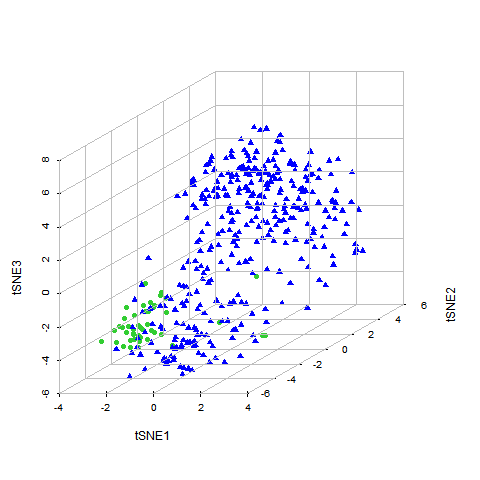
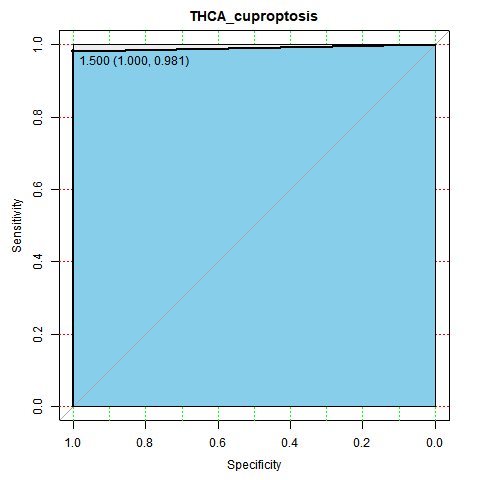
Disease specific cell death gene regulator search
Search Results:
THCA specific cuproptosis gene regulaors.
Different color in column1 means high , low , non-diff expressed gene in disease.
| NO. | Disease | CellDeathType | Genesymbol | Fullname | Control_mean | Disease_mean | log2FC | pvalue | padj | Weight | Detail |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | THCA | cuproptosis | BCL2 | BCL2 apoptosis regulator | 11.87 | 10.09 | -0.23 | 0.0 | 0.0 | 100.0 | Detail |
| 2 | THCA | cuproptosis | AP1S1 | adaptor related protein complex 1 subunit sigma 1 | 9.14 | 9.86 | 0.11 | 0.0 | 0.0 | 98.37 | Detail |
| 3 | THCA | cuproptosis | LOXL2 | lysyl oxidase like 2 | 6.73 | 8.42 | 0.32 | 0.0 | 0.0 | 92.26 | Detail |
| 4 | THCA | cuproptosis | ZKSCAN5 | zinc finger with KRAB and SCAN domains 5 | 8.68 | 8.19 | -0.08 | 0.0 | 0.0 | 90.87 | Detail |
| 5 | THCA | cuproptosis | GSS | glutathione synthetase | 9.68 | 10.27 | 0.09 | 0.0 | 0.0 | 89.65 | Detail |
| 6 | THCA | cuproptosis | ATP6AP1 | ATPase H+ transporting accessory protein 1 | 11.92 | 12.43 | 0.06 | 0.0 | 0.0 | 89.48 | Detail |
| 7 | THCA | cuproptosis | CENPJ | centromere protein J | 8.96 | 7.25 | -0.31 | 0.0 | 0.0 | 88.35 | Detail |
| 8 | THCA | cuproptosis | RAD23B | RAD23 homolog B, nucleotide excision repair protein | 11.53 | 12.48 | 0.11 | 0.0 | 0.0 | 88.25 | Detail |
| 9 | THCA | cuproptosis | ISCA1 | iron-sulfur cluster assembly 1 | 9.93 | 9.3 | -0.09 | 0.0 | 0.0 | 85.62 | Detail |
| 10 | THCA | cuproptosis | SPARC | secreted protein acidic and cysteine rich | 13.62 | 14.48 | 0.09 | 0.0 | 0.0 | 84.88 | Detail |
| 11 | THCA | cuproptosis | SLC33A1 | solute carrier family 33 member 1 | 9.96 | 9.1 | -0.13 | 0.0 | 0.0 | 83.6 | Detail |
| 12 | THCA | cuproptosis | CDKN2A | cyclin dependent kinase inhibitor 2A | 3.06 | 5.51 | 0.85 | 0.0 | 0.0 | 83.29 | Detail |
| 13 | THCA | cuproptosis | MTF1 | metal regulatory transcription factor 1 | 9.1 | 8.48 | -0.1 | 0.0 | 0.0 | 83.06 | Detail |
| 14 | THCA | cuproptosis | UBE2I | ubiquitin conjugating enzyme E2 I | 10.59 | 11.09 | 0.07 | 0.0 | 0.0 | 82.87 | Detail |
| 15 | THCA | cuproptosis | MAD2L2 | mitotic arrest deficient 2 like 2 | 7.92 | 9.02 | 0.19 | 0.0 | 0.0 | 82.65 | Detail |
| 16 | THCA | cuproptosis | FASTKD5 | FAST kinase domains 5 | 8.74 | 8.23 | -0.09 | 0.0 | 0.0 | 82.27 | Detail |
| 17 | THCA | cuproptosis | SERPIND1 | serpin family D member 1 | 4.4 | 1.69 | -1.38 | 0.0 | 0.0 | 81.47 | Detail |
| 18 | THCA | cuproptosis | SOD3 | superoxide dismutase 3 | 11.76 | 9.51 | -0.31 | 0.0 | 0.0 | 80.97 | Detail |
| 19 | THCA | cuproptosis | SEC14L2 | SEC14 like lipid binding 2 | 8.29 | 9.38 | 0.18 | 0.0 | 0.0 | 80.57 | Detail |
| 20 | THCA | cuproptosis | SNCA | synuclein alpha | 7.35 | 5.99 | -0.29 | 0.0 | 0.0 | 80.13 | Detail |
| 21 | THCA | cuproptosis | ISCU | iron-sulfur cluster assembly enzyme | 11.59 | 12.1 | 0.06 | 0.0 | 0.0 | 79.3 | Detail |
| 22 | THCA | cuproptosis | SDHD | succinate dehydrogenase complex subunit D | 11.34 | 10.79 | -0.07 | 0.0 | 0.0 | 79.17 | Detail |
| 23 | THCA | cuproptosis | ABCB10 | ATP binding cassette subfamily B member 10 | 8.61 | 8.05 | -0.1 | 0.0 | 0.0 | 79.1 | Detail |
| 24 | THCA | cuproptosis | FPGS | folylpolyglutamate synthase | 9.72 | 10.26 | 0.08 | 0.0 | 0.0 | 79.02 | Detail |
| 25 | THCA | cuproptosis | ATIC | 5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase | 9.79 | 10.61 | 0.12 | 0.0 | 0.0 | 78.81 | Detail |
| 26 | THCA | cuproptosis | STEAP2 | STEAP2 metalloreductase | 8.36 | 6.67 | -0.32 | 0.0 | 0.0 | 78.15 | Detail |
| 27 | THCA | cuproptosis | ISCA2 | iron-sulfur cluster assembly 2 | 9.2 | 9.75 | 0.08 | 0.0 | 0.0 | 77.68 | Detail |
| 28 | THCA | cuproptosis | MTFR1 | mitochondrial fission regulator 1 | 8.37 | 7.76 | -0.11 | 0.0 | 0.0 | 77.55 | Detail |
| 29 | THCA | cuproptosis | MRPS14 | mitochondrial ribosomal protein S14 | 8.67 | 8.29 | -0.06 | 0.0 | 0.0 | 76.35 | Detail |
| 30 | THCA | cuproptosis | OGDHL | oxoglutarate dehydrogenase L | 10.6 | 8.35 | -0.34 | 0.0 | 0.0 | 76.08 | Detail |
| 31 | THCA | cuproptosis | GPX4 | glutathione peroxidase 4 | 11.6 | 12.39 | 0.1 | 0.0 | 0.0 | 76.04 | Detail |
| 32 | THCA | cuproptosis | FZD8 | frizzled class receptor 8 | 8.92 | 7.78 | -0.2 | 0.0 | 0.0 | 75.36 | Detail |
| 33 | THCA | cuproptosis | NFE2L2 | NFE2 like bZIP transcription factor 2 | 12.22 | 11.74 | -0.06 | 0.0 | 0.0 | 75.06 | Detail |
| 34 | THCA | cuproptosis | FARS2 | phenylalanyl-tRNA synthetase 2, mitochondrial | 8.46 | 7.96 | -0.09 | 0.0 | 0.0 | 74.13 | Detail |
| 35 | THCA | cuproptosis | FADD | Fas associated via death domain | 8.78 | 9.3 | 0.08 | 0.0 | 0.0 | 74.06 | Detail |
| 36 | THCA | cuproptosis | GCSH | glycine cleavage system protein H | 6.95 | 5.54 | -0.33 | 0.0 | 0.0 | 73.79 | Detail |
| 37 | THCA | cuproptosis | DAPK2 | death associated protein kinase 2 | 10.67 | 12.07 | 0.18 | 0.0 | 0.0 | 73.54 | Detail |
| 38 | THCA | cuproptosis | SLFN11 | schlafen family member 11 | 10.14 | 9.34 | -0.12 | 0.0 | 0.0 | 73.47 | Detail |
| 39 | THCA | cuproptosis | IDH1 | isocitrate dehydrogenase (NADP(+)) 1 | 10.12 | 10.71 | 0.08 | 0.0 | 0.0 | 73.26 | Detail |
| 40 | THCA | cuproptosis | MRPL14 | mitochondrial ribosomal protein L14 | 9.0 | 9.78 | 0.12 | 0.0 | 0.0 | 73.09 | Detail |
| 41 | THCA | cuproptosis | ATP7B | ATPase copper transporting beta | 6.78 | 5.77 | -0.23 | 0.0 | 0.0 | 72.6 | Detail |
| 42 | THCA | cuproptosis | FDX1 | ferredoxin 1 | 10.07 | 9.49 | -0.09 | 0.0 | 0.0 | 72.1 | Detail |
| 43 | THCA | cuproptosis | AQP1 | aquaporin 1 (Colton blood group) | 10.97 | 12.04 | 0.13 | 0.0 | 0.0 | 71.73 | Detail |
| 44 | THCA | cuproptosis | TSC22D2 | TSC22 domain family member 2 | 9.5 | 8.72 | -0.12 | 0.0 | 0.0 | 71.33 | Detail |
| 45 | THCA | cuproptosis | GFRA3 | GDNF family receptor alpha 3 | 2.29 | 1.04 | -1.15 | 0.0 | 0.0 | 71.22 | Detail |
| 46 | THCA | cuproptosis | MT1M | metallothionein 1M | 7.75 | 5.43 | -0.51 | 0.0 | 0.0 | 71.12 | Detail |
| 47 | THCA | cuproptosis | PGK1 | phosphoglycerate kinase 1 | 12.66 | 12.22 | -0.05 | 0.0 | 0.0 | 71.01 | Detail |
| 48 | THCA | cuproptosis | RPUSD3 | RNA pseudouridine synthase D3 | 8.48 | 8.94 | 0.08 | 0.0 | 0.0 | 70.85 | Detail |
| 49 | THCA | cuproptosis | EIF3I | eukaryotic translation initiation factor 3 subunit I | 11.74 | 12.1 | 0.04 | 0.0 | 0.0 | 70.78 | Detail |
