0.0 | Detail |
| 1202 | CHOL | autosis | PHLDA1 | pleckstrin homology like domain family A member 1 | 12.02 | 10.52 | -0.19 | 0.05 | 0.08 | 0.0 | Detail |
| 1203 | CHOL | autosis | PTBP3 | polypyrimidine tract binding protein 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1204 | CHOL | autosis | RPS6KB2 | ribosomal protein S6 kinase B2 | 9.93 | 10.05 | 0.02 | 0.86 | 0.89 | 0.0 | Detail |
| 1205 | CHOL | autosis | SIX1 | SIX homeobox 1 | 1.83 | 2.57 | 0.49 | 0.33 | 0.42 | 0.0 | Detail |
| 1206 | CHOL | autosis | SRSF7 | serine and arginine rich splicing factor 7 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1207 | CHOL | autosis | SYF2 | SYF2 pre-mRNA splicing factor | 9.82 | 9.93 | 0.02 | 0.58 | 0.66 | 0.0 | Detail |
| 1208 | CHOL | autosis | TCEA1 | transcription elongation factor A1 | 10.3 | 9.99 | -0.04 | 0.1 | 0.16 | 0.0 | Detail |
| 1209 | CHOL | autosis | TERF1 | telomeric repeat binding factor 1 | 8.81 | 8.71 | -0.02 | 0.77 | 0.83 | 0.0 | Detail |
| 1210 | CHOL | autosis | TP53BP2 | tumor protein p53 binding protein 2 | 9.81 | 10.32 | 0.07 | 0.1 | 0.16 | 0.0 | Detail |
| 1211 | CHOL | autosis | TTC33 | tetratricopeptide repeat domain 33 | 8.84 | 8.46 | -0.06 | 0.09 | 0.14 | 0.0 | Detail |
| 1212 | CHOL | autosis | ZFP36 | ZFP36 ring finger protein | 12.84 | 11.88 | -0.11 | 0.09 | 0.14 | 0.0 | Detail |
| 1213 | CHOL | autosis | HCRT | hypocretin neuropeptide precursor | 0.0 | 0.31 | inf | 0.14 | 0.2 | 0.0 | Detail |
| 1214 | CHOL | autosis | RGS2 | regulator of G protein signaling 2 | 8.44 | 9.48 | 0.17 | 0.16 | 0.23 | 0.0 | Detail |
| 1215 | CHOL | autosis | CLDN11 | claudin 11 | 6.62 | 6.75 | 0.03 | 0.81 | 0.86 | 0.0 | Detail |
| 1216 | CHOL | autosis | HMBOX1 | homeobox containing 1 | 5.63 | 5.75 | 0.03 | 0.77 | 0.83 | 0.0 | Detail |
| 1217 | CHOL | autosis | ITGB6 | integrin subunit beta 6 | 3.4 | 6.35 | 0.9 | 0.04 | 0.07 | 0.0 | Detail |
| 1218 | CHOL | autosis | RNF6 | ring finger protein 6 | 8.93 | 9.0 | 0.01 | 0.55 | 0.63 | 0.0 | Detail |
| 1219 | CHOL | autosis | SOCS4 | suppressor of cytokine signaling 4 | 8.23 | 8.58 | 0.06 | 0.07 | 0.12 | 0.0 | Detail |
| 1220 | CHOL | autosis | TRAF6 | TNF receptor associated factor 6 | 7.24 | 6.86 | -0.08 | 0.26 | 0.34 | 0.0 | Detail |
| 1221 | CHOL | autosis | ADAMTS5 | ADAM metallopeptidase with thrombospondin type 1 motif 5 | 6.24 | 6.48 | 0.05 | 0.66 | 0.72 | 0.0 | Detail |
| 1222 | CHOL | autosis | AKAP12 | A-kinase anchoring protein 12 | 9.51 | 9.5 | -0.0 | 0.98 | 0.98 | 0.0 | Detail |
| 1223 | CHOL | autosis | AREG | amphiregulin | 3.83 | 6.01 | 0.65 | 0.04 | 0.07 | 0.0 | Detail |
| 1224 | CHOL | autosis | ARHGDIB | Rho GDP dissociation inhibitor beta | 10.5 | 10.84 | 0.05 | 0.86 | 0.89 | 0.0 | Detail |
| 1225 | CHOL | autosis | ATP4B | ATPase H+/K+ transporting subunit beta | 0.0 | 0.14 | inf | 0.31 | 0.4 | 0.0 | Detail |
| 1226 | CHOL | autosis | CCN3 | cellular communication network factor 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1227 | CHOL | autosis | CLCN3 | chloride voltage-gated channel 3 | 9.37 | 9.84 | 0.07 | 0.15 | 0.22 | 0.0 | Detail |
| 1228 | CHOL | autosis | CNN2 | calponin 2 | 10.46 | 11.34 | 0.12 | 0.03 | 0.05 | 0.0 | Detail |
| 1229 | CHOL | autosis | CPT1A | carnitine palmitoyltransferase 1A | 12.14 | 11.65 | -0.06 | 0.16 | 0.23 | 0.0 | Detail |
| 1230 | CHOL | autosis | CREBZF | CREB/ATF bZIP transcription factor | 10.06 | 10.25 | 0.03 | 0.21 | 0.29 | 0.0 | Detail |
| 1231 | CHOL | autosis | CXXC1 | CXXC finger protein 1 | 9.56 | 10.03 | 0.07 | 0.13 | 0.19 | 0.0 | Detail |
| 1232 | CHOL | autosis | DHRS11 | dehydrogenase/reductase 11 | 8.08 | 7.52 | -0.1 | 0.19 | 0.26 | 0.0 | Detail |
| 1233 | CHOL | autosis | DLEU1 | deleted in lymphocytic leukemia 1 | 6.64 | 6.38 | -0.06 | 0.26 | 0.34 | 0.0 | Detail |
| 1234 | CHOL | autosis | EFNB1 | ephrin B1 | 9.24 | 9.65 | 0.06 | 0.86 | 0.89 | 0.0 | Detail |
| 1235 | CHOL | autosis | EIF3A | eukaryotic translation initiation factor 3 subunit A | 12.21 | 12.25 | 0.0 | 0.55 | 0.63 | 0.0 | Detail |
| 1236 | CHOL | autosis | EIF6 | eukaryotic translation initiation factor 6 | 11.68 | 11.79 | 0.01 | 0.58 | 0.66 | 0.0 | Detail |
| 1237 | CHOL | autosis | FKTN | fukutin | 8.0 | 8.22 | 0.04 | 0.36 | 0.44 | 0.0 | Detail |
| 1238 | CHOL | autosis | GAREM1 | GRB2 associated regulator of MAPK1 subtype 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1239 | CHOL | autosis | GPI | glucose-6-phosphate isomerase | 12.91 | 12.96 | 0.01 | 1.0 | 1.0 | 0.0 | Detail |
| 1240 | CHOL | autosis | GPRC5A | G protein-coupled receptor class C group 5 member A | 3.65 | 6.28 | 0.78 | 0.03 | 0.05 | 0.0 | Detail |
| 1241 | CHOL | autosis | HINT1 | histidine triad nucleotide binding protein 1 | 12.76 | 12.33 | -0.05 | 0.1 | 0.16 | 0.0 | Detail |
| 1242 | CHOL | autosis | HOXB7 | homeobox B7 | 3.51 | 5.69 | 0.7 | 0.04 | 0.06 | 0.0 | Detail |
| 1243 | CHOL | autosis | HSPA13 | heat shock protein family A (Hsp70) member 13 | 8.4 | 8.97 | 0.09 | 0.05 | 0.09 | 0.0 | Detail |
| 1244 | CHOL | autosis | KLF6 | KLF transcription factor 6 | 12.46 | 12.23 | -0.03 | 0.45 | 0.53 | 0.0 | Detail |
| 1245 | CHOL | autosis | KMT2B | lysine methyltransferase 2B | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1246 | CHOL | autosis | KMT2D | lysine methyltransferase 2D | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1247 | CHOL | autosis | LAMTOR3 | late endosomal/lysosomal adaptor, MAPK and MTOR activator 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1248 | CHOL | autosis | LYN | LYN proto-oncogene, Src family tyrosine kinase | 9.26 | 9.54 | 0.04 | 0.28 | 0.36 | 0.0 | Detail |
| 1249 | CHOL | autosis | MELTF | melanotransferrin | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1250 | CHOL | autosis | MPC1 | mitochondrial pyruvate carrier 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1251 | CHOL | autosis | NCF2 | neutrophil cytosolic factor 2 | 6.86 | 8.66 | 0.34 | 0.03 | 0.05 | 0.0 | Detail |
| 1252 | CHOL | autosis | PCDH7 | protocadherin 7 | 4.99 | 6.46 | 0.37 | 0.18 | 0.24 | 0.0 | Detail |
| 1253 | CHOL | autosis | PON2 | paraoxonase 2 | 11.41 | 11.3 | -0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1254 | CHOL | autosis | PTX3 | pentraxin 3 | 3.62 | 4.52 | 0.32 | 0.24 | 0.31 | 0.0 | Detail |
| 1255 | CHOL | autosis | RABEP1 | rabaptin, RAB GTPase binding effector protein 1 | 10.28 | 9.8 | -0.07 | 0.05 | 0.08 | 0.0 | Detail |
| 1256 | CHOL | autosis | RABGEF1 | RAB guanine nucleotide exchange factor 1 | 9.52 | 9.47 | -0.01 | 0.94 | 0.95 | 0.0 | Detail |
| 1257 | CHOL | autosis | SGCB | sarcoglycan beta | 9.46 | 9.72 | 0.04 | 0.48 | 0.56 | 0.0 | Detail |
| 1258 | CHOL | autosis | SLC16A7 | solute carrier family 16 member 7 | 5.44 | 6.27 | 0.2 | 0.05 | 0.09 | 0.0 | Detail |
| 1259 | CHOL | autosis | TBL1X | transducin beta like 1 X-linked | 10.19 | 9.76 | -0.06 | 0.26 | 0.34 | 0.0 | Detail |
| 1260 | CHOL | autosis | TCF7L2 | transcription factor 7 like 2 | 9.55 | 9.17 | -0.06 | 0.12 | 0.17 | 0.0 | Detail |
| 1261 | CHOL | autosis | TGM2 | transglutaminase 2 | 12.95 | 12.73 | -0.02 | 0.81 | 0.86 | 0.0 | Detail |
| 1262 | CHOL | autosis | TRIM14 | tripartite motif containing 14 | 10.24 | 9.75 | -0.07 | 0.16 | 0.23 | 0.0 | Detail |
| 1263 | CHOL | autosis | TRIM23 | tripartite motif containing 23 | 8.45 | 8.08 | -0.06 | 0.18 | 0.24 | 0.0 | Detail |
| 1264 | CHOL | autosis | TSPAN1 | tetraspanin 1 | 6.49 | 7.58 | 0.22 | 0.14 | 0.21 | 0.0 | Detail |
| 1265 | CHOL | autosis | UBR2 | ubiquitin protein ligase E3 component n-recognin 2 | 9.61 | 9.22 | -0.06 | 0.09 | 0.14 | 0.0 | Detail |
| 1266 | CHOL | autosis | UFM1 | ubiquitin fold modifier 1 | 10.76 | 10.42 | -0.05 | 0.14 | 0.21 | 0.0 | Detail |
| 1267 | CHOL | autosis | YBX3 | Y-box binding protein 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1268 | CHOL | autosis | ZFP36L2 | ZFP36 ring finger protein like 2 | 11.77 | 12.25 | 0.06 | 0.13 | 0.19 | 0.0 | Detail |
| 1269 | CHOL | autosis | CHKA | choline kinase alpha | 9.35 | 9.74 | 0.06 | 0.42 | 0.5 | 0.0 | Detail |
| 1270 | CHOL | autosis | ELP5 | elongator acetyltransferase complex subunit 5 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1271 | CHOL | autosis | MAP2K4 | mitogen-activated protein kinase kinase 4 | 9.7 | 9.31 | -0.06 | 0.05 | 0.08 | 0.0 | Detail |
| 1272 | CHOL | autosis | HIPK3 | homeodomain interacting protein kinase 3 | 9.19 | 8.36 | -0.14 | 0.7 | 0.76 | 0.0 | Detail |
| 1273 | CHOL | autosis | NLK | nemo like kinase | 8.77 | 8.96 | 0.03 | 0.19 | 0.26 | 0.0 | Detail |
| 1274 | CHOL | autosis | ACVR2A | activin A receptor type 2A | 8.58 | 8.44 | -0.02 | 0.9 | 0.92 | 0.0 | Detail |
| 1275 | CHOL | autosis | MFHAS1 | multifunctional ROCO family signaling regulator 1 | 7.17 | 7.93 | 0.15 | 0.13 | 0.19 | 0.0 | Detail |
| 1276 | CHOL | autosis | SNIP1 | Smad nuclear interacting protein 1 | 8.13 | 8.04 | -0.02 | 0.7 | 0.76 | 0.0 | Detail |
| 1277 | CHOL | autosis | ITGA7 | integrin subunit alpha 7 | 8.02 | 8.39 | 0.06 | 0.62 | 0.69 | 0.0 | Detail |
| 1278 | CHOL | autosis | PGPEP1 | pyroglutamyl-peptidase I | 10.56 | 10.29 | -0.04 | 0.39 | 0.47 | 0.0 | Detail |
| 1279 | CHOL | autosis | ATP2A2 | ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 2 | 12.12 | 12.42 | 0.04 | 0.21 | 0.29 | 0.0 | Detail |
| 1280 | CHOL | autosis | CCNA1 | cyclin A1 | 0.67 | 1.56 | 1.22 | 0.22 | 0.3 | 0.0 | Detail |
| 1281 | CHOL | autosis | CCNL1 | cyclin L1 | 10.81 | 10.45 | -0.05 | 0.08 | 0.13 | 0.0 | Detail |
| 1282 | CHOL | autosis | FANCL | FA complementation group L | 8.16 | 8.38 | 0.04 | 0.31 | 0.39 | 0.0 | Detail |
| 1283 | CHOL | autosis | GCC1 | GRIP and coiled-coil domain containing 1 | 9.28 | 9.25 | -0.01 | 0.94 | 0.95 | 0.0 | Detail |
| 1284 | CHOL | autosis | HEY1 | hes related family bHLH transcription factor with YRPW motif 1 | 6.17 | 7.04 | 0.19 | 0.08 | 0.13 | 0.0 | Detail |
| 1285 | CHOL | autosis | IFI16 | interferon gamma inducible protein 16 | 9.37 | 10.47 | 0.16 | 0.04 | 0.07 | 0.0 | Detail |
| 1286 | CHOL | autosis | NAB2 | NGFI-A binding protein 2 | 9.98 | 9.57 | -0.06 | 0.19 | 0.26 | 0.0 | Detail |
| 1287 | CHOL | autosis | NIPBL | NIPBL cohesin loading factor | 10.1 | 10.49 | 0.05 | 0.04 | 0.06 | 0.0 | Detail |
| 1288 | CHOL | autosis | RAD17 | RAD17 checkpoint clamp loader component | 8.98 | 9.19 | 0.03 | 0.05 | 0.08 | 0.0 | Detail |
| 1289 | CHOL | autosis | SLC17A5 | solute carrier family 17 member 5 | 9.43 | 8.84 | -0.09 | 0.18 | 0.24 | 0.0 | Detail |
| 1290 | CHOL | autosis | TRIM33 | tripartite motif containing 33 | 9.48 | 9.79 | 0.05 | 0.12 | 0.17 | 0.0 | Detail |
| 1291 | CHOL | autosis | NMU | neuromedin U | 0.0 | 0.89 | inf | 0.11 | 0.17 | 0.0 | Detail |
| 1292 | CHOL | autosis | DST | dystonin | 12.03 | 12.33 | 0.04 | 0.19 | 0.26 | 0.0 | Detail |
| 1293 | CHOL | autosis | AFF4 | ALF transcription elongation factor 4 | 11.61 | 11.1 | -0.07 | 0.13 | 0.19 | 0.0 | Detail |
| 1294 | CHOL | autosis | ATP2A3 | ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 3 | 8.21 | 9.22 | 0.17 | 0.12 | 0.17 | 0.0 | Detail |
| 1295 | CHOL | autosis | ATP5MC1 | ATP synthase membrane subunit c locus 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1296 | CHOL | autosis | CHAC1 | ChaC glutathione specific gamma-glutamylcyclotransferase 1 | 6.02 | 6.33 | 0.07 | 0.42 | 0.5 | 0.0 | Detail |
| 1297 | CHOL | autosis | CUL3 | cullin 3 | 10.49 | 10.57 | 0.01 | 0.39 | 0.47 | 0.0 | Detail |
| 1298 | CHOL | autosis | GNA13 | G protein subunit alpha 13 | 10.3 | 10.73 | 0.06 | 0.07 | 0.12 | 0.0 | Detail |
| 1299 | CHOL | autosis | COL17A1 | collagen type XVII alpha 1 chain | 1.12 | 3.59 | 1.68 | 0.04 | 0.07 | 0.0 | Detail |
| 1300 | CHOL | autosis | GEMIN4 | gem nuclear organelle associated protein 4 | 8.58 | 9.04 | 0.08 | 0.05 | 0.09 | 0.0 | Detail |
| 1301 | CHOL | autosis | RMDN3 | regulator of microtubule dynamics 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1302 | CHOL | autosis | IRF4 | interferon regulatory factor 4 | 4.52 | 5.03 | 0.15 | 0.66 | 0.72 | 0.0 | Detail |
| 1303 | CHOL | autosis | EML4 | EMAP like 4 | 11.14 | 11.38 | 0.03 | 0.26 | 0.34 | 0.0 | Detail |
| 1304 | CHOL | autosis | LIPA | lipase A, lysosomal acid type | 11.62 | 11.0 | -0.08 | 0.08 | 0.13 | 0.0 | Detail |
| 1305 | CHOL | autosis | BTN3A3 | butyrophilin subfamily 3 member A3 | 8.95 | 8.95 | -0.0 | 0.7 | 0.76 | 0.0 | Detail |
| 1306 | CHOL | autosis | LCK | LCK proto-oncogene, Src family tyrosine kinase | 6.75 | 6.68 | -0.02 | 0.81 | 0.86 | 0.0 | Detail |
| 1307 | CHOL | autosis | MRTFA | myocardin related transcription factor A | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1308 | CHOL | autosis | NFIX | nuclear factor I X | 10.82 | 10.94 | 0.02 | 0.81 | 0.86 | 0.0 | Detail |
| 1309 | CHOL | autosis | IST1 | IST1 factor associated with ESCRT-III | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1310 | CHOL | autosis | SPSB3 | splA/ryanodine receptor domain and SOCS box containing 3 | 10.75 | 10.31 | -0.06 | 0.07 | 0.12 | 0.0 | Detail |
| 1311 | CHOL | autosis | ACP6 | acid phosphatase 6, lysophosphatidic | 8.44 | 8.03 | -0.07 | 0.28 | 0.36 | 0.0 | Detail |
| 1312 | CHOL | autosis | BTF3P11 | basic transcription factor 3 pseudogene 11 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1313 | CHOL | autosis | CDKL3 | cyclin dependent kinase like 3 | 4.28 | 4.79 | 0.16 | 0.12 | 0.17 | 0.0 | Detail |
| 1314 | CHOL | autosis | CNPY2 | canopy FGF signaling regulator 2 | 10.86 | 10.93 | 0.01 | 0.77 | 0.83 | 0.0 | Detail |
| 1315 | CHOL | autosis | DEPTOR | DEP domain containing MTOR interacting protein | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1316 | CHOL | autosis | IL3 | interleukin 3 | 0 | 0 | NA | NA | NA | 0 | Detail |
| 1317 | CHOL | autosis | NUMB | NUMB endocytic adaptor protein | 10.71 | 10.28 | -0.06 | 0.05 | 0.08 | 0.0 | Detail |
| 1318 | CHOL | autosis | POLM | DNA polymerase mu | 8.57 | 9.02 | 0.07 | 0.05 | 0.08 | 0.0 | Detail |
| 1319 | CHOL | autosis | PTRH2 | peptidyl-tRNA hydrolase 2 | 8.64 | 9.0 | 0.06 | 0.07 | 0.12 | 0.0 | Detail |
| 1320 | CHOL | autosis | RRAS2 | RAS related 2 | 9.43 | 9.28 | -0.02 | 0.86 | 0.89 | 0.0 | Detail |
| 1321 | CHOL | autosis | TSPAN4 | tetraspanin 4 | 10.01 | 10.32 | 0.04 | 0.66 | 0.72 | 0.0 | Detail |
| 1322 | CHOL | autosis | ZC3H12A | zinc finger CCCH-type containing 12A | 8.51 | 9.32 | 0.13 | 0.08 | 0.13 | 0.0 | Detail |
| 1323 | CHOL | autosis | LYPD3 | LY6/PLAUR domain containing 3 | 3.91 | 5.09 | 0.38 | 0.03 | 0.06 | 0.0 | Detail |
| 1324 | CHOL | autosis | SENP5 | SUMO specific peptidase 5 | 9.26 | 9.3 | 0.01 | 0.7 | 0.76 | 0.0 | Detail |
| 1325 | CHOL | autosis | SZRD1 | SUZ RNA binding domain containing 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1326 | CHOL | autosis | WDR77 | WD repeat domain 77 | 9.2 | 9.64 | 0.07 | 0.18 | 0.24 | 0.0 | Detail |
| 1327 | CHOL | autosis | RPS6KA5 | ribosomal protein S6 kinase A5 | 5.54 | 6.01 | 0.12 | 0.21 | 0.29 | 0.0 | Detail |
| 1328 | CHOL | autosis | ARHGDIA | Rho GDP dissociation inhibitor alpha | 12.63 | 13.03 | 0.04 | 0.04 | 0.07 | 0.0 | Detail |
| 1329 | CHOL | autosis | FERMT1 | FERM domain containing kindlin 1 | 5.2 | 7.44 | 0.52 | 0.03 | 0.05 | 0.0 | Detail |
| 1330 | CHOL | autosis | IRS2 | insulin receptor substrate 2 | 11.4 | 10.48 | -0.12 | 0.16 | 0.23 | 0.0 | Detail |
| 1331 | CHOL | autosis | PGP | phosphoglycolate phosphatase | 8.35 | 8.86 | 0.09 | 0.04 | 0.07 | 0.0 | Detail |
| 1332 | CHOL | autosis | PMS1 | PMS1 homolog 1, mismatch repair system component | 7.74 | 7.98 | 0.04 | 0.26 | 0.34 | 0.0 | Detail |
| 1333 | CHOL | autosis | PUDP | pseudouridine 5'-phosphatase | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1334 | CHOL | autosis | RGL1 | ral guanine nucleotide dissociation stimulator like 1 | 8.91 | 8.55 | -0.06 | 0.51 | 0.59 | 0.0 | Detail |
| 1335 | CHOL | autosis | SBNO1 | strawberry notch homolog 1 | 7.79 | 7.7 | -0.02 | 0.51 | 0.59 | 0.0 | Detail |
| 1336 | CHOL | autosis | SKIL | SKI like proto-oncogene | 8.25 | 8.67 | 0.07 | 0.18 | 0.24 | 0.0 | Detail |
| 1337 | CHOL | autosis | SLK | STE20 like kinase | 9.95 | 10.22 | 0.04 | 0.18 | 0.24 | 0.0 | Detail |
| 1338 | CHOL | autosis | SOS2 | SOS Ras/Rho guanine nucleotide exchange factor 2 | 9.44 | 9.34 | -0.01 | 0.66 | 0.72 | 0.0 | Detail |
| 1339 | CHOL | autosis | XBP1P1 | X-box binding protein 1 pseudogene 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1340 | CHOL | autosis | PNN | pinin, desmosome associated protein | 10.79 | 10.98 | 0.03 | 0.39 | 0.47 | 0.0 | Detail |
| 1341 | CHOL | autosis | RLF | RLF zinc finger | 8.6 | 8.53 | -0.01 | 0.86 | 0.89 | 0.0 | Detail |
| 1342 | CHOL | autosis | TTBK2 | tau tubulin kinase 2 | 5.39 | 5.26 | -0.04 | 0.9 | 0.92 | 0.0 | Detail |
| 1343 | CHOL | autosis | ATP1A2 | ATPase Na+/K+ transporting subunit alpha 2 | 5.11 | 4.37 | -0.23 | 0.39 | 0.47 | 0.0 | Detail |
| 1344 | CHOL | autosis | PPP3CA | protein phosphatase 3 catalytic subunit alpha | 9.48 | 9.45 | -0.01 | 0.58 | 0.66 | 0.0 | Detail |
| 1345 | CHOL | autosis | SLCO4A1 | solute carrier organic anion transporter family member 4A1 | 4.49 | 6.29 | 0.48 | 0.09 | 0.14 | 0.0 | Detail |
| 1346 | CHOL | autosis | TCEAL9 | transcription elongation factor A like 9 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1347 | CHOL | autosis | NAV3 | neuron navigator 3 | 5.71 | 5.37 | -0.09 | 0.66 | 0.72 | 0.0 | Detail |
| 1348 | CHOL | autosis | FPGT | fucose-1-phosphate guanylyltransferase | 8.55 | 8.65 | 0.02 | 0.33 | 0.41 | 0.0 | Detail |
| 1349 | CHOL | autosis | JADE1 | jade family PHD finger 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1350 | CHOL | autosis | RTL10 | retrotransposon Gag like 10 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1351 | CHOL | autosis | SLC25A11 | solute carrier family 25 member 11 | 10.86 | 10.36 | -0.07 | 0.03 | 0.06 | 0.0 | Detail |
| 1352 | CHOL | autosis | WTAP | WT1 associated protein | 9.96 | 10.23 | 0.04 | 0.28 | 0.36 | 0.0 | Detail |
| 1353 | CHOL | autosis | COL3A1 | collagen type III alpha 1 chain | 12.99 | 13.98 | 0.11 | 0.04 | 0.07 | 0.0 | Detail |
| 1354 | CHOL | autosis | DNAJB1 | DnaJ heat shock protein family (Hsp40) member B1 | 11.02 | 11.42 | 0.05 | 0.21 | 0.29 | 0.0 | Detail |
| 1355 | CHOL | autosis | HILPDA | hypoxia inducible lipid droplet associated | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1356 | CHOL | autosis | MBNL2 | muscleblind like splicing regulator 2 | 11.2 | 10.49 | -0.09 | 0.04 | 0.07 | 0.0 | Detail |
| 1357 | CHOL | autosis | OPTN | optineurin | 11.5 | 11.54 | 0.01 | 0.66 | 0.72 | 0.0 | Detail |
| 1358 | CHOL | autosis | PDE10A | phosphodiesterase 10A | 4.99 | 6.25 | 0.33 | 0.12 | 0.17 | 0.0 | Detail |
| 1359 | CHOL | autosis | PDZD2 | PDZ domain containing 2 | 5.57 | 5.81 | 0.06 | 0.81 | 0.86 | 0.0 | Detail |
| 1360 | CHOL | autosis | PARP2 | poly(ADP-ribose) polymerase 2 | 8.21 | 8.44 | 0.04 | 0.14 | 0.21 | 0.0 | Detail |