| 50 | THCA | cuproptosis | ABCB6 | ATP binding cassette subfamily B member 6 (LAN blood group) | 9.4 | 8.7 | -0.11 | 0.0 | 0.0 | 70.76 | Detail |
| 51 | THCA | cuproptosis | SPATA13 | spermatogenesis associated 13 | 9.08 | 7.7 | -0.24 | 0.0 | 0.0 | 70.32 | Detail |
| 52 | THCA | cuproptosis | TADA2A | transcriptional adaptor 2A | 7.98 | 7.49 | -0.09 | 0.0 | 0.0 | 70.17 | Detail |
| 53 | THCA | cuproptosis | TLR3 | toll like receptor 3 | 7.77 | 6.71 | -0.21 | 0.0 | 0.0 | 69.33 | Detail |
| 54 | THCA | cuproptosis | UBE2D3 | ubiquitin conjugating enzyme E2 D3 | 12.5 | 12.28 | -0.03 | 0.0 | 0.0 | 68.76 | Detail |
| 55 | THCA | cuproptosis | NNAT | neuronatin | 3.29 | 2.16 | -0.61 | 0.0 | 0.0 | 68.33 | Detail |
| 56 | THCA | cuproptosis | MT1F | metallothionein 1F | 12.2 | 9.57 | -0.35 | 0.0 | 0.0 | 67.72 | Detail |
| 57 | THCA | cuproptosis | COQ7 | coenzyme Q7, hydroxylase | 9.25 | 8.93 | -0.05 | 0.0 | 0.0 | 67.46 | Detail |
| 58 | THCA | cuproptosis | SHMT2 | serine hydroxymethyltransferase 2 | 9.4 | 10.09 | 0.1 | 0.0 | 0.0 | 67.16 | Detail |
| 59 | THCA | cuproptosis | BECN1 | beclin 1 | 10.37 | 10.6 | 0.03 | 0.0 | 0.0 | 66.69 | Detail |
| 60 | THCA | cuproptosis | MT1A | metallothionein 1A | 4.8 | 2.98 | -0.69 | 0.0 | 0.0 | 66.68 | Detail |
| 61 | THCA | cuproptosis | CYCS | cytochrome c, somatic | 11.33 | 10.7 | -0.08 | 0.0 | 0.0 | 66.44 | Detail |
| 62 | THCA | cuproptosis | MT1H | metallothionein 1H | 9.13 | 5.13 | -0.83 | 0.0 | 0.0 | 66.21 | Detail |
| 63 | THCA | cuproptosis | MRPL17 | mitochondrial ribosomal protein L17 | 9.83 | 10.35 | 0.07 | 0.0 | 0.0 | 65.93 | Detail |
| 64 | THCA | cuproptosis | TANK | TRAF family member associated NFKB activator | 9.74 | 10.12 | 0.06 | 0.0 | 0.0 | 65.76 | Detail |
| 65 | THCA | cuproptosis | NUBPL | NUBP iron-sulfur cluster assembly factor, mitochondrial | 8.25 | 7.84 | -0.07 | 0.0 | 0.0 | 65.0 | Detail |
| 66 | THCA | cuproptosis | MT1DP | metallothionein 1D, pseudogene | 3.86 | 2.32 | -0.73 | 0.0 | 0.0 | 64.99 | Detail |
| 67 | THCA | cuproptosis | COX15 | cytochrome c oxidase assembly homolog COX15 | 9.98 | 9.68 | -0.05 | 0.0 | 0.0 | 64.68 | Detail |
| 68 | THCA | cuproptosis | MAP1LC3A | microtubule associated protein 1 light chain 3 alpha | 9.25 | 8.29 | -0.16 | 0.0 | 0.0 | 64.29 | Detail |
| 69 | THCA | cuproptosis | LONP1 | lon peptidase 1, mitochondrial | 10.62 | 11.09 | 0.06 | 0.0 | 0.0 | 63.57 | Detail |
| 70 | THCA | cuproptosis | ATM | ATM serine/threonine kinase | 10.56 | 9.96 | -0.08 | 0.0 | 0.0 | 63.33 | Detail |
| 71 | THCA | cuproptosis | PTCD3 | pentatricopeptide repeat domain 3 | 9.92 | 9.67 | -0.04 | 0.0 | 0.0 | 62.8 | Detail |
| 72 | THCA | cuproptosis | ELF3 | E74 like ETS transcription factor 3 | 8.13 | 9.77 | 0.27 | 0.0 | 0.0 | 61.71 | Detail |
| 73 | THCA | cuproptosis | MT1G | metallothionein 1G | 12.71 | 8.87 | -0.52 | 0.0 | 0.0 | 61.51 | Detail |
| 74 | THCA | cuproptosis | HEPHL1 | hephaestin like 1 | 2.97 | 1.64 | -0.86 | 0.0 | 0.0 | 60.38 | Detail |
| 75 | THCA | cuproptosis | DOT1L | DOT1 like histone lysine methyltransferase | 9.3 | 8.83 | -0.07 | 0.0 | 0.0 | 60.37 | Detail |
| 76 | THCA | cuproptosis | MRPS25 | mitochondrial ribosomal protein S25 | 10.21 | 10.71 | 0.07 | 0.0 | 0.0 | 59.98 | Detail |
| 77 | THCA | cuproptosis | DYNC1LI2 | dynein cytoplasmic 1 light intermediate chain 2 | 11.81 | 11.57 | -0.03 | 0.0 | 0.0 | 59.92 | Detail |
| 78 | THCA | cuproptosis | FDXR | ferredoxin reductase | 8.13 | 8.79 | 0.11 | 0.0 | 0.0 | 59.0 | Detail |
| 79 | THCA | cuproptosis | CDC27 | cell division cycle 27 | 11.32 | 10.91 | -0.05 | 0.0 | 0.0 | 58.94 | Detail |
| 80 | THCA | cuproptosis | AFG3L2 | AFG3 like matrix AAA peptidase subunit 2 | 10.81 | 10.38 | -0.06 | 0.0 | 0.0 | 58.74 | Detail |
| 81 | THCA | cuproptosis | NFS1 | NFS1 cysteine desulfurase | 8.89 | 8.61 | -0.05 | 0.0 | 0.0 | 58.56 | Detail |
| 82 | THCA | cuproptosis | LIPT2 | lipoyl(octanoyl) transferase 2 | 5.38 | 4.8 | -0.17 | 0.0 | 0.0 | 57.3 | Detail |
| 83 | THCA | cuproptosis | CIAO1 | cytosolic iron-sulfur assembly component 1 | 10.82 | 10.97 | 0.02 | 0.0 | 0.0 | 57.12 | Detail |
| 84 | THCA | cuproptosis | STARD7 | StAR related lipid transfer domain containing 7 | 11.89 | 11.55 | -0.04 | 0.0 | 0.0 | 56.32 | Detail |
| 85 | THCA | cuproptosis | LIPT1 | lipoyltransferase 1 | 7.77 | 7.37 | -0.08 | 0.0 | 0.0 | 56.27 | Detail |
| 86 | THCA | cuproptosis | STEAP4 | STEAP4 metalloreductase | 9.19 | 10.02 | 0.12 | 0.0 | 0.0 | 56.26 | Detail |
| 87 | THCA | cuproptosis | HSF1 | heat shock transcription factor 1 | 10.62 | 10.91 | 0.04 | 0.0 | 0.0 | 55.67 | Detail |
| 88 | THCA | cuproptosis | CP | ceruloplasmin | 7.64 | 5.43 | -0.49 | 0.0 | 0.0 | 55.56 | Detail |
| 89 | THCA | cuproptosis | DBR1 | debranching RNA lariats 1 | 8.52 | 8.25 | -0.05 | 0.0 | 0.0 | 55.43 | Detail |
| 90 | THCA | cuproptosis | POT1 | protection of telomeres 1 | 9.25 | 8.96 | -0.05 | 0.0 | 0.0 | 55.25 | Detail |
| 91 | THCA | cuproptosis | DNTTIP2 | deoxynucleotidyltransferase terminal interacting protein 2 | 10.38 | 10.14 | -0.03 | 0.0 | 0.0 | 55.07 | Detail |
| 92 | THCA | cuproptosis | FXN | frataxin | 8.02 | 7.68 | -0.06 | 0.0 | 0.0 | 54.85 | Detail |
| 93 | THCA | cuproptosis | NFKBIA | NFKB inhibitor alpha | 11.59 | 10.82 | -0.1 | 0.0 | 0.0 | 54.64 | Detail |
| 94 | THCA | cuproptosis | SPDYE1 | speedy/RINGO cell cycle regulator family member E1 | 3.42 | 2.8 | -0.29 | 0.0 | 0.0 | 54.43 | Detail |
| 95 | THCA | cuproptosis | DBT | dihydrolipoamide branched chain transacylase E2 | 9.78 | 9.37 | -0.06 | 0.0 | 0.0 | 54.39 | Detail |
| 96 | THCA | cuproptosis | SLC25A32 | solute carrier family 25 member 32 | 8.69 | 8.34 | -0.06 | 0.0 | 0.0 | 54.27 | Detail |
| 97 | THCA | cuproptosis | ETFDH | electron transfer flavoprotein dehydrogenase | 8.7 | 8.34 | -0.06 | 0.0 | 0.0 | 54.17 | Detail |
| 98 | THCA | cuproptosis | RIPK1 | receptor interacting serine/threonine kinase 1 | 9.97 | 9.73 | -0.04 | 0.0 | 0.0 | 54.06 | Detail |
| 99 | THCA | cuproptosis | FKBP4 | FKBP prolyl isomerase 4 | 11.69 | 11.26 | -0.05 | 0.0 | 0.0 | 53.94 | Detail |
| 100 | THCA | cuproptosis | RHOV | ras homolog family member V | 4.25 | 5.65 | 0.41 | 0.0 | 0.0 | 53.77 | Detail |
| 101 | THCA | cuproptosis | MYO1B | myosin IB | 9.62 | 10.18 | 0.08 | 0.0 | 0.0 | 53.55 | Detail |
| 102 | THCA | cuproptosis | DLD | dihydrolipoamide dehydrogenase | 10.67 | 10.36 | -0.04 | 0.0 | 0.0 | 52.7 | Detail |