| 1361 | CHOL | autosis | EIF1 | eukaryotic translation initiation factor 1 | 13.45 | 13.77 | 0.03 | 0.07 | 0.12 | 0.0 | Detail |
| 1362 | CHOL | autosis | APPBP2 | amyloid beta precursor protein binding protein 2 | 9.22 | 9.54 | 0.05 | 0.06 | 0.1 | 0.0 | Detail |
| 1363 | CHOL | autosis | ARFGAP3 | ADP ribosylation factor GTPase activating protein 3 | 10.34 | 10.38 | 0.01 | 0.66 | 0.72 | 0.0 | Detail |
| 1364 | CHOL | autosis | ARL8B | ADP ribosylation factor like GTPase 8B | 10.54 | 10.16 | -0.05 | 0.04 | 0.06 | 0.0 | Detail |
| 1365 | CHOL | autosis | ATRIP | ATR interacting protein | 7.17 | 7.22 | 0.01 | 0.94 | 0.95 | 0.0 | Detail |
| 1366 | CHOL | autosis | BAG5 | BAG cochaperone 5 | 9.35 | 9.66 | 0.05 | 0.13 | 0.19 | 0.0 | Detail |
| 1367 | CHOL | autosis | CCNK | cyclin K | 9.13 | 9.2 | 0.01 | 0.98 | 0.98 | 0.0 | Detail |
| 1368 | CHOL | autosis | ELL | elongation factor for RNA polymerase II | 8.9 | 8.95 | 0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1369 | CHOL | autosis | HBP1 | HMG-box transcription factor 1 | 10.93 | 10.55 | -0.05 | 0.04 | 0.07 | 0.0 | Detail |
| 1370 | CHOL | autosis | HIVEP1 | HIVEP zinc finger 1 | 9.57 | 8.81 | -0.12 | 0.03 | 0.06 | 0.0 | Detail |
| 1371 | CHOL | autosis | LCOR | ligand dependent nuclear receptor corepressor | 5.63 | 5.43 | -0.05 | 0.94 | 0.95 | 0.0 | Detail |
| 1372 | CHOL | autosis | PPP1R8 | protein phosphatase 1 regulatory subunit 8 | 9.23 | 9.28 | 0.01 | 0.66 | 0.72 | 0.0 | Detail |
| 1373 | CHOL | autosis | PPP2R2A | protein phosphatase 2 regulatory subunit Balpha | 9.5 | 9.51 | 0.0 | 0.9 | 0.92 | 0.0 | Detail |
| 1374 | CHOL | autosis | PSG1 | pregnancy specific beta-1-glycoprotein 1 | 0.0 | 0.76 | inf | 0.06 | 0.09 | 0.0 | Detail |
| 1375 | CHOL | autosis | PTPRN2 | protein tyrosine phosphatase receptor type N2 | 6.91 | 7.0 | 0.02 | 0.86 | 0.89 | 0.0 | Detail |
| 1376 | CHOL | autosis | RB1CC1 | RB1 inducible coiled-coil 1 | 10.21 | 10.05 | -0.02 | 0.74 | 0.79 | 0.0 | Detail |
| 1377 | CHOL | autosis | RBM25 | RNA binding motif protein 25 | 10.18 | 10.3 | 0.02 | 0.74 | 0.79 | 0.0 | Detail |
| 1378 | CHOL | autosis | SCAND1 | SCAN domain containing 1 | 10.78 | 10.39 | -0.05 | 0.09 | 0.14 | 0.0 | Detail |
| 1379 | CHOL | autosis | TTLL12 | tubulin tyrosine ligase like 12 | 10.29 | 10.57 | 0.04 | 0.26 | 0.34 | 0.0 | Detail |
| 1380 | CHOL | autosis | WDR19 | WD repeat domain 19 | 8.66 | 8.53 | -0.02 | 0.86 | 0.89 | 0.0 | Detail |
| 1381 | CHOL | autosis | ZBTB18 | zinc finger and BTB domain containing 18 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1382 | CHOL | autosis | B3GNT2 | UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 2 | 9.17 | 9.14 | -0.01 | 0.94 | 0.95 | 0.0 | Detail |
| 1383 | CHOL | autosis | SLC18A2 | solute carrier family 18 member A2 | 2.2 | 2.62 | 0.26 | 0.83 | 0.87 | 0.0 | Detail |
| 1384 | CHOL | autosis | BACH1 | BTB domain and CNC homolog 1 | 9.7 | 9.41 | -0.04 | 0.42 | 0.5 | 0.0 | Detail |
| 1385 | CHOL | autosis | BAZ2B | bromodomain adjacent to zinc finger domain 2B | 9.14 | 9.24 | 0.02 | 0.39 | 0.47 | 0.0 | Detail |
| 1386 | CHOL | autosis | EFEMP2 | EGF containing fibulin extracellular matrix protein 2 | 8.53 | 8.99 | 0.08 | 0.36 | 0.44 | 0.0 | Detail |
| 1387 | CHOL | autosis | ELF1 | E74 like ETS transcription factor 1 | 10.06 | 10.06 | 0.0 | 0.81 | 0.86 | 0.0 | Detail |
| 1388 | CHOL | autosis | ETV3 | ETS variant transcription factor 3 | 5.56 | 6.19 | 0.16 | 0.21 | 0.29 | 0.0 | Detail |
| 1389 | CHOL | autosis | FOXD1 | forkhead box D1 | 1.27 | 2.92 | 1.2 | 0.05 | 0.09 | 0.0 | Detail |
| 1390 | CHOL | autosis | FRS2 | fibroblast growth factor receptor substrate 2 | 8.81 | 8.82 | 0.0 | 0.86 | 0.89 | 0.0 | Detail |
| 1391 | CHOL | autosis | GLIPR1 | GLI pathogenesis related 1 | 7.62 | 8.14 | 0.09 | 0.42 | 0.5 | 0.0 | Detail |
| 1392 | CHOL | autosis | HSPA6 | heat shock protein family A (Hsp70) member 6 | 6.15 | 7.13 | 0.21 | 0.13 | 0.19 | 0.0 | Detail |
| 1393 | CHOL | autosis | HSPBAP1 | HSPB1 associated protein 1 | 6.92 | 7.61 | 0.14 | 0.06 | 0.1 | 0.0 | Detail |
| 1394 | CHOL | autosis | IFIT5 | interferon induced protein with tetratricopeptide repeats 5 | 8.9 | 8.91 | 0.0 | 0.94 | 0.95 | 0.0 | Detail |
| 1395 | CHOL | autosis | JMJD1C | jumonji domain containing 1C | 10.48 | 10.18 | -0.04 | 0.31 | 0.39 | 0.0 | Detail |
| 1396 | CHOL | autosis | KCNA5 | potassium voltage-gated channel subfamily A member 5 | 2.86 | 4.11 | 0.52 | 0.12 | 0.17 | 0.0 | Detail |
| 1397 | CHOL | autosis | LCP1 | lymphocyte cytosolic protein 1 | 10.77 | 10.85 | 0.01 | 0.98 | 0.98 | 0.0 | Detail |
| 1398 | CHOL | autosis | MLPH | melanophilin | 9.53 | 8.79 | -0.12 | 0.7 | 0.76 | 0.0 | Detail |
| 1399 | CHOL | autosis | NCOR1 | nuclear receptor corepressor 1 | 10.6 | 10.66 | 0.01 | 0.86 | 0.89 | 0.0 | Detail |
| 1400 | CHOL | autosis | NT5C3A | 5'-nucleotidase, cytosolic IIIA | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1401 | CHOL | autosis | PLEKHF1 | pleckstrin homology and FYVE domain containing 1 | 8.33 | 7.26 | -0.2 | 0.06 | 0.1 | 0.0 | Detail |
| 1402 | CHOL | autosis | POP5 | POP5 homolog, ribonuclease P/MRP subunit | 8.7 | 8.57 | -0.02 | 0.48 | 0.56 | 0.0 | Detail |
| 1403 | CHOL | autosis | PSG7 | pregnancy specific beta-1-glycoprotein 7 | 0.21 | 0.38 | 0.83 | 1.0 | 1.0 | 0.0 | Detail |
| 1404 | CHOL | autosis | PXMP4 | peroxisomal membrane protein 4 | 9.0 | 8.65 | -0.06 | 0.05 | 0.08 | 0.0 | Detail |
| 1405 | CHOL | autosis | RIDA | reactive intermediate imine deaminase A homolog | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1406 | CHOL | autosis | SENP6 | SUMO specific peptidase 6 | 10.14 | 10.06 | -0.01 | 0.7 | 0.76 | 0.0 | Detail |
| 1407 | CHOL | autosis | SERPINA2 | serpin family A member 2 (gene/pseudogene) | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1408 | CHOL | autosis | SETD5 | SET domain containing 5 | 10.34 | 10.48 | 0.02 | 0.31 | 0.39 | 0.0 | Detail |
| 1409 | CHOL | autosis | SLC1A4 | solute carrier family 1 member 4 | 8.43 | 8.99 | 0.09 | 0.12 | 0.17 | 0.0 | Detail |
| 1410 | CHOL | autosis | TMF1 | TATA element modulatory factor 1 | 9.4 | 9.25 | -0.02 | 0.86 | 0.89 | 0.0 | Detail |
| 1411 | CHOL | autosis | TSPYL2 | TSPY like 2 | 8.56 | 9.08 | 0.09 | 0.42 | 0.5 | 0.0 | Detail |
| 1412 | CHOL | autosis | WDR48 | WD repeat domain 48 | 9.52 | 9.31 | -0.03 | 0.16 | 0.23 | 0.0 | Detail |
| 1413 | CHOL | autosis | CBLL1 | Cbl proto-oncogene like 1 | 8.7 | 9.13 | 0.07 | 0.05 | 0.09 | 0.0 | Detail |
| 1414 | CHOL | autosis | OGA | O-GlcNAcase | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1415 | CHOL | autosis | HMBS | hydroxymethylbilane synthase | 8.98 | 8.95 | -0.01 | 0.77 | 0.83 | 0.0 | Detail |
| 1416 | CHOL | autosis | HUS1 | HUS1 checkpoint clamp component | 8.51 | 8.18 | -0.06 | 0.16 | 0.23 | 0.0 | Detail |
| 1417 | CHOL | autosis | BEX3 | brain expressed X-linked 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1418 | CHOL | autosis | MAP3K14 | mitogen-activated protein kinase kinase kinase 14 | 9.08 | 9.75 | 0.1 | 0.16 | 0.23 | 0.0 | Detail |
| 1419 | CHOL | autosis | CCDC68 | coiled-coil domain containing 68 | 8.07 | 7.81 | -0.05 | 0.86 | 0.89 | 0.0 | Detail |
| 1420 | CHOL | autosis | PALLD | palladin, cytoskeletal associated protein | 9.62 | 10.08 | 0.07 | 0.26 | 0.34 | 0.0 | Detail |
| 1421 | CHOL | autosis | CTPS1 | CTP synthase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1422 | CHOL | autosis | ICMT | isoprenylcysteine carboxyl methyltransferase | 10.42 | 10.58 | 0.02 | 0.45 | 0.53 | 0.0 | Detail |
| 1423 | CHOL | autosis | PIAS4 | protein inhibitor of activated STAT 4 | 8.09 | 8.46 | 0.06 | 0.04 | 0.06 | 0.0 | Detail |
| 1424 | CHOL | autosis | TNFAIP1 | TNF alpha induced protein 1 | 10.85 | 10.92 | 0.01 | 0.48 | 0.56 | 0.0 | Detail |
| 1425 | CHOL | autosis | MEF2A | myocyte enhancer factor 2A | 10.15 | 9.86 | -0.04 | 0.33 | 0.41 | 0.0 | Detail |
| 1426 | CHOL | autosis | MTPAP | mitochondrial poly(A) polymerase | 8.86 | 8.84 | -0.0 | 0.77 | 0.83 | 0.0 | Detail |
| 1427 | CHOL | autosis | NFAT5 | nuclear factor of activated T cells 5 | 9.78 | 10.3 | 0.07 | 0.07 | 0.12 | 0.0 | Detail |
| 1428 | CHOL | autosis | PSRC1 | proline and serine rich coiled-coil 1 | 5.63 | 6.93 | 0.3 | 0.33 | 0.41 | 0.0 | Detail |
| 1429 | CHOL | autosis | RCBTB1 | RCC1 and BTB domain containing protein 1 | 8.43 | 8.7 | 0.04 | 0.24 | 0.31 | 0.0 | Detail |
| 1430 | CHOL | autosis | RPS6KC1 | ribosomal protein S6 kinase C1 | 8.26 | 8.74 | 0.08 | 0.04 | 0.07 | 0.0 | Detail |
| 1431 | CHOL | autosis | PHF1 | PHD finger protein 1 | 9.68 | 10.13 | 0.07 | 0.08 | 0.13 | 0.0 | Detail |
| 1432 | CHOL | autosis | TADA3 | transcriptional adaptor 3 | 10.73 | 10.49 | -0.03 | 0.21 | 0.29 | 0.0 | Detail |
| 1433 | CHOL | autosis | CCNT1 | cyclin T1 | 5.64 | 5.58 | -0.01 | 0.7 | 0.76 | 0.0 | Detail |
| 1434 | CHOL | autosis | CCDC85B | coiled-coil domain containing 85B | 8.35 | 8.04 | -0.06 | 0.31 | 0.39 | 0.0 | Detail |
| 1435 | CHOL | autosis | DOK4 | docking protein 4 | 9.87 | 10.79 | 0.13 | 0.05 | 0.08 | 0.0 | Detail |
| 1436 | CHOL | autosis | PUM1 | pumilio RNA binding family member 1 | 10.5 | 10.6 | 0.01 | 0.81 | 0.86 | 0.0 | Detail |
| 1437 | CHOL | autosis | RBM39 | RNA binding motif protein 39 | 11.65 | 11.99 | 0.04 | 0.06 | 0.1 | 0.0 | Detail |
| 1438 | CHOL | autosis | TUBB2A | tubulin beta 2A class IIa | 10.92 | 10.07 | -0.12 | 0.05 | 0.09 | 0.0 | Detail |
| 1439 | CHOL | autosis | DNAJB4 | DnaJ heat shock protein family (Hsp40) member B4 | 8.68 | 8.01 | -0.12 | 0.07 | 0.12 | 0.0 | Detail |