| 103 | THCA | cuproptosis | RPAP1 | RNA polymerase II associated protein 1 | 9.14 | 8.85 | -0.05 | 0.0 | 0.0 | 51.84 | Detail |
| 104 | THCA | cuproptosis | RPL28 | ribosomal protein L28 | 13.34 | 13.86 | 0.06 | 0.0 | 0.0 | 51.49 | Detail |
| 105 | THCA | cuproptosis | PLOD1 | procollagen-lysine,2-oxoglutarate 5-dioxygenase 1 | 11.05 | 11.44 | 0.05 | 0.0 | 0.0 | 50.7 | Detail |
| 106 | THCA | cuproptosis | TP53 | tumor protein p53 | 9.71 | 10.05 | 0.05 | 0.0 | 0.0 | 50.54 | Detail |
| 107 | THCA | cuproptosis | GCLC | glutamate-cysteine ligase catalytic subunit | 10.17 | 9.7 | -0.07 | 0.0 | 0.0 | 50.42 | Detail |
| 108 | THCA | cuproptosis | STEAP3 | STEAP3 metalloreductase | 8.12 | 8.92 | 0.13 | 0.0 | 0.0 | 50.21 | Detail |
| 109 | THCA | cuproptosis | DDX20 | DEAD-box helicase 20 | 8.63 | 8.36 | -0.05 | 0.0 | 0.0 | 49.44 | Detail |
| 110 | THCA | cuproptosis | MADD | MAP kinase activating death domain | 10.05 | 9.82 | -0.03 | 0.0 | 0.0 | 49.42 | Detail |
| 111 | THCA | cuproptosis | PPP1R15A | protein phosphatase 1 regulatory subunit 15A | 11.79 | 10.72 | -0.14 | 0.0 | 0.0 | 49.32 | Detail |
| 112 | THCA | cuproptosis | CCDC22 | coiled-coil domain containing 22 | 9.0 | 9.25 | 0.04 | 0.0 | 0.0 | 49.17 | Detail |
| 113 | THCA | cuproptosis | RPL7A | ribosomal protein L7a | 13.82 | 14.17 | 0.04 | 0.0 | 0.0 | 48.81 | Detail |
| 114 | THCA | cuproptosis | QRSL1 | glutaminyl-tRNA amidotransferase subunit QRSL1 | 9.12 | 8.86 | -0.04 | 0.0 | 0.0 | 48.79 | Detail |
| 115 | THCA | cuproptosis | CCL20 | C-C motif chemokine ligand 20 | 1.78 | 3.84 | 1.11 | 0.0 | 0.0 | 47.95 | Detail |
| 116 | THCA | cuproptosis | HARS2 | histidyl-tRNA synthetase 2, mitochondrial | 9.17 | 9.39 | 0.03 | 0.0 | 0.0 | 47.79 | Detail |
| 117 | THCA | cuproptosis | SNAPIN | SNAP associated protein | 9.26 | 9.47 | 0.03 | 0.0 | 0.0 | 47.4 | Detail |
| 118 | THCA | cuproptosis | HNRNPA2B1 | heterogeneous nuclear ribonucleoprotein A2/B1 | 13.77 | 13.54 | -0.02 | 0.0 | 0.0 | 47.29 | Detail |
| 119 | THCA | cuproptosis | DTYMK | deoxythymidylate kinase | 7.94 | 8.26 | 0.06 | 0.0 | 0.0 | 47.06 | Detail |
| 120 | THCA | cuproptosis | SDSL | serine dehydratase like | 9.9 | 9.35 | -0.08 | 0.0 | 0.0 | 46.94 | Detail |
| 121 | THCA | cuproptosis | DNM1L | dynamin 1 like | 10.31 | 10.12 | -0.03 | 0.0 | 0.0 | 46.87 | Detail |
| 122 | THCA | cuproptosis | SLC30A10 | solute carrier family 30 member 10 | 0.94 | 0.42 | -1.17 | 0.0 | 0.0 | 46.73 | Detail |
| 123 | THCA | cuproptosis | TKT | transketolase | 12.61 | 13.03 | 0.05 | 0.0 | 0.0 | 46.61 | Detail |
| 124 | THCA | cuproptosis | COQ2 | coenzyme Q2, polyprenyltransferase | 7.61 | 7.91 | 0.06 | 0.0 | 0.0 | 46.0 | Detail |
| 125 | THCA | cuproptosis | DHX15 | DEAH-box helicase 15 | 11.04 | 10.78 | -0.03 | 0.0 | 0.0 | 45.97 | Detail |
| 126 | THCA | cuproptosis | PRDX1 | peroxiredoxin 1 | 13.66 | 13.2 | -0.05 | 0.0 | 0.0 | 45.85 | Detail |
| 127 | THCA | cuproptosis | FAM111A | FAM111 trypsin like peptidase A | 9.77 | 10.26 | 0.07 | 0.0 | 0.0 | 45.15 | Detail |
| 128 | THCA | cuproptosis | MUC20 | mucin 20, cell surface associated | 7.72 | 8.6 | 0.16 | 0.0 | 0.0 | 45.08 | Detail |
| 129 | THCA | cuproptosis | MECR | mitochondrial trans-2-enoyl-CoA reductase | 8.3 | 8.57 | 0.05 | 0.0 | 0.0 | 44.08 | Detail |
| 130 | THCA | cuproptosis | ACO1 | aconitase 1 | 10.49 | 11.04 | 0.07 | 0.0 | 0.0 | 43.99 | Detail |
| 131 | THCA | cuproptosis | PRND | prion like protein doppel | 1.31 | 2.65 | 1.02 | 0.0 | 0.0 | 43.61 | Detail |
| 132 | THCA | cuproptosis | UQCRC2 | ubiquinol-cytochrome c reductase core protein 2 | 11.57 | 11.39 | -0.02 | 0.0 | 0.0 | 42.95 | Detail |
| 133 | THCA | cuproptosis | MT1X | metallothionein 1X | 12.04 | 10.99 | -0.13 | 0.0 | 0.0 | 42.86 | Detail |
| 134 | THCA | cuproptosis | SDHC | succinate dehydrogenase complex subunit C | 10.8 | 10.56 | -0.03 | 0.0 | 0.0 | 42.67 | Detail |
| 135 | THCA | cuproptosis | AHR | aryl hydrocarbon receptor | 10.39 | 11.18 | 0.11 | 0.0 | 0.0 | 42.35 | Detail |
| 136 | THCA | cuproptosis | ISG15 | ISG15 ubiquitin like modifier | 8.41 | 9.26 | 0.14 | 0.0 | 0.0 | 42.04 | Detail |
| 137 | THCA | cuproptosis | XRN1 | 5'-3' exoribonuclease 1 | 9.88 | 9.52 | -0.05 | 0.0 | 0.0 | 41.95 | Detail |
| 138 | THCA | cuproptosis | HSD17B10 | hydroxysteroid 17-beta dehydrogenase 10 | 10.66 | 10.27 | -0.05 | 0.0 | 0.0 | 41.83 | Detail |
| 139 | THCA | cuproptosis | SCO1 | synthesis of cytochrome C oxidase 1 | 8.77 | 8.51 | -0.04 | 0.0 | 0.0 | 41.52 | Detail |
| 140 | THCA | cuproptosis | N6AMT1 | N-6 adenine-specific DNA methyltransferase 1 | 8.13 | 7.84 | -0.05 | 0.0 | 0.0 | 41.5 | Detail |
| 141 | THCA | cuproptosis | FAS | Fas cell surface death receptor | 8.37 | 9.16 | 0.13 | 0.0 | 0.0 | 41.49 | Detail |
| 142 | THCA | cuproptosis | CMPK1 | cytidine/uridine monophosphate kinase 1 | 11.91 | 11.69 | -0.03 | 0.0 | 0.0 | 39.9 | Detail |
| 143 | THCA | cuproptosis | FAM96A | cytosolic iron-sulfur assembly component 2A | 9.91 | 10.13 | 0.03 | 0.0 | 0.0 | 39.81 | Detail |
| 144 | THCA | cuproptosis | SARS2 | seryl-tRNA synthetase 2, mitochondrial | 8.77 | 9.05 | 0.05 | 0.0 | 0.0 | 38.89 | Detail |
| 145 | THCA | cuproptosis | CUTC | cutC copper transporter | 8.11 | 8.34 | 0.04 | 0.0 | 0.0 | 38.74 | Detail |
| 146 | THCA | cuproptosis | MT1E | metallothionein 1E | 11.64 | 10.56 | -0.14 | 0.0 | 0.0 | 38.33 | Detail |
| 147 | THCA | cuproptosis | PHF5A | PHD finger protein 5A | 8.74 | 8.47 | -0.04 | 0.0 | 0.0 | 37.76 | Detail |
| 148 | THCA | cuproptosis | ELP3 | elongator acetyltransferase complex subunit 3 | 9.73 | 9.54 | -0.03 | 0.0 | 0.0 | 37.26 | Detail |
| 149 | THCA | cuproptosis | PDZD4 | PDZ domain containing 4 | 6.11 | 5.42 | -0.17 | 0.0 | 0.0 | 37.11 | Detail |
| 150 | THCA | cuproptosis | MRPS30 | mitochondrial ribosomal protein S30 | 8.69 | 8.5 | -0.03 | 0.0 | 0.0 | 36.85 | Detail |
| 151 | THCA | cuproptosis | POLD1 | DNA polymerase delta 1, catalytic subunit | 8.76 | 9.11 | 0.06 | 0.0 | 0.0 | 36.66 | Detail |
| 152 | THCA | cuproptosis | UBA2 | ubiquitin like modifier activating enzyme 2 | 10.64 | 10.81 | 0.02 | 0.0 | 0.0 | 36.64 | Detail |
| 153 | THCA | cuproptosis | IDH3B | isocitrate dehydrogenase (NAD(+)) 3 non-catalytic subunit beta | 10.54 | 10.33 | -0.03 | 0.0 | 0.0 | 36.42 | Detail |
| 154 | THCA | cuproptosis | UQCRC1 | ubiquinol-cytochrome c reductase core protein 1 | 11.54 | 11.82 | 0.04 | 0.0 | 0.0 | 36.14 | Detail |
| 155 | THCA | cuproptosis | ANKRD49 | ankyrin repeat domain 49 | 8.63 | 8.31 | -0.05 | 0.0 | 0.0 | 35.9 | Detail |