| 1440 | CHOL | autosis | AMPH | amphiphysin | 3.66 | 3.6 | -0.03 | 0.58 | 0.66 | 0.0 | Detail |
| 1441 | CHOL | autosis | ATP8A1 | ATPase phospholipid transporting 8A1 | 7.02 | 6.5 | -0.11 | 0.21 | 0.29 | 0.0 | Detail |
| 1442 | CHOL | autosis | GMEB1 | glucocorticoid modulatory element binding protein 1 | 6.29 | 6.79 | 0.11 | 0.04 | 0.07 | 0.0 | Detail |
| 1443 | CHOL | autosis | GSTT2 | glutathione S-transferase theta 2 (gene/pseudogene) | 7.2 | 6.09 | -0.24 | 0.33 | 0.41 | 0.0 | Detail |
| 1444 | CHOL | autosis | HERC4 | HECT and RLD domain containing E3 ubiquitin protein ligase 4 | 9.7 | 9.68 | -0.0 | 0.81 | 0.86 | 0.0 | Detail |
| 1445 | CHOL | autosis | LZTS3 | leucine zipper tumor suppressor family member 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1446 | CHOL | autosis | MKNK2 | MAPK interacting serine/threonine kinase 2 | 11.74 | 11.45 | -0.04 | 0.08 | 0.13 | 0.0 | Detail |
| 1447 | CHOL | autosis | PARD6A | par-6 family cell polarity regulator alpha | 6.92 | 6.63 | -0.06 | 0.74 | 0.79 | 0.0 | Detail |
| 1448 | CHOL | autosis | POGLUT1 | protein O-glucosyltransferase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1449 | CHOL | autosis | PRMT7 | protein arginine methyltransferase 7 | 9.11 | 9.36 | 0.04 | 0.26 | 0.34 | 0.0 | Detail |
| 1450 | CHOL | autosis | RNF38 | ring finger protein 38 | 9.22 | 9.35 | 0.02 | 0.81 | 0.86 | 0.0 | Detail |
| 1451 | CHOL | autosis | SHTN1 | shootin 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1452 | CHOL | autosis | TRMO | tRNA methyltransferase O | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1453 | CHOL | autosis | YPEL5 | yippee like 5 | 10.84 | 10.82 | -0.0 | 0.85 | 0.89 | 0.0 | Detail |
| 1454 | CHOL | autosis | ATP1A4 | ATPase Na+/K+ transporting subunit alpha 4 | 0.42 | 0.88 | 1.05 | 0.53 | 0.61 | 0.0 | Detail |
| 1455 | CHOL | autosis | SSX2IP | SSX family member 2 interacting protein | 9.14 | 9.06 | -0.01 | 0.98 | 0.98 | 0.0 | Detail |
| 1456 | CHOL | autosis | ATG14 | autophagy related 14 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1457 | CHOL | autosis | ELOF1 | elongation factor 1 | 9.96 | 10.13 | 0.02 | 0.45 | 0.53 | 0.0 | Detail |
| 1458 | CHOL | autosis | MAP2K6 | mitogen-activated protein kinase kinase 6 | 6.51 | 6.77 | 0.06 | 0.51 | 0.59 | 0.0 | Detail |
| 1459 | CHOL | autosis | UBAP1 | ubiquitin associated protein 1 | 10.83 | 10.55 | -0.04 | 0.12 | 0.17 | 0.0 | Detail |
| 1460 | CHOL | autosis | B4GAT1 | beta-1,4-glucuronyltransferase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1461 | CHOL | autosis | ITSN2 | intersectin 2 | 9.94 | 10.13 | 0.03 | 0.42 | 0.5 | 0.0 | Detail |
| 1462 | CHOL | autosis | TANK | TRAF family member associated NFKB activator | 10.02 | 9.62 | -0.06 | 0.09 | 0.14 | 0.0 | Detail |
| 1463 | CHOL | autosis | AMOTL2 | angiomotin like 2 | 10.44 | 10.59 | 0.02 | 1.0 | 1.0 | 0.0 | Detail |
| 1464 | CHOL | autosis | CCNL2 | cyclin L2 | 11.3 | 11.25 | -0.01 | 0.74 | 0.79 | 0.0 | Detail |
| 1465 | CHOL | autosis | FEM1B | fem-1 homolog B | 10.35 | 10.0 | -0.05 | 0.14 | 0.21 | 0.0 | Detail |
| 1466 | CHOL | autosis | IPP | intracisternal A particle-promoted polypeptide | 7.24 | 7.42 | 0.04 | 0.39 | 0.47 | 0.0 | Detail |
| 1467 | CHOL | autosis | MED13L | mediator complex subunit 13L | 9.73 | 9.92 | 0.03 | 0.31 | 0.39 | 0.0 | Detail |
| 1468 | CHOL | autosis | SRSF11 | serine and arginine rich splicing factor 11 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1469 | CHOL | autosis | ATP2B4 | ATPase plasma membrane Ca2+ transporting 4 | 9.66 | 10.32 | 0.09 | 0.18 | 0.24 | 0.0 | Detail |
| 1470 | CHOL | autosis | MALL | mal, T cell differentiation protein like | 7.46 | 8.57 | 0.2 | 0.21 | 0.29 | 0.0 | Detail |
| 1471 | CHOL | autosis | MBP | myelin basic protein | 10.04 | 9.72 | -0.05 | 0.21 | 0.29 | 0.0 | Detail |
| 1472 | CHOL | autosis | MRPL36 | mitochondrial ribosomal protein L36 | 8.76 | 8.52 | -0.04 | 0.18 | 0.24 | 0.0 | Detail |
| 1473 | CHOL | autosis | NAV2 | neuron navigator 2 | 10.32 | 9.84 | -0.07 | 0.7 | 0.76 | 0.0 | Detail |
| 1474 | CHOL | autosis | NHLRC2 | NHL repeat containing 2 | 7.12 | 6.23 | -0.19 | 0.05 | 0.08 | 0.0 | Detail |
| 1475 | CHOL | autosis | ORC6 | origin recognition complex subunit 6 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1476 | CHOL | autosis | PCNX1 | pecanex 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1477 | CHOL | autosis | PPOX | protoporphyrinogen oxidase | 8.09 | 8.6 | 0.09 | 0.16 | 0.23 | 0.0 | Detail |
| 1478 | CHOL | autosis | SAC3D1 | SAC3 domain containing 1 | 7.61 | 8.07 | 0.08 | 0.16 | 0.23 | 0.0 | Detail |
| 1479 | CHOL | autosis | ZNF91 | zinc finger protein 91 | 7.92 | 8.18 | 0.05 | 0.19 | 0.26 | 0.0 | Detail |
| 1480 | CHOL | autosis | GATC | glutamyl-tRNA amidotransferase subunit C | 5.03 | 4.94 | -0.03 | 0.77 | 0.83 | 0.0 | Detail |
| 1481 | CHOL | autosis | SMC5 | structural maintenance of chromosomes 5 | 9.74 | 9.42 | -0.05 | 0.16 | 0.23 | 0.0 | Detail |
| 1482 | CHOL | autosis | CHTOP | chromatin target of PRMT1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1483 | CHOL | autosis | NDUFS6 | NADH:ubiquinone oxidoreductase subunit S6 | 10.56 | 10.58 | 0.0 | 0.7 | 0.76 | 0.0 | Detail |
| 1484 | CHOL | autosis | PSMB10 | proteasome 20S subunit beta 10 | 10.39 | 10.51 | 0.02 | 0.66 | 0.72 | 0.0 | Detail |
| 1485 | CHOL | autosis | SNAP23 | synaptosome associated protein 23 | 10.03 | 10.2 | 0.02 | 0.13 | 0.19 | 0.0 | Detail |
| 1486 | CHOL | autosis | BRD1 | bromodomain containing 1 | 9.42 | 9.79 | 0.06 | 0.09 | 0.14 | 0.0 | Detail |
| 1487 | CHOL | autosis | CNOT4 | CCR4-NOT transcription complex subunit 4 | 8.59 | 8.57 | -0.0 | 0.9 | 0.92 | 0.0 | Detail |
| 1488 | CHOL | autosis | DARS2 | aspartyl-tRNA synthetase 2, mitochondrial | 8.85 | 9.26 | 0.07 | 0.13 | 0.19 | 0.0 | Detail |
| 1489 | CHOL | autosis | MRPL24 | mitochondrial ribosomal protein L24 | 10.79 | 10.79 | 0.0 | 0.77 | 0.83 | 0.0 | Detail |
| 1490 | CHOL | autosis | MRPS11 | mitochondrial ribosomal protein S11 | 9.47 | 9.29 | -0.03 | 0.19 | 0.26 | 0.0 | Detail |
| 1491 | CHOL | autosis | NFE2L1 | NFE2 like bZIP transcription factor 1 | 12.97 | 12.78 | -0.02 | 0.42 | 0.5 | 0.0 | Detail |
| 1492 | CHOL | autosis | PI4K2A | phosphatidylinositol 4-kinase type 2 alpha | 9.6 | 9.59 | -0.0 | 1.0 | 1.0 | 0.0 | Detail |
| 1493 | CHOL | autosis | PIGK | phosphatidylinositol glycan anchor biosynthesis class K | 9.46 | 9.56 | 0.02 | 0.45 | 0.53 | 0.0 | Detail |
| 1494 | CHOL | autosis | PPP2R2D | protein phosphatase 2 regulatory subunit Bdelta | 9.46 | 9.03 | -0.07 | 0.04 | 0.06 | 0.0 | Detail |
| 1495 | CHOL | autosis | RPL7A | ribosomal protein L7a | 13.15 | 13.73 | 0.06 | 0.33 | 0.41 | 0.0 | Detail |
| 1496 | CHOL | autosis | TAF9 | TATA-box binding protein associated factor 9 | 10.05 | 10.05 | 0.0 | 1.0 | 1.0 | 0.0 | Detail |
| 1497 | CHOL | autosis | ZNF24 | zinc finger protein 24 | 10.73 | 10.54 | -0.03 | 0.48 | 0.56 | 0.0 | Detail |
| 1498 | CHOL | autosis | DSE | dermatan sulfate epimerase | 8.5 | 9.09 | 0.1 | 0.18 | 0.24 | 0.0 | Detail |
| 1499 | CHOL | autosis | ENC1 | ectodermal-neural cortex 1 | 9.56 | 9.59 | 0.0 | 0.94 | 0.95 | 0.0 | Detail |
| 1500 | CHOL | autosis | HECA | hdc homolog, cell cycle regulator | 9.6 | 9.03 | -0.09 | 0.05 | 0.09 | 0.0 | Detail |
| 1501 | CHOL | autosis | MAFF | MAF bZIP transcription factor F | 9.48 | 10.08 | 0.09 | 0.31 | 0.39 | 0.0 | Detail |
| 1502 | CHOL | autosis | N4BP2L2 | NEDD4 binding protein 2 like 2 | 10.01 | 9.93 | -0.01 | 0.74 | 0.79 | 0.0 | Detail |
| 1503 | CHOL | autosis | ORC1 | origin recognition complex subunit 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1504 | CHOL | autosis | PNRC1 | proline rich nuclear receptor coactivator 1 | 11.9 | 10.97 | -0.12 | 0.16 | 0.23 | 0.0 | Detail |
| 1505 | CHOL | autosis | PTPN18 | protein tyrosine phosphatase non-receptor type 18 | 10.42 | 10.71 | 0.04 | 0.09 | 0.14 | 0.0 | Detail |
| 1506 | CHOL | autosis | TOB2 | transducer of ERBB2, 2 | 10.23 | 10.83 | 0.08 | 0.04 | 0.06 | 0.0 | Detail |
| 1507 | CHOL | autosis | TREX1 | three prime repair exonuclease 1 | 8.08 | 8.05 | -0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1508 | CHOL | autosis | UBE2H | ubiquitin conjugating enzyme E2 H | 9.77 | 9.84 | 0.01 | 0.74 | 0.79 | 0.0 | Detail |
| 1509 | CHOL | autosis | AKAP17A | A-kinase anchoring protein 17A | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1510 | CHOL | autosis | B3GAT3 | beta-1,3-glucuronyltransferase 3 | 9.71 | 10.1 | 0.06 | 0.26 | 0.34 | 0.0 | Detail |
| 1511 | CHOL | autosis | ENSA | endosulfine alpha | 11.58 | 11.93 | 0.04 | 0.08 | 0.13 | 0.0 | Detail |
| 1512 | CHOL | autosis | ATP8B1 | ATPase phospholipid transporting 8B1 | 9.76 | 10.27 | 0.07 | 0.06 | 0.1 | 0.0 | Detail |
| 1513 | CHOL | autosis | TRAM2 | translocation associated membrane protein 2 | 9.84 | 9.15 | -0.1 | 0.03 | 0.06 | 0.0 | Detail |
| 1514 | CHOL | autosis | ADAMTS3 | ADAM metallopeptidase with thrombospondin type 1 motif 3 | 3.46 | 3.75 | 0.12 | 0.86 | 0.89 | 0.0 | Detail |
| 1515 | CHOL | autosis | CBY1 | chibby 1, beta catenin antagonist | 8.74 | 9.28 | 0.09 | 0.03 | 0.05 | 0.0 | Detail |
| 1516 | CHOL | autosis | PWWP3A | PWWP domain containing 3A, DNA repair factor | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1517 | CHOL | autosis | ATP2B1 | ATPase plasma membrane Ca2+ transporting 1 | 9.45 | 9.31 | -0.02 | 0.7 | 0.76 | 0.0 | Detail |
| 1518 | CHOL | autosis | UGDH | UDP-glucose 6-dehydrogenase | 11.41 | 10.18 | -0.16 | 0.04 | 0.06 | 0.0 | Detail |
| 1519 | CHOL | autosis | SETMAR | SET domain and mariner transposase fusion gene | 7.69 | 7.42 | -0.05 | 0.39 | 0.47 | 0.0 | Detail |
| 1520 | CHOL | autosis | ATP5ME | ATP synthase membrane subunit e | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1521 | CHOL | autosis | RPA2 | replication protein A2 | 9.09 | 9.33 | 0.04 | 0.42 | 0.5 | 0.0 | Detail |
| 1522 | CHOL | autosis | DDAH2 | DDAH family member 2, ADMA-independent | 10.02 | 10.91 | 0.12 | 0.03 | 0.06 | 0.0 | Detail |
| 1523 | CHOL | autosis | N6AMT1 | N-6 adenine-specific DNA methyltransferase 1 | 7.44 | 7.83 | 0.08 | 0.13 | 0.19 | 0.0 | Detail |
| 1524 | CHOL | autosis | SLC5A6 | solute carrier family 5 member 6 | 10.27 | 9.71 | -0.08 | 0.14 | 0.21 | 0.0 | Detail |
| 1525 | CHOL | autosis | GMPR2 | guanosine monophosphate reductase 2 | 10.15 | 10.2 | 0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1526 | CHOL | autosis | PLPP1 | phospholipid phosphatase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1527 | CHOL | autosis | COA1 | cytochrome c oxidase assembly factor 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1528 | CHOL | autosis | FAM102A | estrogen-induced osteoclastogenesis regulator 1 | 11.39 | 11.6 | 0.03 | 0.7 | 0.76 | 0.0 | Detail |
| 1529 | CHOL | autosis | INHBB | inhibin subunit beta B | 9.77 | 8.9 | -0.13 | 0.19 | 0.26 | 0.0 | Detail |
| 1530 | CHOL | autosis | RNASET2 | ribonuclease T2 | 9.73 | 9.85 | 0.02 | 0.98 | 0.98 | 0.0 | Detail |
| 1531 | CHOL | autosis | CGRRF1 | cell growth regulator with ring finger domain 1 | 6.94 | 7.06 | 0.02 | 0.51 | 0.59 | 0.0 | Detail |
| 1532 | CHOL | autosis | TOX4 | TOX high mobility group box family member 4 | 10.22 | 10.39 | 0.02 | 0.26 | 0.34 | 0.0 | Detail |
| 1533 | CHOL | autosis | ANKRD12 | ankyrin repeat domain 12 | 9.65 | 9.47 | -0.03 | 0.47 | 0.55 | 0.0 | Detail |
| 1534 | CHOL | autosis | ELOA | elongin A | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1535 | CHOL | autosis | ORC2 | origin recognition complex subunit 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1536 | CHOL | autosis | ARL5A | ADP ribosylation factor like GTPase 5A | 9.68 | 9.62 | -0.01 | 0.58 | 0.66 | 0.0 | Detail |
| 1537 | CHOL | autosis | COPS2 | COP9 signalosome subunit 2 | 10.27 | 10.4 | 0.02 | 0.33 | 0.41 | 0.0 | Detail |
| 1538 | CHOL | autosis | EXOSC4 | exosome component 4 | 8.98 | 9.08 | 0.02 | 0.86 | 0.89 | 0.0 | Detail |
| 1539 | CHOL | autosis | SEPHS1 | selenophosphate synthetase 1 | 10.12 | 10.07 | -0.01 | 0.98 | 0.98 | 0.0 | Detail |
| 1540 | CHOL | autosis | NME3 | NME/NM23 nucleoside diphosphate kinase 3 | 9.34 | 9.98 | 0.09 | 0.03 | 0.06 | 0.0 | Detail |
| 1541 | CHOL | autosis | ADCY8 | adenylate cyclase 8 | 1.14 | 2.11 | 0.88 | 0.49 | 0.58 | 0.0 | Detail |
| 1542 | CHOL | autosis | B4GALT6 | beta-1,4-galactosyltransferase 6 | 6.27 | 6.59 | 0.07 | 0.51 | 0.59 | 0.0 | Detail |
| 1543 | CHOL | autosis | CA6 | carbonic anhydrase 6 | 0.0 | 0.14 | inf | 0.46 | 0.55 | 0.0 | Detail |
| 1544 | CHOL | autosis | CHPF2 | chondroitin polymerizing factor 2 | 9.92 | 10.39 | 0.07 | 0.04 | 0.06 | 0.0 | Detail |
| 1545 | CHOL | autosis | FOCAD | focadhesin | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1546 | CHOL | autosis | GJD3 | gap junction protein delta 3 | 6.45 | 6.06 | -0.09 | 0.51 | 0.59 | 0.0 | Detail |
| 1547 | CHOL | autosis | PIGN | phosphatidylinositol glycan anchor biosynthesis class N | 8.77 | 8.97 | 0.03 | 0.36 | 0.44 | 0.0 | Detail |
| 1548 | CHOL | autosis | PIGO | phosphatidylinositol glycan anchor biosynthesis class O | 9.57 | 9.6 | 0.0 | 0.9 | 0.92 | 0.0 | Detail |
| 1549 | CHOL | autosis | SLC18A1 | solute carrier family 18 member A1 | 0.33 | 1.07 | 1.69 | 0.39 | 0.47 | 0.0 | Detail |
| 1550 | CHOL | autosis | SPIDR | scaffold protein involved in DNA repair | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1551 | CHOL | autosis | ZNF442 | zinc finger protein 442 | 4.79 | 3.83 | -0.32 | 0.03 | 0.06 | 0.0 | Detail |
| 1552 | CHOL | autosis | MPPE1 | metallophosphoesterase 1 | 9.5 | 9.62 | 0.02 | 0.58 | 0.66 | 0.0 | Detail |
| 1553 | CHOL | autosis | PPP1R7 | protein phosphatase 1 regulatory subunit 7 | 10.2 | 10.33 | 0.02 | 0.36 | 0.44 | 0.0 | Detail |
| 1554 | CHOL | autosis | WAPL | WAPL cohesin release factor | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1555 | CHOL | autosis | TMCC1 | transmembrane and coiled-coil domain family 1 | 9.46 | 9.24 | -0.03 | 0.45 | 0.53 | 0.0 | Detail |
| 1556 | CHOL | autosis | ATP11B | ATPase phospholipid transporting 11B (putative) | 9.79 | 9.91 | 0.02 | 0.74 | 0.79 | 0.0 | Detail |
| 1557 | CHOL | autosis | N4BP1 | NEDD4 binding protein 1 | 9.42 | 9.76 | 0.05 | 0.05 | 0.08 | 0.0 | Detail |
| 1558 | CHOL | autosis | SPSB1 | splA/ryanodine receptor domain and SOCS box containing 1 | 9.94 | 10.22 | 0.04 | 0.55 | 0.63 | 0.0 | Detail |
| 1559 | CHOL | autosis | GM2A | ganglioside GM2 activator | 10.33 | 10.58 | 0.04 | 0.55 | 0.63 | 0.0 | Detail |
| 1560 | CHOL | autosis | RFC5 | replication factor C subunit 5 | 8.06 | 8.52 | 0.08 | 0.05 | 0.09 | 0.0 | Detail |
| 1561 | CHOL | autosis | SNU13 | small nuclear ribonucleoprotein 13 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1562 | CHOL | autosis | MPZL2 | myelin protein zero like 2 | 9.17 | 9.29 | 0.02 | 0.94 | 0.95 | 0.0 | Detail |
| 1563 | CHOL | autosis | NUP58 | nucleoporin 58 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1564 | CHOL | autosis | RETREG2 | reticulophagy regulator family member 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1565 | CHOL | autosis | ZSCAN31 | zinc finger and SCAN domain containing 31 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1566 | CHOL | autosis | LAPTM5 | lysosomal protein transmembrane 5 | 10.68 | 11.45 | 0.1 | 0.21 | 0.29 | 0.0 | Detail |
| 1567 | CHOL | autosis | PPP2CB | protein phosphatase 2 catalytic subunit beta | 10.98 | 10.73 | -0.03 | 0.36 | 0.44 | 0.0 | Detail |
| 1568 | CHOL | autosis | CEBPG | CCAAT enhancer binding protein gamma | 10.82 | 10.38 | -0.06 | 0.06 | 0.1 | 0.0 | Detail |
| 1569 | CHOL | autosis | ARFGEF2 | ADP ribosylation factor guanine nucleotide exchange factor 2 | 10.5 | 10.37 | -0.02 | 0.66 | 0.72 | 0.0 | Detail |
| 1570 | CHOL | autosis | ATP6V0A2 | ATPase H+ transporting V0 subunit a2 | 8.27 | 8.46 | 0.03 | 0.62 | 0.69 | 0.0 | Detail |
| 1571 | CHOL | autosis | CHORDC1 | cysteine and histidine rich domain containing 1 | 8.22 | 8.31 | 0.02 | 0.58 | 0.66 | 0.0 | Detail |
| 1572 | CHOL | autosis | HTT | huntingtin | 10.42 | 10.65 | 0.03 | 0.24 | 0.31 | 0.0 | Detail |
| 1573 | CHOL | autosis | NREP | neuronal regeneration related protein | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1574 | CHOL | autosis | PGM3 | phosphoglucomutase 3 | 9.03 | 8.18 | -0.14 | 0.04 | 0.07 | 0.0 | Detail |
| 1575 | CHOL | autosis | POMP | proteasome maturation protein | 10.84 | 10.6 | -0.03 | 0.18 | 0.24 | 0.0 | Detail |
| 1576 | CHOL | autosis | PSMD3 | proteasome 26S subunit, non-ATPase 3 | 11.41 | 11.39 | -0.0 | 0.94 | 0.95 | 0.0 | Detail |
| 1577 | CHOL | autosis | SYNPO | synaptopodin | 11.7 | 11.81 | 0.01 | 0.7 | 0.76 | 0.0 | Detail |
| 1578 | CHOL | autosis | GNB1 | G protein subunit beta 1 | 12.23 | 12.53 | 0.03 | 0.39 | 0.47 | 0.0 | Detail |
| 1579 | CHOL | autosis | AGO3 | argonaute RISC catalytic component 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1580 | CHOL | autosis | AP2A2 | adaptor related protein complex 2 subunit alpha 2 | 11.18 | 11.26 | 0.01 | 0.81 | 0.86 | 0.0 | Detail |
| 1581 | CHOL | autosis | ATP2B3 | ATPase plasma membrane Ca2+ transporting 3 | 0.42 | 0.33 | -0.35 | 0.55 | 0.63 | 0.0 | Detail |
| 1582 | CHOL | autosis | CISD1 | CDGSH iron sulfur domain 1 | 9.26 | 8.79 | -0.07 | 0.18 | 0.24 | 0.0 | Detail |
| 1583 | CHOL | autosis | CSTF1 | cleavage stimulation factor subunit 1 | 9.24 | 9.15 | -0.01 | 0.48 | 0.56 | 0.0 | Detail |
| 1584 | CHOL | autosis | EIF2B1 | eukaryotic translation initiation factor 2B subunit alpha | 9.81 | 10.08 | 0.04 | 0.04 | 0.07 | 0.0 | Detail |
| 1585 | CHOL | autosis | HIVEP2 | HIVEP zinc finger 2 | 8.62 | 9.3 | 0.11 | 0.09 | 0.14 | 0.0 | Detail |
| 1586 | CHOL | autosis | INAVA | innate immunity activator | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1587 | CHOL | autosis | KCNK1 | potassium two pore domain channel subfamily K member 1 | 7.76 | 8.45 | 0.12 | 0.09 | 0.14 | 0.0 | Detail |
| 1588 | CHOL | autosis | USPL1 | ubiquitin specific peptidase like 1 | 8.64 | 8.11 | -0.09 | 0.04 | 0.06 | 0.0 | Detail |
| 1589 | CHOL | autosis | VPS4B | vacuolar protein sorting 4 homolog B | 10.08 | 10.14 | 0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1590 | CHOL | autosis | ZDHHC3 | zinc finger DHHC-type palmitoyltransferase 3 | 10.09 | 9.87 | -0.03 | 0.08 | 0.13 | 0.0 | Detail |
| 1591 | CHOL | autosis | SF1 | splicing factor 1 | 11.77 | 12.09 | 0.04 | 0.03 | 0.05 | 0.0 | Detail |
| 1592 | CHOL | autosis | EN1 | engrailed homeobox 1 | 0.53 | 2.85 | 2.42 | 0.03 | 0.05 | 0.0 | Detail |
| 1593 | CHOL | autosis | RIOK3 | RIO kinase 3 | 11.19 | 10.7 | -0.07 | 0.03 | 0.06 | 0.0 | Detail |
| 1594 | CHOL | autosis | EFHD2 | EF-hand domain family member D2 | 10.21 | 10.63 | 0.06 | 0.33 | 0.41 | 0.0 | Detail |
| 1595 | CHOL | autosis | POLE2 | DNA polymerase epsilon 2, accessory subunit | 4.66 | 5.01 | 0.1 | 0.86 | 0.89 | 0.0 | Detail |