| 156 | THCA | cuproptosis | EPC2 | enhancer of polycomb homolog 2 | 9.76 | 9.49 | -0.04 | 0.0 | 0.0 | 35.57 | Detail |
| 157 | THCA | cuproptosis | MRPS23 | mitochondrial ribosomal protein S23 | 9.75 | 9.98 | 0.03 | 0.0 | 0.0 | 35.46 | Detail |
| 158 | THCA | cuproptosis | TFAM | transcription factor A, mitochondrial | 9.09 | 8.86 | -0.04 | 0.0 | 0.0 | 35.42 | Detail |
| 159 | THCA | cuproptosis | GFM1 | G elongation factor mitochondrial 1 | 10.14 | 9.88 | -0.04 | 0.0 | 0.0 | 35.39 | Detail |
| 160 | THCA | cuproptosis | NRF1 | nuclear respiratory factor 1 | 8.92 | 8.76 | -0.03 | 0.0 | 0.0 | 35.21 | Detail |
| 161 | THCA | cuproptosis | GADD45GIP1 | GADD45G interacting protein 1 | 9.69 | 10.12 | 0.06 | 0.0 | 0.0 | 34.9 | Detail |
| 162 | THCA | cuproptosis | MRPL51 | mitochondrial ribosomal protein L51 | 10.55 | 10.74 | 0.03 | 0.0 | 0.0 | 34.72 | Detail |
| 163 | THCA | cuproptosis | MRPS22 | mitochondrial ribosomal protein S22 | 8.97 | 8.82 | -0.03 | 0.0 | 0.0 | 34.21 | Detail |
| 164 | THCA | cuproptosis | RANGAP1 | Ran GTPase activating protein 1 | 10.41 | 10.66 | 0.03 | 0.0 | 0.0 | 33.68 | Detail |
| 165 | THCA | cuproptosis | PRDM10 | PR/SET domain 10 | 7.88 | 7.65 | -0.04 | 0.0 | 0.0 | 33.59 | Detail |
| 166 | THCA | cuproptosis | RPS27A | ribosomal protein S27a | 13.34 | 13.04 | -0.03 | 0.0 | 0.0 | 33.59 | Detail |
| 167 | THCA | cuproptosis | ANLN | anillin, actin binding protein | 4.61 | 5.47 | 0.25 | 0.0 | 0.0 | 33.42 | Detail |
| 168 | THCA | cuproptosis | GATC | glutamyl-tRNA amidotransferase subunit C | 5.11 | 4.63 | -0.14 | 0.0 | 0.0 | 33.33 | Detail |
| 169 | THCA | cuproptosis | PURG | purine rich element binding protein G | 1.94 | 1.4 | -0.47 | 0.0 | 0.0 | 32.6 | Detail |
| 170 | THCA | cuproptosis | APP | amyloid beta precursor protein | 14.79 | 14.55 | -0.02 | 0.0 | 0.0 | 32.38 | Detail |
| 171 | THCA | cuproptosis | LIAS | lipoic acid synthetase | 7.77 | 7.53 | -0.04 | 0.0 | 0.0 | 32.29 | Detail |
| 172 | THCA | cuproptosis | YARS2 | tyrosyl-tRNA synthetase 2 | 7.79 | 7.93 | 0.02 | 0.0 | 0.0 | 31.82 | Detail |
| 173 | THCA | cuproptosis | BUD31 | BUD31 homolog | 10.17 | 10.39 | 0.03 | 0.0 | 0.0 | 31.35 | Detail |
| 174 | THCA | cuproptosis | TNF | tumor necrosis factor | 4.04 | 2.84 | -0.51 | 0.0 | 0.0 | 31.21 | Detail |
| 175 | THCA | cuproptosis | HNRNPU | heterogeneous nuclear ribonucleoprotein U | 13.34 | 13.16 | -0.02 | 0.0 | 0.0 | 31.2 | Detail |
| 176 | THCA | cuproptosis | MAP2K2 | mitogen-activated protein kinase kinase 2 | 10.91 | 11.15 | 0.03 | 0.0 | 0.0 | 31.17 | Detail |
| 177 | THCA | cuproptosis | NDUFAB1 | NADH:ubiquinone oxidoreductase subunit AB1 | 9.57 | 9.83 | 0.04 | 0.0 | 0.0 | 31.12 | Detail |
| 178 | THCA | cuproptosis | SLC16A1 | solute carrier family 16 member 1 | 7.27 | 7.78 | 0.1 | 0.0 | 0.0 | 30.22 | Detail |
| 179 | THCA | cuproptosis | LACTB2 | lactamase beta 2 | 7.98 | 8.29 | 0.06 | 0.0 | 0.0 | 29.81 | Detail |
| 180 | THCA | cuproptosis | RUVBL2 | RuvB like AAA ATPase 2 | 10.47 | 10.71 | 0.03 | 0.0 | 0.0 | 29.75 | Detail |
| 181 | THCA | cuproptosis | RPS19 | ribosomal protein S19 | 13.24 | 13.56 | 0.03 | 0.0 | 0.0 | 29.53 | Detail |
| 182 | THCA | cuproptosis | ZNF341 | zinc finger protein 341 | 6.76 | 7.05 | 0.06 | 0.0 | 0.0 | 29.45 | Detail |
| 183 | THCA | cuproptosis | HCCS | holocytochrome c synthase | 8.62 | 8.48 | -0.02 | 0.0 | 0.0 | 28.82 | Detail |
| 184 | THCA | cuproptosis | SCAP | SREBF chaperone | 10.86 | 11.04 | 0.02 | 0.0 | 0.0 | 28.61 | Detail |
| 185 | THCA | cuproptosis | CIT | citron rho-interacting serine/threonine kinase | 6.21 | 6.9 | 0.15 | 0.0 | 0.0 | 27.92 | Detail |
| 186 | THCA | cuproptosis | ATOX1 | antioxidant 1 copper chaperone | 10.11 | 9.78 | -0.05 | 0.0 | 0.0 | 27.77 | Detail |
| 187 | THCA | cuproptosis | PITRM1 | pitrilysin metallopeptidase 1 | 10.52 | 10.35 | -0.02 | 0.0 | 0.0 | 27.63 | Detail |
| 188 | THCA | cuproptosis | C7orf26 | integrator complex subunit 15 | 9.19 | 9.03 | -0.03 | 0.0 | 0.0 | 27.61 | Detail |
| 189 | THCA | cuproptosis | SSBP1 | single stranded DNA binding protein 1 | 9.87 | 10.07 | 0.03 | 0.0 | 0.0 | 27.53 | Detail |
| 190 | THCA | cuproptosis | AOC2 | amine oxidase copper containing 2 | 3.87 | 3.31 | -0.22 | 0.0 | 0.0 | 27.47 | Detail |
| 191 | THCA | cuproptosis | MBTPS2 | membrane bound transcription factor peptidase, site 2 | 8.63 | 8.94 | 0.05 | 0.0 | 0.0 | 27.15 | Detail |
| 192 | THCA | cuproptosis | BACE1 | beta-secretase 1 | 10.09 | 10.4 | 0.04 | 0.0 | 0.0 | 27.06 | Detail |
| 193 | THCA | cuproptosis | SCO2 | synthesis of cytochrome C oxidase 2 | 8.19 | 8.56 | 0.06 | 0.0 | 0.0 | 26.99 | Detail |
| 194 | THCA | cuproptosis | MRPL15 | mitochondrial ribosomal protein L15 | 9.27 | 9.08 | -0.03 | 0.0 | 0.0 | 26.96 | Detail |
| 195 | THCA | cuproptosis | NFYC | nuclear transcription factor Y subunit gamma | 10.33 | 10.18 | -0.02 | 0.0 | 0.0 | 26.61 | Detail |
| 196 | THCA | cuproptosis | ATF4 | activating transcription factor 4 | 12.47 | 12.15 | -0.04 | 0.0 | 0.0 | 26.37 | Detail |
| 197 | THCA | cuproptosis | PPIE | peptidylprolyl isomerase E | 9.92 | 10.09 | 0.03 | 0.0 | 0.0 | 26.3 | Detail |
| 198 | THCA | cuproptosis | CLPP | caseinolytic mitochondrial matrix peptidase proteolytic subunit | 9.38 | 9.62 | 0.04 | 0.0 | 0.0 | 26.07 | Detail |
| 199 | THCA | cuproptosis | PRC1 | protein regulator of cytokinesis 1 | 6.68 | 7.03 | 0.07 | 0.0 | 0.0 | 25.91 | Detail |
| 200 | THCA | cuproptosis | RPLP0 | ribosomal protein lateral stalk subunit P0 | 14.01 | 14.23 | 0.02 | 0.0 | 0.0 | 25.75 | Detail |
| 201 | THCA | cuproptosis | IDH2 | isocitrate dehydrogenase (NADP(+)) 2 | 9.45 | 9.81 | 0.05 | 0.0 | 0.0 | 25.44 | Detail |
| 202 | THCA | cuproptosis | DNAJC11 | DnaJ heat shock protein family (Hsp40) member C11 | 9.81 | 9.7 | -0.02 | 0.0 | 0.0 | 25.41 | Detail |
| 203 | THCA | cuproptosis | CISD1 | CDGSH iron sulfur domain 1 | 8.58 | 8.33 | -0.04 | 0.0 | 0.0 | 25.23 | Detail |
| 204 | THCA | cuproptosis | MRPS12 | mitochondrial ribosomal protein S12 | 8.33 | 8.62 | 0.05 | 0.0 | 0.0 | 25.13 | Detail |
| 205 | THCA | cuproptosis | MAP2K1 | mitogen-activated protein kinase kinase 1 | 9.72 | 9.89 | 0.03 | 0.0 | 0.0 | 24.95 | Detail |
| 206 | THCA | cuproptosis | DBH | dopamine beta-hydroxylase | 4.55 | 3.82 | -0.25 | 0.0 | 0.0 | 24.86 | Detail |
| 207 | THCA | cuproptosis | MMGT1 | membrane magnesium transporter 1 | 10.04 | 9.75 | -0.04 | 0.0 | 0.0 | 24.83 | Detail |
| 208 | THCA | cuproptosis | CDC16 | cell division cycle 16 | 10.42 | 10.53 | 0.01 | 0.0 | 0.0 | 24.49 | Detail |
| 209 | THCA | cuproptosis | MRPL36 | mitochondrial ribosomal protein L36 | 7.72 | 7.97 | 0.05 | 0.0 | 0.0 | 24.39 | Detail |
| 210 | THCA | cuproptosis | SUPT16H | SPT16 homolog, facilitates chromatin remodeling subunit | 11.31 | 11.16 | -0.02 | 0.0 | 0.0 | 24.29 | Detail |
| 211 | THCA | cuproptosis | FOXM1 | forkhead box M1 | 5.53 | 6.07 | 0.13 | 0.0 | 0.0 | 24.21 | Detail |
| 212 | THCA | cuproptosis | HSPA9 | heat shock protein family A (Hsp70) member 9 | 12.39 | 12.27 | -0.01 | 0.0 | 0.0 | 24.14 | Detail |
| 213 | THCA | cuproptosis | SPATA5 | AFG2 AAA ATPase homolog A | 7.61 | 7.36 | -0.05 | 0.0 | 0.0 | 24.06 | Detail |
| 214 | THCA | cuproptosis | ANKRD9 | ankyrin repeat domain 9 | 6.04 | 5.37 | -0.17 | 0.0 | 0.0 | 23.67 | Detail |
| 215 | THCA | cuproptosis | POLR2B | RNA polymerase II subunit B | 10.83 | 10.68 | -0.02 | 0.0 | 0.0 | 23.62 | Detail |
| 216 | THCA | cuproptosis | MRPS11 | mitochondrial ribosomal protein S11 | 9.42 | 9.18 | -0.04 | 0.0 | 0.0 | 23.53 | Detail |
| 217 | THCA | cuproptosis | STK11 | serine/threonine kinase 11 | 10.1 | 10.32 | 0.03 | 0.0 | 0.0 | 23.5 | Detail |
| 218 | THCA | cuproptosis | COX18 | cytochrome c oxidase assembly factor COX18 | 8.25 | 8.46 | 0.04 | 0.0 | 0.0 | 23.09 | Detail |
| 219 | THCA | cuproptosis | SYMPK | symplekin scaffold protein | 10.95 | 11.11 | 0.02 | 0.0 | 0.0 | 22.91 | Detail |
| 220 | THCA | cuproptosis | VEGFA | vascular endothelial growth factor A | 13.09 | 12.54 | -0.06 | 0.0 | 0.0 | 22.68 | Detail |
| 221 | THCA | cuproptosis | ATPAF2 | ATP synthase mitochondrial F1 complex assembly factor 2 | 8.44 | 8.64 | 0.03 | 0.0 | 0.0 | 22.55 | Detail |
| 222 | THCA | cuproptosis | ATP8B1 | ATPase phospholipid transporting 8B1 | 10.74 | 10.48 | -0.04 | 0.0 | 0.0 | 22.44 | Detail |
| 223 | THCA | cuproptosis | NDUFA11 | NADH:ubiquinone oxidoreductase subunit A11 | 10.72 | 10.98 | 0.03 | 0.0 | 0.0 | 21.33 | Detail |
| 224 | THCA | cuproptosis | OXA1L | OXA1L mitochondrial inner membrane protein | 11.16 | 11.32 | 0.02 | 0.0 | 0.0 | 21.21 | Detail |
| 225 | THCA | cuproptosis | ABCB11 | ATP binding cassette subfamily B member 11 | 0.77 | 0.2 | -1.93 | 0.0 | 0.0 | 21.06 | Detail |
| 226 | THCA | cuproptosis | RPL3 | ribosomal protein L3 | 15.35 | 15.51 | 0.02 | 0.0 | 0.0 | 20.98 | Detail |
| 227 | THCA | cuproptosis | GLRX5 | glutaredoxin 5 | 9.85 | 9.67 | -0.03 | 0.0 | 0.0 | 20.92 | Detail |
| 228 | THCA | cuproptosis | CYP1A1 | cytochrome P450 family 1 subfamily A member 1 | 1.61 | 1.01 | -0.68 | 0.0 | 0.0 | 20.58 | Detail |
| 229 | THCA | cuproptosis | IDH3A | isocitrate dehydrogenase (NAD(+)) 3 catalytic subunit alpha | 9.76 | 9.65 | -0.02 | 0.0 | 0.0 | 20.36 | Detail |
| 230 | THCA | cuproptosis | SAMM50 | SAMM50 sorting and assembly machinery component | 9.75 | 9.58 | -0.03 | 0.0 | 0.0 | 20.16 | Detail |
| 231 | THCA | cuproptosis | ABCB8 | ATP binding cassette subfamily B member 8 | 9.74 | 9.52 | -0.03 | 0.0 | 0.0 | 19.53 | Detail |
| 232 | THCA | cuproptosis | LARS2 | leucyl-tRNA synthetase 2, mitochondrial | 8.9 | 8.75 | -0.02 | 0.0 | 0.0 | 19.39 | Detail |
| 233 | THCA | cuproptosis | HSPA8 | heat shock protein family A (Hsp70) member 8 | 14.25 | 13.86 | -0.04 | 0.0 | 0.0 | 19.19 | Detail |
| 234 | THCA | cuproptosis | PCBP2 | poly(rC) binding protein 2 | 13.64 | 13.52 | -0.01 | 0.0 | 0.0 | 19.05 | Detail |
| 235 | THCA | cuproptosis | TPK1 | thiamin pyrophosphokinase 1 | 6.61 | 6.83 | 0.05 | 0.0 | 0.0 | 18.81 | Detail |
| 236 | THCA | cuproptosis | PTPMT1 | protein tyrosine phosphatase mitochondrial 1 | 9.56 | 9.73 | 0.03 | 0.0 | 0.0 | 18.79 | Detail |
| 237 | THCA | cuproptosis | REXO2 | RNA exonuclease 2 | 8.94 | 9.15 | 0.03 | 0.0 | 0.0 | 18.67 | Detail |
| 238 | THCA | cuproptosis | GBF1 | golgi brefeldin A resistant guanine nucleotide exchange factor 1 | 11.06 | 11.21 | 0.02 | 0.0 | 0.0 | 18.26 | Detail |
| 239 | THCA | cuproptosis | POLE | DNA polymerase epsilon, catalytic subunit | 8.25 | 8.46 | 0.04 | 0.0 | 0.0 | 17.57 | Detail |
| 240 | THCA | cuproptosis | CDK5RAP1 | CDK5 regulatory subunit associated protein 1 | 8.91 | 9.05 | 0.02 | 0.0 | 0.0 | 17.54 | Detail |
| 241 | THCA | cuproptosis | MRPL12 | mitochondrial ribosomal protein L12 | 9.16 | 9.42 | 0.04 | 0.0 | 0.0 | 16.76 | Detail |
| 242 | THCA | cuproptosis | ZFAT | zinc finger and AT-hook domain containing | 7.84 | 7.94 | 0.02 | 0.0 | 0.0 | 16.33 | Detail |
| 243 | THCA | cuproptosis | COX11 | cytochrome c oxidase copper chaperone COX11 | 9.85 | 9.72 | -0.02 | 0.0 | 0.0 | 16.01 | Detail |
| 244 | THCA | cuproptosis | COX6B1 | cytochrome c oxidase subunit 6B1 | 11.26 | 11.49 | 0.03 | 0.0 | 0.0 | 15.77 | Detail |
| 245 | THCA | cuproptosis | CS | citrate synthase | 11.11 | 11.29 | 0.02 | 0.0 | 0.0 | 15.65 | Detail |
| 246 | THCA | cuproptosis | DPYD | dihydropyrimidine dehydrogenase | 9.11 | 9.3 | 0.03 | 0.0 | 0.0 | 15.65 | Detail |
| 247 | THCA | cuproptosis | MCM5 | minichromosome maintenance complex component 5 | 9.52 | 9.73 | 0.03 | 0.0 | 0.0 | 15.6 | Detail |
| 248 | THCA | cuproptosis | MBTPS1 | membrane bound transcription factor peptidase, site 1 | 12.04 | 11.92 | -0.01 | 0.0 | 0.0 | 15.56 | Detail |
| 249 | THCA | cuproptosis | PSMB5 | proteasome 20S subunit beta 5 | 10.63 | 10.76 | 0.02 | 0.0 | 0.0 | 15.35 | Detail |
| 250 | THCA | cuproptosis | NDUFS1 | NADH:ubiquinone oxidoreductase core subunit S1 | 10.11 | 9.9 | -0.03 | 0.0 | 0.0 | 14.81 | Detail |
| 251 | THCA | cuproptosis | NUBP2 | NUBP iron-sulfur cluster assembly factor 2, cytosolic | 9.59 | 9.8 | 0.03 | 0.0 | 0.0 | 14.74 | Detail |
| 252 | THCA | cuproptosis | PRNP | prion protein (Kanno blood group) | 11.89 | 12.07 | 0.02 | 0.0 | 0.0 | 14.62 | Detail |
| 253 | THCA | cuproptosis | MLKL | mixed lineage kinase domain like pseudokinase | 6.74 | 7.13 | 0.08 | 0.0 | 0.0 | 14.58 | Detail |
| 254 | THCA | cuproptosis | PDE12 | phosphodiesterase 12 | 9.07 | 8.88 | -0.03 | 0.0 | 0.0 | 13.84 | Detail |
| 255 | THCA | cuproptosis | RPL19 | ribosomal protein L19 | 14.25 | 14.39 | 0.01 | 0.0 | 0.0 | 13.82 | Detail |
| 256 | THCA | cuproptosis | MTIF3 | mitochondrial translational initiation factor 3 | 9.54 | 9.37 | -0.03 | 0.0 | 0.0 | 13.69 | Detail |
| 257 | THCA | cuproptosis | RPL10A | ribosomal protein L10a | 13.47 | 13.6 | 0.01 | 0.0 | 0.0 | 13.65 | Detail |