| 1596 | CHOL | autosis | WDCP | WD repeat and coiled coil containing | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1597 | CHOL | autosis | GABBR2 | gamma-aminobutyric acid type B receptor subunit 2 | 0.77 | 2.0 | 1.37 | 0.03 | 0.06 | 0.0 | Detail |
| 1598 | CHOL | autosis | TIMM10 | translocase of inner mitochondrial membrane 10 | 9.35 | 9.03 | -0.05 | 0.26 | 0.34 | 0.0 | Detail |
| 1599 | CHOL | autosis | ADAT1 | adenosine deaminase tRNA specific 1 | 6.81 | 7.37 | 0.11 | 0.13 | 0.19 | 0.0 | Detail |
| 1600 | CHOL | autosis | DNAJB14 | DnaJ heat shock protein family (Hsp40) member B14 | 7.35 | 7.52 | 0.03 | 0.33 | 0.41 | 0.0 | Detail |
| 1601 | CHOL | autosis | PRRC1 | proline rich coiled-coil 1 | 10.37 | 10.7 | 0.05 | 0.16 | 0.23 | 0.0 | Detail |
| 1602 | CHOL | autosis | TM7SF3 | transmembrane 7 superfamily member 3 | 11.4 | 11.58 | 0.02 | 0.48 | 0.56 | 0.0 | Detail |
| 1603 | CHOL | autosis | ALG6 | ALG6 alpha-1,3-glucosyltransferase | 7.88 | 8.33 | 0.08 | 0.03 | 0.06 | 0.0 | Detail |
| 1604 | CHOL | autosis | ALG8 | ALG8 alpha-1,3-glucosyltransferase | 9.73 | 9.38 | -0.05 | 0.09 | 0.14 | 0.0 | Detail |
| 1605 | CHOL | autosis | AP3S2 | adaptor related protein complex 3 subunit sigma 2 | 10.93 | 10.6 | -0.04 | 0.09 | 0.14 | 0.0 | Detail |
| 1606 | CHOL | autosis | ARSJ | arylsulfatase family member J | 6.51 | 7.63 | 0.23 | 0.04 | 0.06 | 0.0 | Detail |
| 1607 | CHOL | autosis | ATP6V1B2 | ATPase H+ transporting V1 subunit B2 | 10.79 | 10.7 | -0.01 | 0.51 | 0.59 | 0.0 | Detail |
| 1608 | CHOL | autosis | ATP9A | ATPase phospholipid transporting 9A (putative) | 9.98 | 10.86 | 0.12 | 0.03 | 0.06 | 0.0 | Detail |
| 1609 | CHOL | autosis | ATXN10 | ataxin 10 | 11.08 | 11.04 | -0.0 | 0.9 | 0.92 | 0.0 | Detail |
| 1610 | CHOL | autosis | BAHD1 | bromo adjacent homology domain containing 1 | 9.58 | 9.32 | -0.04 | 0.42 | 0.5 | 0.0 | Detail |
| 1611 | CHOL | autosis | BIN3 | bridging integrator 3 | 8.71 | 9.03 | 0.05 | 0.31 | 0.39 | 0.0 | Detail |
| 1612 | CHOL | autosis | BSCL2 | BSCL2 lipid droplet biogenesis associated, seipin | 10.26 | 10.76 | 0.07 | 0.09 | 0.14 | 0.0 | Detail |
| 1613 | CHOL | autosis | CCDC15 | coiled-coil domain containing 15 | 5.12 | 5.2 | 0.02 | 0.7 | 0.76 | 0.0 | Detail |
| 1614 | CHOL | autosis | CCNT2 | cyclin T2 | 9.28 | 9.46 | 0.03 | 0.39 | 0.47 | 0.0 | Detail |
| 1615 | CHOL | autosis | CDK17 | cyclin dependent kinase 17 | 8.87 | 8.95 | 0.01 | 0.48 | 0.56 | 0.0 | Detail |
| 1616 | CHOL | autosis | CDKN2AIP | CDKN2A interacting protein | 8.5 | 8.19 | -0.05 | 0.26 | 0.34 | 0.0 | Detail |
| 1617 | CHOL | autosis | CHIC2 | cysteine rich hydrophobic domain 2 | 8.13 | 7.95 | -0.03 | 0.62 | 0.69 | 0.0 | Detail |
| 1618 | CHOL | autosis | CLCN6 | chloride voltage-gated channel 6 | 8.59 | 8.08 | -0.09 | 0.04 | 0.07 | 0.0 | Detail |
| 1619 | CHOL | autosis | CLK4 | CDC like kinase 4 | 8.77 | 8.7 | -0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1620 | CHOL | autosis | CTNS | cystinosin, lysosomal cystine transporter | 8.73 | 8.63 | -0.02 | 0.9 | 0.92 | 0.0 | Detail |
| 1621 | CHOL | autosis | DCUN1D2 | defective in cullin neddylation 1 domain containing 2 | 7.64 | 7.8 | 0.03 | 0.66 | 0.72 | 0.0 | Detail |
| 1622 | CHOL | autosis | DENND4C | DENN domain containing 4C | 10.03 | 9.5 | -0.08 | 0.28 | 0.36 | 0.0 | Detail |
| 1623 | CHOL | autosis | DOLK | dolichol kinase | 8.71 | 8.55 | -0.03 | 0.19 | 0.26 | 0.0 | Detail |
| 1624 | CHOL | autosis | DPM2 | dolichyl-phosphate mannosyltransferase subunit 2, regulatory | 9.32 | 9.87 | 0.08 | 0.07 | 0.12 | 0.0 | Detail |
| 1625 | CHOL | autosis | DPY19L1 | dpy-19 like C-mannosyltransferase 1 | 9.72 | 9.26 | -0.07 | 0.1 | 0.16 | 0.0 | Detail |
| 1626 | CHOL | autosis | DYNC1LI1 | dynein cytoplasmic 1 light intermediate chain 1 | 8.91 | 9.0 | 0.01 | 0.77 | 0.83 | 0.0 | Detail |
| 1627 | CHOL | autosis | EEF1AKMT3 | EEF1A lysine methyltransferase 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1628 | CHOL | autosis | EMC9 | ER membrane protein complex subunit 9 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1629 | CHOL | autosis | EML3 | EMAP like 3 | 9.82 | 10.03 | 0.03 | 0.74 | 0.79 | 0.0 | Detail |
| 1630 | CHOL | autosis | ERG28 | ergosterol biosynthesis 28 homolog | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1631 | CHOL | autosis | FAM189A2 | endosomal transmembrane epsin interactor 1 | 2.86 | 4.92 | 0.78 | 0.03 | 0.05 | 0.0 | Detail |
| 1632 | CHOL | autosis | FBXL12 | F-box and leucine rich repeat protein 12 | 8.41 | 8.61 | 0.03 | 0.1 | 0.16 | 0.0 | Detail |
| 1633 | CHOL | autosis | FBXO38 | F-box protein 38 | 9.54 | 9.56 | 0.0 | 0.86 | 0.89 | 0.0 | Detail |
| 1634 | CHOL | autosis | FBXO9 | F-box protein 9 | 9.97 | 9.75 | -0.03 | 0.21 | 0.28 | 0.0 | Detail |
| 1635 | CHOL | autosis | FEM1C | fem-1 homolog C | 9.83 | 9.3 | -0.08 | 0.1 | 0.16 | 0.0 | Detail |
| 1636 | CHOL | autosis | GEMIN6 | gem nuclear organelle associated protein 6 | 7.47 | 7.97 | 0.09 | 0.04 | 0.06 | 0.0 | Detail |
| 1637 | CHOL | autosis | INTS5 | integrator complex subunit 5 | 9.36 | 9.42 | 0.01 | 0.62 | 0.69 | 0.0 | Detail |
| 1638 | CHOL | autosis | KCTD14 | potassium channel tetramerization domain containing 14 | 6.96 | 5.59 | -0.32 | 0.18 | 0.24 | 0.0 | Detail |
| 1639 | CHOL | autosis | KHNYN | KH and NYN domain containing | 9.68 | 10.12 | 0.06 | 0.05 | 0.09 | 0.0 | Detail |
| 1640 | CHOL | autosis | KLHL24 | kelch like family member 24 | 10.29 | 10.01 | -0.04 | 0.39 | 0.47 | 0.0 | Detail |
| 1641 | CHOL | autosis | LAGE3 | L antigen family member 3 | 8.22 | 8.56 | 0.06 | 0.14 | 0.21 | 0.0 | Detail |
| 1642 | CHOL | autosis | LIPT1 | lipoyltransferase 1 | 6.75 | 6.82 | 0.01 | 0.81 | 0.86 | 0.0 | Detail |
| 1643 | CHOL | autosis | LMBR1L | limb development membrane protein 1 like | 8.61 | 9.03 | 0.07 | 0.1 | 0.16 | 0.0 | Detail |
| 1644 | CHOL | autosis | LRIF1 | ligand dependent nuclear receptor interacting factor 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1645 | CHOL | autosis | MID2 | midline 2 | 8.35 | 8.23 | -0.02 | 0.51 | 0.59 | 0.0 | Detail |
| 1646 | CHOL | autosis | MRPS34 | mitochondrial ribosomal protein S34 | 10.44 | 10.71 | 0.04 | 0.24 | 0.31 | 0.0 | Detail |
| 1647 | CHOL | autosis | MYO9A | myosin IXA | 6.21 | 6.02 | -0.05 | 0.9 | 0.92 | 0.0 | Detail |
| 1648 | CHOL | autosis | NRGN | neurogranin | 5.75 | 6.74 | 0.23 | 0.09 | 0.14 | 0.0 | Detail |
| 1649 | CHOL | autosis | OSER1 | oxidative stress responsive serine rich 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1650 | CHOL | autosis | PEX11B | peroxisomal biogenesis factor 11 beta | 9.43 | 9.55 | 0.02 | 0.62 | 0.69 | 0.0 | Detail |
| 1651 | CHOL | autosis | PGAP2 | post-GPI attachment to proteins 2 | 9.45 | 9.31 | -0.02 | 0.55 | 0.63 | 0.0 | Detail |
| 1652 | CHOL | autosis | POLR2I | RNA polymerase II subunit I | 9.26 | 9.32 | 0.01 | 0.98 | 0.98 | 0.0 | Detail |
| 1653 | CHOL | autosis | POLR2L | RNA polymerase II, I and III subunit L | 11.02 | 10.7 | -0.04 | 0.31 | 0.39 | 0.0 | Detail |
| 1654 | CHOL | autosis | PPP2R5B | protein phosphatase 2 regulatory subunit B'beta | 8.02 | 8.55 | 0.09 | 0.08 | 0.13 | 0.0 | Detail |
| 1655 | CHOL | autosis | PSG6 | pregnancy specific beta-1-glycoprotein 6 | 0.0 | 0.29 | inf | 0.31 | 0.4 | 0.0 | Detail |
| 1656 | CHOL | autosis | PTCD2 | pentatricopeptide repeat domain 2 | 6.75 | 6.46 | -0.06 | 0.28 | 0.36 | 0.0 | Detail |
| 1657 | CHOL | autosis | PTDSS1 | phosphatidylserine synthase 1 | 10.71 | 10.54 | -0.02 | 0.33 | 0.41 | 0.0 | Detail |
| 1658 | CHOL | autosis | R3HCC1L | R3H domain and coiled-coil containing 1 like | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1659 | CHOL | autosis | RAB29 | RAB29, member RAS oncogene family | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1660 | CHOL | autosis | RBM48 | RNA binding motif protein 48 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1661 | CHOL | autosis | RIPOR1 | RHO family interacting cell polarization regulator 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1662 | CHOL | autosis | RPL38 | ribosomal protein L38 | 12.11 | 12.77 | 0.08 | 0.05 | 0.09 | 0.0 | Detail |
| 1663 | CHOL | autosis | RRAGA | Ras related GTP binding A | 10.73 | 10.33 | -0.05 | 0.05 | 0.09 | 0.0 | Detail |
| 1664 | CHOL | autosis | RRAGC | Ras related GTP binding C | 7.34 | 7.97 | 0.12 | 0.05 | 0.09 | 0.0 | Detail |
| 1665 | CHOL | autosis | RRAGD | Ras related GTP binding D | 8.44 | 7.73 | -0.13 | 0.24 | 0.31 | 0.0 | Detail |
| 1666 | CHOL | autosis | SFSWAP | splicing factor SWAP | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1667 | CHOL | autosis | SNRNP25 | small nuclear ribonucleoprotein U11/U12 subunit 25 | 9.24 | 8.8 | -0.07 | 0.1 | 0.16 | 0.0 | Detail |
| 1668 | CHOL | autosis | SPTSSA | serine palmitoyltransferase small subunit A | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1669 | CHOL | autosis | SRSF8 | serine and arginine rich splicing factor 8 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1670 | CHOL | autosis | TAF10 | TATA-box binding protein associated factor 10 | 9.94 | 10.01 | 0.01 | 0.86 | 0.89 | 0.0 | Detail |
| 1671 | CHOL | autosis | TBC1D2 | TBC1 domain family member 2 | 8.57 | 8.09 | -0.08 | 0.19 | 0.26 | 0.0 | Detail |
| 1672 | CHOL | autosis | TBC1D2B | TBC1 domain family member 2B | 10.97 | 10.94 | -0.0 | 0.81 | 0.86 | 0.0 | Detail |
| 1673 | CHOL | autosis | TCTN3 | tectonic family member 3 | 9.75 | 10.1 | 0.05 | 0.04 | 0.06 | 0.0 | Detail |