| 258 | THCA | cuproptosis | RMND1 | required for meiotic nuclear division 1 homolog | 9.16 | 9.33 | 0.03 | 0.0 | 0.0 | 13.61 | Detail |
| 259 | THCA | cuproptosis | IMP4 | IMP U3 small nucleolar ribonucleoprotein 4 | 8.99 | 9.09 | 0.02 | 0.0 | 0.0 | 13.44 | Detail |
| 260 | THCA | cuproptosis | CNN1 | calponin 1 | 7.27 | 6.7 | -0.12 | 0.0 | 0.0 | 13.34 | Detail |
| 261 | THCA | cuproptosis | PDE3B | phosphodiesterase 3B | 5.16 | 4.37 | -0.24 | 0.0 | 0.0 | 13.16 | Detail |
| 262 | THCA | cuproptosis | SDHA | succinate dehydrogenase complex flavoprotein subunit A | 10.85 | 10.73 | -0.02 | 0.0 | 0.0 | 13.16 | Detail |
| 263 | THCA | cuproptosis | COPZ1 | COPI coat complex subunit zeta 1 | 12.11 | 11.99 | -0.01 | 0.0 | 0.0 | 12.96 | Detail |
| 264 | THCA | cuproptosis | SLC25A26 | solute carrier family 25 member 26 | 9.14 | 9.01 | -0.02 | 0.0 | 0.0 | 12.96 | Detail |
| 265 | THCA | cuproptosis | DYNLRB1 | dynein light chain roadblock-type 1 | 11.19 | 11.38 | 0.02 | 0.0 | 0.0 | 12.82 | Detail |
| 266 | THCA | cuproptosis | YEATS2 | YEATS domain containing 2 | 9.97 | 9.79 | -0.03 | 0.0 | 0.0 | 12.7 | Detail |
| 267 | THCA | cuproptosis | USE1 | unconventional SNARE in the ER 1 | 8.5 | 8.67 | 0.03 | 0.0 | 0.0 | 12.55 | Detail |
| 268 | THCA | cuproptosis | NDUFA1 | NADH:ubiquinone oxidoreductase subunit A1 | 10.72 | 10.46 | -0.04 | 0.0 | 0.0 | 12.32 | Detail |
| 269 | THCA | cuproptosis | NDUFB10 | NADH:ubiquinone oxidoreductase subunit B10 | 10.27 | 10.48 | 0.03 | 0.0 | 0.0 | 12.11 | Detail |
| 270 | THCA | cuproptosis | CCL5 | C-C motif chemokine ligand 5 | 8.99 | 8.08 | -0.15 | 0.0 | 0.0 | 11.75 | Detail |
| 271 | THCA | cuproptosis | UBE2D4 | ubiquitin conjugating enzyme E2 D4 (putative) | 8.39 | 8.23 | -0.03 | 0.0 | 0.0 | 11.68 | Detail |
| 272 | THCA | cuproptosis | PSMA6 | proteasome 20S subunit alpha 6 | 10.51 | 10.61 | 0.01 | 0.0 | 0.0 | 11.42 | Detail |
| 273 | THCA | cuproptosis | HEPH | hephaestin | 7.52 | 7.17 | -0.07 | 0.0 | 0.0 | 11.37 | Detail |
| 274 | THCA | cuproptosis | CHCHD1 | coiled-coil-helix-coiled-coil-helix domain containing 1 | 9.27 | 9.41 | 0.02 | 0.0 | 0.01 | 10.97 | Detail |
| 275 | THCA | cuproptosis | NDUFA4 | NDUFA4 mitochondrial complex associated | 11.84 | 12.04 | 0.03 | 0.0 | 0.01 | 10.9 | Detail |
| 276 | THCA | cuproptosis | UBE2D2 | ubiquitin conjugating enzyme E2 D2 | 10.82 | 10.91 | 0.01 | 0.0 | 0.01 | 10.62 | Detail |
| 277 | THCA | cuproptosis | GCLM | glutamate-cysteine ligase modifier subunit | 9.16 | 8.91 | -0.04 | 0.0 | 0.01 | 10.6 | Detail |
| 278 | THCA | cuproptosis | GGPS1 | geranylgeranyl diphosphate synthase 1 | 9.81 | 9.72 | -0.01 | 0.0 | 0.01 | 10.59 | Detail |
| 279 | THCA | cuproptosis | CDK1 | cyclin dependent kinase 1 | 6.54 | 6.73 | 0.04 | 0.0 | 0.01 | 10.52 | Detail |
| 280 | THCA | cuproptosis | SLC25A19 | solute carrier family 25 member 19 | 7.18 | 7.39 | 0.04 | 0.0 | 0.01 | 10.32 | Detail |
| 281 | THCA | cuproptosis | SERPINE1 | serpin family E member 1 | 8.62 | 9.31 | 0.11 | 0.0 | 0.01 | 9.87 | Detail |
| 282 | THCA | cuproptosis | SLC25A3 | solute carrier family 25 member 3 | 12.62 | 12.73 | 0.01 | 0.0 | 0.01 | 9.76 | Detail |
| 283 | THCA | cuproptosis | EGLN1 | egl-9 family hypoxia inducible factor 1 | 10.21 | 10.09 | -0.02 | 0.0 | 0.01 | 9.53 | Detail |
| 284 | THCA | cuproptosis | MRPL41 | mitochondrial ribosomal protein L41 | 8.88 | 9.21 | 0.05 | 0.0 | 0.01 | 9.33 | Detail |
| 285 | THCA | cuproptosis | DLST | dihydrolipoamide S-succinyltransferase | 11.1 | 11.21 | 0.01 | 0.0 | 0.01 | 9.25 | Detail |
| 286 | THCA | cuproptosis | SOX2 | SRY-box transcription factor 2 | 1.13 | 0.87 | -0.38 | 0.0 | 0.01 | 8.99 | Detail |
| 287 | THCA | cuproptosis | SLC31A1 | solute carrier family 31 member 1 | 9.48 | 9.31 | -0.02 | 0.0 | 0.01 | 8.91 | Detail |
| 288 | THCA | cuproptosis | ALAS1 | 5'-aminolevulinate synthase 1 | 10.16 | 10.01 | -0.02 | 0.0 | 0.01 | 8.83 | Detail |
| 289 | THCA | cuproptosis | PDHA1 | pyruvate dehydrogenase E1 subunit alpha 1 | 10.79 | 10.72 | -0.01 | 0.0 | 0.01 | 8.8 | Detail |
| 290 | THCA | cuproptosis | MRPL55 | mitochondrial ribosomal protein L55 | 9.09 | 9.31 | 0.03 | 0.01 | 0.01 | 8.55 | Detail |
| 291 | THCA | cuproptosis | CCDC28B | coiled-coil domain containing 28B | 6.66 | 6.96 | 0.06 | 0.01 | 0.01 | 8.38 | Detail |
| 292 | THCA | cuproptosis | POLR2K | RNA polymerase II, I and III subunit K | 9.77 | 9.65 | -0.02 | 0.01 | 0.01 | 8.04 | Detail |
| 293 | THCA | cuproptosis | UBAP2L | ubiquitin associated protein 2 like | 11.31 | 11.42 | 0.01 | 0.01 | 0.01 | 7.68 | Detail |
| 294 | THCA | cuproptosis | BARX1 | BARX homeobox 1 | 2.62 | 2.23 | -0.23 | 0.01 | 0.01 | 7.64 | Detail |
| 295 | THCA | cuproptosis | MRPS33 | mitochondrial ribosomal protein S33 | 9.55 | 9.43 | -0.02 | 0.01 | 0.01 | 7.56 | Detail |
| 296 | THCA | cuproptosis | CENPW | centromere protein W | 4.61 | 4.85 | 0.07 | 0.01 | 0.01 | 7.38 | Detail |
| 297 | THCA | cuproptosis | NDOR1 | NADPH dependent diflavin oxidoreductase 1 | 8.95 | 9.11 | 0.03 | 0.01 | 0.01 | 7.24 | Detail |
| 298 | THCA | cuproptosis | IDH3G | isocitrate dehydrogenase (NAD(+)) 3 non-catalytic subunit gamma | 10.07 | 10.2 | 0.02 | 0.01 | 0.01 | 7.22 | Detail |
| 299 | THCA | cuproptosis | YAP1 | Yes1 associated transcriptional regulator | 11.73 | 11.94 | 0.03 | 0.01 | 0.01 | 7.21 | Detail |
| 300 | THCA | cuproptosis | ALB | albumin | 2.85 | 2.56 | -0.16 | 0.01 | 0.01 | 7.13 | Detail |
| 301 | THCA | cuproptosis | ZBTB11 | zinc finger and BTB domain containing 11 | 9.24 | 9.11 | -0.02 | 0.01 | 0.01 | 7.08 | Detail |
| 302 | THCA | cuproptosis | MRPS16 | mitochondrial ribosomal protein S16 | 10.35 | 10.45 | 0.01 | 0.01 | 0.01 | 7.03 | Detail |
| 303 | THCA | cuproptosis | SLC25A39 | solute carrier family 25 member 39 | 11.48 | 11.69 | 0.03 | 0.01 | 0.01 | 7.01 | Detail |
| 304 | THCA | cuproptosis | HAUS5 | HAUS augmin like complex subunit 5 | 8.41 | 8.53 | 0.02 | 0.01 | 0.01 | 6.76 | Detail |
| 305 | THCA | cuproptosis | OSBPL9 | oxysterol binding protein like 9 | 11.05 | 10.97 | -0.01 | 0.01 | 0.01 | 6.65 | Detail |
| 306 | THCA | cuproptosis | SLC16A6 | solute carrier family 16 member 6 | 5.26 | 4.89 | -0.11 | 0.01 | 0.01 | 6.24 | Detail |
| 307 | THCA | cuproptosis | MPI | mannose phosphate isomerase | 9.74 | 9.65 | -0.01 | 0.01 | 0.02 | 5.95 | Detail |
| 308 | THCA | cuproptosis | MAN2C1 | mannosidase alpha class 2C member 1 | 10.69 | 10.83 | 0.02 | 0.01 | 0.02 | 5.95 | Detail |
| 309 | THCA | cuproptosis | CLEC11A | C-type lectin domain containing 11A | 7.08 | 7.45 | 0.07 | 0.01 | 0.02 | 5.82 | Detail |
| 310 | THCA | cuproptosis | FASLG | Fas ligand | 3.66 | 2.98 | -0.3 | 0.01 | 0.02 | 5.78 | Detail |
| 311 | THCA | cuproptosis | LDLR | low density lipoprotein receptor | 8.14 | 8.62 | 0.08 | 0.01 | 0.02 | 5.78 | Detail |
| 312 | THCA | cuproptosis | PCMT1 | protein-L-isoaspartate (D-aspartate) O-methyltransferase | 10.48 | 10.54 | 0.01 | 0.01 | 0.02 | 5.43 | Detail |
| 313 | THCA | cuproptosis | PPA2 | inorganic pyrophosphatase 2 | 10.03 | 10.17 | 0.02 | 0.01 | 0.02 | 5.38 | Detail |
| 314 | THCA | cuproptosis | DLAT | dihydrolipoamide S-acetyltransferase | 10.04 | 9.91 | -0.02 | 0.01 | 0.02 | 5.26 | Detail |
| 315 | THCA | cuproptosis | MOXD1 | monooxygenase DBH like 1 | 6.7 | 6.1 | -0.14 | 0.01 | 0.02 | 5.23 | Detail |
| 316 | THCA | cuproptosis | SRP54 | signal recognition particle 54 | 10.16 | 10.05 | -0.01 | 0.01 | 0.02 | 5.16 | Detail |
| 317 | THCA | cuproptosis | FAU | FAU ubiquitin like and ribosomal protein S30 fusion | 12.94 | 13.08 | 0.02 | 0.01 | 0.02 | 5.01 | Detail |
| 318 | THCA | cuproptosis | CHMP4B | charged multivesicular body protein 4B | 11.77 | 11.86 | 0.01 | 0.01 | 0.02 | 4.94 | Detail |
| 319 | THCA | cuproptosis | DIP2B | disco interacting protein 2 homolog B | 9.64 | 9.52 | -0.02 | 0.01 | 0.02 | 4.92 | Detail |
| 320 | THCA | cuproptosis | NDUFB6 | NADH:ubiquinone oxidoreductase subunit B6 | 9.62 | 9.44 | -0.03 | 0.01 | 0.02 | 4.88 | Detail |
| 321 | THCA | cuproptosis | NUP133 | nucleoporin 133 | 10.09 | 9.98 | -0.02 | 0.01 | 0.02 | 4.82 | Detail |
| 322 | THCA | cuproptosis | COX19 | cytochrome c oxidase assembly factor COX19 | 8.39 | 8.24 | -0.03 | 0.01 | 0.02 | 4.8 | Detail |
| 323 | THCA | cuproptosis | PFKP | phosphofructokinase, platelet | 11.27 | 11.44 | 0.02 | 0.01 | 0.02 | 4.64 | Detail |
| 324 | THCA | cuproptosis | MRPL30 | mitochondrial ribosomal protein L30 | 9.34 | 9.25 | -0.01 | 0.01 | 0.02 | 4.37 | Detail |
| 325 | THCA | cuproptosis | MRPL39 | mitochondrial ribosomal protein L39 | 8.28 | 8.18 | -0.02 | 0.01 | 0.02 | 4.26 | Detail |
| 326 | THCA | cuproptosis | MRRF | mitochondrial ribosome recycling factor | 8.69 | 8.6 | -0.02 | 0.01 | 0.02 | 4.0 | Detail |
| 327 | THCA | cuproptosis | ABCE1 | ATP binding cassette subfamily E member 1 | 10.0 | 9.88 | -0.02 | 0.02 | 0.02 | 3.68 | Detail |
| 328 | THCA | cuproptosis | NDUFB9 | NADH:ubiquinone oxidoreductase subunit B9 | 11.28 | 11.44 | 0.02 | 0.02 | 0.03 | 3.35 | Detail |
| 329 | THCA | cuproptosis | FGD6 | FYVE, RhoGEF and PH domain containing 6 | 9.32 | 9.12 | -0.03 | 0.02 | 0.03 | 3.31 | Detail |
| 330 | THCA | cuproptosis | PDK1 | pyruvate dehydrogenase kinase 1 | 7.79 | 7.57 | -0.04 | 0.02 | 0.03 | 3.3 | Detail |
| 331 | THCA | cuproptosis | ULK2 | unc-51 like autophagy activating kinase 2 | 9.06 | 8.96 | -0.02 | 0.02 | 0.03 | 3.04 | Detail |
| 332 | THCA | cuproptosis | CCS | copper chaperone for superoxide dismutase | 9.55 | 9.41 | -0.02 | 0.02 | 0.03 | 3.01 | Detail |
| 333 | THCA | cuproptosis | NDUFV1 | NADH:ubiquinone oxidoreductase core subunit V1 | 11.31 | 11.23 | -0.01 | 0.02 | 0.03 | 2.7 | Detail |
| 334 | THCA | cuproptosis | MRPS18A | mitochondrial ribosomal protein S18A | 9.33 | 9.42 | 0.01 | 0.02 | 0.03 | 2.68 | Detail |
| 335 | THCA | cuproptosis | MAL2 | mal, T cell differentiation protein 2 | 12.53 | 12.35 | -0.02 | 0.02 | 0.03 | 2.62 | Detail |
| 336 | THCA | cuproptosis | GTPBP4 | GTP binding protein 4 | 9.54 | 9.45 | -0.01 | 0.02 | 0.03 | 2.45 | Detail |
| 337 | THCA | cuproptosis | NOP2 | NOP2 nucleolar protein | 8.98 | 9.13 | 0.02 | 0.02 | 0.03 | 2.37 | Detail |
| 338 | THCA | cuproptosis | CDKN3 | cyclin dependent kinase inhibitor 3 | 3.29 | 3.57 | 0.12 | 0.02 | 0.03 | 2.27 | Detail |
| 339 | THCA | cuproptosis | SLC7A5 | solute carrier family 7 member 5 | 7.32 | 7.81 | 0.09 | 0.02 | 0.03 | 2.09 | Detail |
| 340 | THCA | cuproptosis | MRPL52 | mitochondrial ribosomal protein L52 | 9.4 | 9.52 | 0.02 | 0.02 | 0.03 | 1.57 | Detail |
| 341 | THCA | cuproptosis | RAD9A | RAD9 checkpoint clamp component A | 8.11 | 7.96 | -0.03 | 0.02 | 0.04 | 1.51 | Detail |
| 342 | THCA | cuproptosis | TUBGCP4 | tubulin gamma complex component 4 | 7.78 | 7.67 | -0.02 | 0.02 | 0.04 | 1.49 | Detail |
| 343 | THCA | cuproptosis | TUT1 | terminal uridylyl transferase 1, U6 snRNA-specific | 9.53 | 9.45 | -0.01 | 0.02 | 0.04 | 1.45 | Detail |
| 344 | THCA | cuproptosis | PRKDC | protein kinase, DNA-activated, catalytic subunit | 11.36 | 11.18 | -0.02 | 0.02 | 0.04 | 1.37 | Detail |
| 345 | THCA | cuproptosis | ACD | ACD shelterin complex subunit and telomerase recruitment factor | 8.61 | 8.45 | -0.03 | 0.03 | 0.04 | 1.18 | Detail |
| 346 | THCA | cuproptosis | RPL27 | ribosomal protein L27 | 12.67 | 12.82 | 0.02 | 0.03 | 0.04 | 1.18 | Detail |
| 347 | THCA | cuproptosis | MARS2 | methionyl-tRNA synthetase 2, mitochondrial | 7.04 | 6.85 | -0.04 | 0.03 | 0.04 | 1.07 | Detail |
| 348 | THCA | cuproptosis | PDHB | pyruvate dehydrogenase E1 subunit beta | 10.09 | 10.18 | 0.01 | 0.03 | 0.04 | 0.16 | Detail |
| 349 | THCA | cuproptosis | MRPS21 | mitochondrial ribosomal protein S21 | 10.28 | 10.4 | 0.02 | 0.03 | 0.04 | 0.16 | Detail |
| 350 | THCA | cuproptosis | SLC11A2 | solute carrier family 11 member 2 | 11.12 | 11.06 | -0.01 | 0.03 | 0.05 | 0.08 | Detail |
| 351 | THCA | cuproptosis | RTEL1 | regulator of telomere elongation helicase 1 | 8.66 | 8.54 | -0.02 | 0.03 | 0.05 | 0.05 | Detail |
| 352 | THCA | cuproptosis | ARF1 | ADP ribosylation factor 1 | 13.3 | 13.36 | 0.01 | 0.08 | 0.11 | 0.0 | Detail |
| 353 | THCA | cuproptosis | ATP7A | ATPase copper transporting alpha | 8.73 | 8.64 | -0.02 | 0.08 | 0.1 | 0.0 | Detail |
| 354 | THCA | cuproptosis | MT2A | metallothionein 2A | 12.14 | 11.94 | -0.02 | 0.3 | 0.34 | 0.0 | Detail |
| 355 | THCA | cuproptosis | SLC31A2 | solute carrier family 31 member 2 | 7.77 | 7.61 | -0.03 | 0.15 | 0.18 | 0.0 | Detail |
| 356 | THCA | cuproptosis | AOC1 | amine oxidase copper containing 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 357 | THCA | cuproptosis | AQP2 | aquaporin 2 | 1.12 | 1.08 | -0.06 | 0.36 | 0.4 | 0.0 | Detail |
| 358 | THCA | cuproptosis | DAXX | death domain associated protein | 10.32 | 10.24 | -0.01 | 0.05 | 0.07 | 0.0 | Detail |
| 359 | THCA | cuproptosis | MT1B | metallothionein 1B | 0.04 | 0.02 | -0.79 | 0.18 | 0.22 | 0.0 | Detail |