| 1674 | CHOL | autosis | THYN1 | thymocyte nuclear protein 1 | 9.52 | 9.02 | -0.08 | 0.03 | 0.06 | 0.0 | Detail |
| 1675 | CHOL | autosis | TMEM223 | transmembrane protein 223 | 8.81 | 8.68 | -0.02 | 0.42 | 0.5 | 0.0 | Detail |
| 1676 | CHOL | autosis | TSEN2 | tRNA splicing endonuclease subunit 2 | 6.74 | 7.0 | 0.05 | 0.16 | 0.23 | 0.0 | Detail |
| 1677 | CHOL | autosis | TXNL4B | thioredoxin like 4B | 8.47 | 8.61 | 0.02 | 0.31 | 0.39 | 0.0 | Detail |
| 1678 | CHOL | autosis | UBXN7 | UBX domain protein 7 | 6.69 | 7.08 | 0.08 | 0.39 | 0.47 | 0.0 | Detail |
| 1679 | CHOL | autosis | UGGT2 | UDP-glucose glycoprotein glucosyltransferase 2 | 8.05 | 8.38 | 0.06 | 0.19 | 0.26 | 0.0 | Detail |
| 1680 | CHOL | autosis | UPF2 | UPF2 regulator of nonsense mediated mRNA decay | 9.46 | 9.76 | 0.05 | 0.09 | 0.14 | 0.0 | Detail |
| 1681 | CHOL | autosis | UQCR10 | ubiquinol-cytochrome c reductase, complex III subunit X | 10.97 | 10.54 | -0.06 | 0.09 | 0.14 | 0.0 | Detail |
| 1682 | CHOL | autosis | USB1 | U6 snRNA biogenesis phosphodiesterase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1683 | CHOL | autosis | VCPKMT | valosin containing protein lysine methyltransferase | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1684 | CHOL | autosis | WDR5B | WD repeat domain 5B | 7.88 | 8.13 | 0.05 | 0.28 | 0.36 | 0.0 | Detail |
| 1685 | CHOL | autosis | WIPI1 | WD repeat domain, phosphoinositide interacting 1 | 8.9 | 9.51 | 0.1 | 0.19 | 0.26 | 0.0 | Detail |
| 1686 | CHOL | autosis | ZBED8 | family with sequence similarity 200 member C | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1687 | CHOL | autosis | ZBTB25 | zinc finger and BTB domain containing 25 | 5.92 | 6.11 | 0.05 | 0.58 | 0.66 | 0.0 | Detail |
| 1688 | CHOL | autosis | ZBTB43 | zinc finger and BTB domain containing 43 | 8.69 | 8.16 | -0.09 | 0.05 | 0.09 | 0.0 | Detail |
| 1689 | CHOL | autosis | ZC2HC1A | zinc finger C2HC-type containing 1A | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1690 | CHOL | autosis | ZC3H7A | zinc finger CCCH-type containing 7A | 9.63 | 9.91 | 0.04 | 0.07 | 0.12 | 0.0 | Detail |
| 1691 | CHOL | autosis | ZMYM5 | zinc finger MYM-type containing 5 | 7.55 | 7.8 | 0.05 | 0.33 | 0.41 | 0.0 | Detail |
| 1692 | CHOL | autosis | ZNF140 | zinc finger protein 140 | 8.38 | 8.66 | 0.05 | 0.18 | 0.24 | 0.0 | Detail |
| 1693 | CHOL | autosis | ZNF212 | zinc finger protein 212 | 8.16 | 8.47 | 0.05 | 0.08 | 0.13 | 0.0 | Detail |
| 1694 | CHOL | autosis | ZNF394 | zinc finger protein 394 | 8.56 | 8.85 | 0.05 | 0.19 | 0.26 | 0.0 | Detail |
| 1695 | CHOL | autosis | PLCL2 | phospholipase C like 2 | 8.75 | 8.26 | -0.08 | 0.14 | 0.21 | 0.0 | Detail |
| 1696 | CHOL | autosis | TMEM160 | transmembrane protein 160 | 7.16 | 6.52 | -0.13 | 0.04 | 0.06 | 0.0 | Detail |
| 1697 | CHOL | autosis | BLOC1S1 | biogenesis of lysosomal organelles complex 1 subunit 1 | 11.04 | 10.6 | -0.06 | 0.06 | 0.1 | 0.0 | Detail |
| 1698 | CHOL | autosis | ABHD3 | abhydrolase domain containing 3, phospholipase | 9.64 | 10.48 | 0.12 | 0.03 | 0.05 | 0.0 | Detail |
| 1699 | CHOL | autosis | ANAPC13 | anaphase promoting complex subunit 13 | 10.22 | 9.97 | -0.04 | 0.28 | 0.36 | 0.0 | Detail |
| 1700 | CHOL | autosis | ANAPC15 | anaphase promoting complex subunit 15 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1701 | CHOL | autosis | ANKFY1 | ankyrin repeat and FYVE domain containing 1 | 10.64 | 10.3 | -0.05 | 0.05 | 0.09 | 0.0 | Detail |
| 1702 | CHOL | autosis | AP1AR | adaptor related protein complex 1 associated regulatory protein | 9.17 | 8.55 | -0.1 | 0.16 | 0.23 | 0.0 | Detail |
| 1703 | CHOL | autosis | ARHGAP45 | Rho GTPase activating protein 45 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1704 | CHOL | autosis | ART4 | ADP-ribosyltransferase 4 (inactive) (Dombrock blood group) | 8.94 | 7.29 | -0.29 | 0.08 | 0.13 | 0.0 | Detail |
| 1705 | CHOL | autosis | ASTE1 | asteroid homolog 1 | 7.41 | 7.12 | -0.06 | 0.19 | 0.26 | 0.0 | Detail |
| 1706 | CHOL | autosis | ATP10B | ATPase phospholipid transporting 10B (putative) | 2.63 | 4.22 | 0.68 | 0.38 | 0.46 | 0.0 | Detail |
| 1707 | CHOL | autosis | ATP10D | ATPase phospholipid transporting 10D (putative) | 7.56 | 7.86 | 0.06 | 0.51 | 0.59 | 0.0 | Detail |
| 1708 | CHOL | autosis | ATP8B3 | ATPase phospholipid transporting 8B3 | 4.56 | 5.72 | 0.33 | 0.04 | 0.06 | 0.0 | Detail |
| 1709 | CHOL | autosis | ATP9B | ATPase phospholipid transporting 9B (putative) | 8.55 | 8.51 | -0.01 | 0.42 | 0.5 | 0.0 | Detail |
| 1710 | CHOL | autosis | BET1 | Bet1 golgi vesicular membrane trafficking protein | 9.08 | 9.14 | 0.01 | 0.9 | 0.92 | 0.0 | Detail |
| 1711 | CHOL | autosis | BOLA1 | bolA family member 1 | 8.33 | 8.35 | 0.0 | 0.9 | 0.92 | 0.0 | Detail |
| 1712 | CHOL | autosis | C6orf89 | chromosome 6 open reading frame 89 | 11.12 | 11.18 | 0.01 | 0.51 | 0.59 | 0.0 | Detail |
| 1713 | CHOL | autosis | C9orf40 | chromosome 9 open reading frame 40 | 6.38 | 6.1 | -0.07 | 0.51 | 0.59 | 0.0 | Detail |
| 1714 | CHOL | autosis | CCSER2 | coiled-coil serine rich protein 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1715 | CHOL | autosis | CENPS | centromere protein S | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1716 | CHOL | autosis | CEP112 | centrosomal protein 112 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1717 | CHOL | autosis | CHST2 | carbohydrate sulfotransferase 2 | 6.21 | 6.91 | 0.15 | 0.13 | 0.19 | 0.0 | Detail |
| 1718 | CHOL | autosis | CLN6 | CLN6 transmembrane ER protein | 9.7 | 10.13 | 0.06 | 0.06 | 0.1 | 0.0 | Detail |
| 1719 | CHOL | autosis | CMC4 | C-X9-C motif containing 4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1720 | CHOL | autosis | COA4 | cytochrome c oxidase assembly factor 4 homolog | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1721 | CHOL | autosis | CSGALNACT2 | chondroitin sulfate N-acetylgalactosaminyltransferase 2 | 8.6 | 8.77 | 0.03 | 0.18 | 0.24 | 0.0 | Detail |
| 1722 | CHOL | autosis | CSTF2T | cleavage stimulation factor subunit 2 tau variant | 9.5 | 9.62 | 0.02 | 0.9 | 0.92 | 0.0 | Detail |
| 1723 | CHOL | autosis | CSTF3 | cleavage stimulation factor subunit 3 | 8.97 | 9.15 | 0.03 | 0.16 | 0.23 | 0.0 | Detail |
| 1724 | CHOL | autosis | DCAF8 | DDB1 and CUL4 associated factor 8 | 11.9 | 11.51 | -0.05 | 0.03 | 0.05 | 0.0 | Detail |
| 1725 | CHOL | autosis | DET1 | DET1 partner of COP1 E3 ubiquitin ligase | 6.7 | 6.46 | -0.05 | 0.31 | 0.39 | 0.0 | Detail |
| 1726 | CHOL | autosis | DNAAF2 | dynein axonemal assembly factor 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1727 | CHOL | autosis | DNPH1 | 2'-deoxynucleoside 5'-phosphate N-hydrolase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1728 | CHOL | autosis | EEF2KMT | eukaryotic elongation factor 2 lysine methyltransferase | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1729 | CHOL | autosis | EID1 | EP300 interacting inhibitor of differentiation 1 | 11.75 | 11.39 | -0.05 | 0.13 | 0.19 | 0.0 | Detail |
| 1730 | CHOL | autosis | EXOG | exo/endonuclease G | 6.27 | 6.5 | 0.05 | 0.58 | 0.66 | 0.0 | Detail |
| 1731 | CHOL | autosis | EXTL2 | exostosin like glycosyltransferase 2 | 8.12 | 8.04 | -0.01 | 0.98 | 0.98 | 0.0 | Detail |
| 1732 | CHOL | autosis | FAM169A | family with sequence similarity 169 member A | 6.98 | 6.7 | -0.06 | 0.7 | 0.76 | 0.0 | Detail |
| 1733 | CHOL | autosis | FICD | FIC domain protein adenylyltransferase | 5.45 | 5.6 | 0.04 | 0.66 | 0.72 | 0.0 | Detail |
| 1734 | CHOL | autosis | FMO1 | flavin containing dimethylaniline monoxygenase 1 | 5.59 | 5.63 | 0.01 | 0.86 | 0.89 | 0.0 | Detail |
| 1735 | CHOL | autosis | FNTB | farnesyltransferase, CAAX box, subunit beta | 8.51 | 8.65 | 0.02 | 0.42 | 0.5 | 0.0 | Detail |
| 1736 | CHOL | autosis | GPATCH3 | G-patch domain containing 3 | 8.25 | 8.06 | -0.03 | 0.33 | 0.41 | 0.0 | Detail |
| 1737 | CHOL | autosis | GPBP1L1 | GC-rich promoter binding protein 1 like 1 | 11.55 | 11.34 | -0.03 | 0.58 | 0.66 | 0.0 | Detail |
| 1738 | CHOL | autosis | GPR137B | G protein-coupled receptor 137B | 8.35 | 9.07 | 0.12 | 0.13 | 0.19 | 0.0 | Detail |
| 1739 | CHOL | autosis | GPR176 | G protein-coupled receptor 176 | 4.97 | 6.04 | 0.28 | 0.12 | 0.17 | 0.0 | Detail |
| 1740 | CHOL | autosis | HS1BP3 | HCLS1 binding protein 3 | 9.93 | 10.44 | 0.07 | 0.05 | 0.08 | 0.0 | Detail |
| 1741 | CHOL | autosis | INPP5E | inositol polyphosphate-5-phosphatase E | 7.96 | 8.47 | 0.09 | 0.07 | 0.12 | 0.0 | Detail |
| 1742 | CHOL | autosis | KIAA1549L | KIAA1549 like | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1743 | CHOL | autosis | KIZ | kizuna centrosomal protein | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1744 | CHOL | autosis | KPNA5 | karyopherin subunit alpha 5 | 4.26 | 4.92 | 0.21 | 0.07 | 0.12 | 0.0 | Detail |
| 1745 | CHOL | autosis | LCMT2 | leucine carboxyl methyltransferase 2 | 7.94 | 7.98 | 0.01 | 0.86 | 0.89 | 0.0 | Detail |
| 1746 | CHOL | autosis | LIN37 | lin-37 DREAM MuvB core complex component | 7.11 | 7.35 | 0.05 | 0.45 | 0.53 | 0.0 | Detail |
| 1747 | CHOL | autosis | LINC01963 | long intergenic non-protein coding RNA 1963 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1748 | CHOL | autosis | LYST | lysosomal trafficking regulator | 8.84 | 8.14 | -0.12 | 0.14 | 0.21 | 0.0 | Detail |
| 1749 | CHOL | autosis | MANSC1 | MANSC domain containing 1 | 8.26 | 9.63 | 0.22 | 0.03 | 0.05 | 0.0 | Detail |
| 1750 | CHOL | autosis | MAP4K2 | mitogen-activated protein kinase kinase kinase kinase 2 | 7.64 | 8.29 | 0.12 | 0.05 | 0.08 | 0.0 | Detail |
| 1751 | CHOL | autosis | MDM1 | Mdm1 nuclear protein | 7.1 | 7.44 | 0.07 | 0.08 | 0.13 | 0.0 | Detail |
| 1752 | CHOL | autosis | MEAK7 | MTOR associated protein, eak-7 homolog | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1753 | CHOL | autosis | METTL18 | methyltransferase 18, RPL3 N3(tau)-histidine | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1754 | CHOL | autosis | MRPL57 | mitochondrial ribosomal protein L57 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1755 | CHOL | autosis | MRPS14 | mitochondrial ribosomal protein S14 | 8.45 | 8.64 | 0.03 | 0.42 | 0.5 | 0.0 | Detail |
| 1756 | CHOL | autosis | MXD4 | MAX dimerization protein 4 | 10.34 | 10.72 | 0.05 | 0.08 | 0.13 | 0.0 | Detail |
| 1757 | CHOL | autosis | MYT1 | myelin transcription factor 1 | 0.84 | 1.64 | 0.97 | 0.17 | 0.24 | 0.0 | Detail |
| 1758 | CHOL | autosis | NDUFA3 | NADH:ubiquinone oxidoreductase subunit A3 | 9.61 | 9.18 | -0.07 | 0.33 | 0.41 | 0.0 | Detail |
| 1759 | CHOL | autosis | NDUFA7 | NADH:ubiquinone oxidoreductase subunit A7 | 9.85 | 9.71 | -0.02 | 0.36 | 0.44 | 0.0 | Detail |
| 1760 | CHOL | autosis | NDUFAF7 | NADH:ubiquinone oxidoreductase complex assembly factor 7 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1761 | CHOL | autosis | NFRKB | nuclear factor related to kappaB binding protein | 8.81 | 9.17 | 0.06 | 0.04 | 0.07 | 0.0 | Detail |
| 1762 | CHOL | autosis | NPIPA1 | nuclear pore complex interacting protein family member A1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1763 | CHOL | autosis | NUDT19 | nudix hydrolase 19 | 8.35 | 8.57 | 0.04 | 0.39 | 0.47 | 0.0 | Detail |
| 1764 | CHOL | autosis | NUDT4 | nudix hydrolase 4 | 9.52 | 9.39 | -0.02 | 0.38 | 0.46 | 0.0 | Detail |
| 1765 | CHOL | autosis | NUP50 | nucleoporin 50 | 10.09 | 10.25 | 0.02 | 0.28 | 0.36 | 0.0 | Detail |
| 1766 | CHOL | autosis | ORC3 | origin recognition complex subunit 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1767 | CHOL | autosis | ORC4 | origin recognition complex subunit 4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1768 | CHOL | autosis | ORC5 | origin recognition complex subunit 5 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1769 | CHOL | autosis | PELO | pelota mRNA surveillance and ribosome rescue factor | 8.67 | 9.19 | 0.08 | 0.03 | 0.06 | 0.0 | Detail |
| 1770 | CHOL | autosis | PEX12 | peroxisomal biogenesis factor 12 | 8.02 | 7.56 | -0.09 | 0.04 | 0.06 | 0.0 | Detail |
| 1771 | CHOL | autosis | PHF3 | PHD finger protein 3 | 10.75 | 10.41 | -0.05 | 0.36 | 0.44 | 0.0 | Detail |
| 1772 | CHOL | autosis | PLCXD1 | phosphatidylinositol specific phospholipase C X domain containing 1 | 9.54 | 9.02 | -0.08 | 0.16 | 0.23 | 0.0 | Detail |
| 1773 | CHOL | autosis | PLEKHO2 | pleckstrin homology domain containing O2 | 8.92 | 9.47 | 0.09 | 0.55 | 0.63 | 0.0 | Detail |
| 1774 | CHOL | autosis | PLGRKT | plasminogen receptor with a C-terminal lysine | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1775 | CHOL | autosis | PNISR | PNN interacting serine and arginine rich protein | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1776 | CHOL | autosis | PNPLA4 | patatin like phospholipase domain containing 4 | 9.37 | 8.83 | -0.09 | 0.28 | 0.36 | 0.0 | Detail |
| 1777 | CHOL | autosis | POGLUT2 | protein O-glucosyltransferase 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1778 | CHOL | autosis | POLR1B | RNA polymerase I subunit B | 8.88 | 9.0 | 0.02 | 0.58 | 0.66 | 0.0 | Detail |
| 1779 | CHOL | autosis | PRRC2B | proline rich coiled-coil 2B | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1780 | CHOL | autosis | PYCR3 | pyrroline-5-carboxylate reductase 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1781 | CHOL | autosis | RAB8B | RAB8B, member RAS oncogene family | 8.75 | 9.31 | 0.09 | 0.06 | 0.1 | 0.0 | Detail |
| 1782 | CHOL | autosis | RABGGTB | Rab geranylgeranyltransferase subunit beta | 10.1 | 10.57 | 0.06 | 0.04 | 0.07 | 0.0 | Detail |
| 1783 | CHOL | autosis | RANBP6 | RAN binding protein 6 | 9.27 | 9.04 | -0.04 | 0.18 | 0.24 | 0.0 | Detail |
| 1784 | CHOL | autosis | RGS7 | regulator of G protein signaling 7 | 1.52 | 1.39 | -0.13 | 0.89 | 0.92 | 0.0 | Detail |
| 1785 | CHOL | autosis | RNASE6 | ribonuclease A family member k6 | 6.88 | 7.53 | 0.13 | 0.26 | 0.34 | 0.0 | Detail |
| 1786 | CHOL | autosis | RNASEH1P1 | ribonuclease H1 pseudogene 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1787 | CHOL | autosis | RP2 | RP2 activator of ARL3 GTPase | 8.71 | 8.36 | -0.06 | 0.36 | 0.45 | 0.0 | Detail |
| 1788 | CHOL | autosis | RTL6 | retrotransposon Gag like 6 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1789 | CHOL | autosis | RWDD2A | RWD domain containing 2A | 6.47 | 6.78 | 0.07 | 0.36 | 0.44 | 0.0 | Detail |
| 1790 | CHOL | autosis | SCGB1D1 | secretoglobin family 1D member 1 | 0 | 0 | NA | NA | NA | 0 | Detail |
| 1791 | CHOL | autosis | SDHAF3 | succinate dehydrogenase complex assembly factor 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1792 | CHOL | autosis | SLC30A5 | solute carrier family 30 member 5 | 9.65 | 9.97 | 0.05 | 0.1 | 0.16 | 0.0 | Detail |
| 1793 | CHOL | autosis | SLC35B3 | solute carrier family 35 member B3 | 9.19 | 9.22 | 0.01 | 1.0 | 1.0 | 0.0 | Detail |
| 1794 | CHOL | autosis | SLF2 | SMC5-SMC6 complex localization factor 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1795 | CHOL | autosis | SMARCD2 | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 | 11.07 | 11.28 | 0.03 | 0.31 | 0.39 | 0.0 | Detail |
| 1796 | CHOL | autosis | SMCO4 | single-pass membrane protein with coiled-coil domains 4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1797 | CHOL | autosis | SMIM29 | small integral membrane protein 29 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1798 | CHOL | autosis | SNX16 | sorting nexin 16 | 6.89 | 6.83 | -0.01 | 0.98 | 0.98 | 0.0 | Detail |
| 1799 | CHOL | autosis | SREK1 | splicing regulatory glutamic acid and lysine rich protein 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1800 | CHOL | autosis | STK19B | serine/threonine kinase 19B (pseudogene) | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1801 | CHOL | autosis | STX5 | syntaxin 5 | 10.07 | 9.95 | -0.02 | 0.48 | 0.56 | 0.0 | Detail |
| 1802 | CHOL | autosis | STX8 | syntaxin 8 | 8.13 | 8.41 | 0.05 | 0.21 | 0.29 | 0.0 | Detail |
| 1803 | CHOL | autosis | SURF2 | surfeit 2 | 8.13 | 8.24 | 0.02 | 0.94 | 0.95 | 0.0 | Detail |
| 1804 | CHOL | autosis | SYNC | syncoilin, intermediate filament protein | 4.75 | 6.24 | 0.39 | 0.09 | 0.14 | 0.0 | Detail |
| 1805 | CHOL | autosis | TBC1D15 | TBC1 domain family member 15 | 10.09 | 9.78 | -0.05 | 0.09 | 0.14 | 0.0 | Detail |
| 1806 | CHOL | autosis | TCEA1P2 | transcription elongation factor A1 pseudogene 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1807 | CHOL | autosis | TEX261 | testis expressed 261 | 11.03 | 11.12 | 0.01 | 0.58 | 0.66 | 0.0 | Detail |
| 1808 | CHOL | autosis | THTPA | thiamine triphosphatase | 8.0 | 8.19 | 0.03 | 1.0 | 1.0 | 0.0 | Detail |
| 1809 | CHOL | autosis | TMEM177 | transmembrane protein 177 | 8.72 | 8.22 | -0.09 | 0.13 | 0.19 | 0.0 | Detail |
| 1810 | CHOL | autosis | TMEM251 | lysosomal enzyme trafficking factor | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1811 | CHOL | autosis | TMEM39A | transmembrane protein 39A | 8.4 | 8.87 | 0.08 | 0.04 | 0.06 | 0.0 | Detail |
| 1812 | CHOL | autosis | TMX4 | thioredoxin related transmembrane protein 4 | 9.51 | 9.39 | -0.02 | 0.66 | 0.72 | 0.0 | Detail |
| 1813 | CHOL | autosis | TVP23B | trans-golgi network vesicle protein 23 homolog B | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1814 | CHOL | autosis | UROS | uroporphyrinogen III synthase | 10.19 | 9.4 | -0.12 | 0.03 | 0.05 | 0.0 | Detail |
| 1815 | CHOL | autosis | UTP3 | UTP3 small subunit processome component | 9.48 | 9.34 | -0.02 | 0.58 | 0.66 | 0.0 | Detail |
| 1816 | CHOL | autosis | VSIG2 | V-set and immunoglobulin domain containing 2 | 6.74 | 5.05 | -0.42 | 0.13 | 0.19 | 0.0 | Detail |
| 1817 | CHOL | autosis | WFS1 | wolframin ER transmembrane glycoprotein | 9.11 | 9.5 | 0.06 | 0.07 | 0.12 | 0.0 | Detail |
| 1818 | CHOL | autosis | YRDC | yrdC N6-threonylcarbamoyltransferase domain containing | 8.29 | 8.55 | 0.04 | 0.24 | 0.31 | 0.0 | Detail |
| 1819 | CHOL | autosis | ZBTB14 | zinc finger and BTB domain containing 14 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1820 | CHOL | autosis | ZDHHC4 | zinc finger DHHC-type palmitoyltransferase 4 | 9.75 | 10.15 | 0.06 | 0.05 | 0.08 | 0.0 | Detail |
| 1821 | CHOL | autosis | ZNF112 | zinc finger protein 112 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1822 | CHOL | autosis | ZNF254 | zinc finger protein 254 | 8.26 | 8.0 | -0.05 | 0.24 | 0.31 | 0.0 | Detail |
| 1823 | CHOL | autosis | ZNF274 | zinc finger protein 274 | 9.36 | 8.8 | -0.09 | 0.14 | 0.21 | 0.0 | Detail |
| 1824 | CHOL | autosis | ZNF322P1 | zinc finger protein 322 pseudogene 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 1825 | CHOL | autosis | ZNF426 | zinc finger protein 426 | 4.94 | 5.46 | 0.14 | 0.31 | 0.39 | 0.0 | Detail |
| 1826 | CHOL | autosis | ZNF512B | zinc finger protein 512B | 9.56 | 9.86 | 0.04 | 0.19 | 0.26 | 0.0 | Detail |
Ensemble learning visualization:
Ensemble learning for CHOL based on specific autosis gene regulaors.