| 360 | THCA | cuproptosis | MT1HL1 | metallothionein 1H like 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 361 | THCA | cuproptosis | MT3 | metallothionein 3 | 2.41 | 2.4 | -0.01 | 0.24 | 0.28 | 0.0 | Detail |
| 362 | THCA | cuproptosis | MT4 | metallothionein 4 | 0 | 0 | NA | NA | NA | 0 | Detail |
| 363 | THCA | cuproptosis | COMMD1 | copper metabolism domain containing 1 | 8.43 | 8.39 | -0.01 | 0.92 | 0.93 | 0.0 | Detail |
| 364 | THCA | cuproptosis | XIAP | X-linked inhibitor of apoptosis | 10.82 | 10.83 | 0.0 | 0.91 | 0.92 | 0.0 | Detail |
| 365 | THCA | cuproptosis | COX17 | cytochrome c oxidase copper chaperone COX17 | 9.19 | 9.22 | 0.0 | 0.45 | 0.5 | 0.0 | Detail |
| 366 | THCA | cuproptosis | PARK7 | Parkinsonism associated deglycase | 12.3 | 12.22 | -0.01 | 0.21 | 0.24 | 0.0 | Detail |
| 367 | THCA | cuproptosis | RIPK3 | receptor interacting serine/threonine kinase 3 | 8.2 | 8.16 | -0.01 | 0.47 | 0.51 | 0.0 | Detail |
| 368 | THCA | cuproptosis | GLS | glutaminase | 11.08 | 11.2 | 0.02 | 0.07 | 0.1 | 0.0 | Detail |
| 369 | THCA | cuproptosis | AASDHPPT | aminoadipate-semialdehyde dehydrogenase-phosphopantetheinyl transferase | 9.66 | 9.63 | -0.01 | 0.26 | 0.31 | 0.0 | Detail |
| 370 | THCA | cuproptosis | ABCB4 | ATP binding cassette subfamily B member 4 | 3.94 | 3.59 | -0.13 | 0.15 | 0.18 | 0.0 | Detail |
| 371 | THCA | cuproptosis | ABCB7 | ATP binding cassette subfamily B member 7 | 8.65 | 8.69 | 0.01 | 0.34 | 0.38 | 0.0 | Detail |
| 372 | THCA | cuproptosis | ACLY | ATP citrate lyase | 11.0 | 11.02 | 0.0 | 0.44 | 0.48 | 0.0 | Detail |
| 373 | THCA | cuproptosis | ACO2 | aconitase 2 | 11.68 | 11.6 | -0.01 | 0.04 | 0.05 | 0.0 | Detail |
| 374 | THCA | cuproptosis | ACP1 | acid phosphatase 1 | 10.65 | 10.63 | -0.0 | 0.83 | 0.85 | 0.0 | Detail |
| 375 | THCA | cuproptosis | ACTR10 | actin related protein 10 | 10.26 | 10.32 | 0.01 | 0.05 | 0.07 | 0.0 | Detail |
| 376 | THCA | cuproptosis | ADGRE5 | adhesion G protein-coupled receptor E5 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 377 | THCA | cuproptosis | AOC3 | amine oxidase copper containing 3 | 8.97 | 8.7 | -0.04 | 0.15 | 0.18 | 0.0 | Detail |
| 378 | THCA | cuproptosis | API5 | apoptosis inhibitor 5 | 11.17 | 11.12 | -0.01 | 0.29 | 0.33 | 0.0 | Detail |
| 379 | THCA | cuproptosis | ARFGEF1-DT | ARFGEF1 divergent transcript | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 380 | THCA | cuproptosis | ARHGEF39 | Rho guanine nucleotide exchange factor 39 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 381 | THCA | cuproptosis | ARMH3 | armadillo like helical domain containing 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 382 | THCA | cuproptosis | ATL2 | atlastin GTPase 2 | 10.99 | 11.07 | 0.01 | 0.22 | 0.25 | 0.0 | Detail |
| 383 | THCA | cuproptosis | ATP5PD | ATP synthase peripheral stalk subunit d | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 384 | THCA | cuproptosis | ATP7BP1 | ATPase copper transporting beta pseudogene 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 385 | THCA | cuproptosis | BCKDH | protein phosphatase, Mg2+/Mn2+ dependent 1K | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 386 | THCA | cuproptosis | BRCA1 | BRCA1 DNA repair associated | 7.6 | 7.35 | -0.05 | 0.18 | 0.22 | 0.0 | Detail |
| 387 | THCA | cuproptosis | BRPF1 | bromodomain and PHD finger containing 1 | 9.35 | 9.3 | -0.01 | 0.04 | 0.06 | 0.0 | Detail |
| 388 | THCA | cuproptosis | C1QBP | complement C1q binding protein | 10.06 | 10.09 | 0.01 | 0.62 | 0.66 | 0.0 | Detail |
| 389 | THCA | cuproptosis | C6orf136 | chromosome 6 open reading frame 136 | 8.31 | 8.31 | -0.0 | 0.59 | 0.63 | 0.0 | Detail |
| 390 | THCA | cuproptosis | C6ORF136 | chromosome 6 open reading frame 136 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 391 | THCA | cuproptosis | CAPRIN1 | cell cycle associated protein 1 | 12.03 | 12.1 | 0.01 | 0.05 | 0.07 | 0.0 | Detail |
| 392 | THCA | cuproptosis | CAT | catalase | 12.29 | 12.18 | -0.01 | 0.23 | 0.28 | 0.0 | Detail |
| 393 | THCA | cuproptosis | CCDC115 | coiled-coil domain containing 115 | 10.35 | 10.35 | -0.0 | 1.0 | 1.0 | 0.0 | Detail |
| 394 | THCA | cuproptosis | CCDC137 | coiled-coil domain containing 137 | 8.43 | 8.53 | 0.02 | 0.05 | 0.07 | 0.0 | Detail |
| 395 | THCA | cuproptosis | CCL8 | C-C motif chemokine ligand 8 | 4.47 | 4.05 | -0.14 | 0.09 | 0.12 | 0.0 | Detail |
| 396 | THCA | cuproptosis | CD274 | CD274 molecule | 5.6 | 5.39 | -0.05 | 0.62 | 0.66 | 0.0 | Detail |
| 397 | THCA | cuproptosis | CEPT1 | choline/ethanolamine phosphotransferase 1 | 9.29 | 9.26 | -0.01 | 0.63 | 0.67 | 0.0 | Detail |
| 398 | THCA | cuproptosis | CERS2 | ceramide synthase 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 399 | THCA | cuproptosis | CERT1 | ceramide transporter 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 400 | THCA | cuproptosis | CHTOP | chromatin target of PRMT1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 401 | THCA | cuproptosis | CIAO2A | cytosolic iron-sulfur assembly component 2A | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 402 | THCA | cuproptosis | CIAO2B | cytosolic iron-sulfur assembly component 2B | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 403 | THCA | cuproptosis | CIAPIN1 | cytokine induced apoptosis inhibitor 1 | 9.3 | 9.23 | -0.01 | 0.09 | 0.11 | 0.0 | Detail |
| 404 | THCA | cuproptosis | CLPX | caseinolytic mitochondrial matrix peptidase chaperone subunit X | 9.34 | 9.38 | 0.01 | 0.09 | 0.12 | 0.0 | Detail |
| 405 | THCA | cuproptosis | CLUH | clustered mitochondria homolog | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 406 | THCA | cuproptosis | CMC1 | C-X9-C motif containing 1 | 7.0 | 6.87 | -0.03 | 0.14 | 0.17 | 0.0 | Detail |
| 407 | THCA | cuproptosis | COA5 | cytochrome c oxidase assembly factor 5 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 408 | THCA | cuproptosis | COA6 | cytochrome c oxidase assembly factor 6 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 409 | THCA | cuproptosis | COAS | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 410 | THCA | cuproptosis | COPB2 | COPI coat complex subunit beta 2 | 11.66 | 11.64 | -0.0 | 0.7 | 0.73 | 0.0 | Detail |
| 411 | THCA | cuproptosis | copper鈥搝inc alloy (CZA) | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 412 | THCA | cuproptosis | COPS5 | COP9 signalosome subunit 5 | 9.88 | 9.82 | -0.01 | 0.1 | 0.13 | 0.0 | Detail |
| 413 | THCA | cuproptosis | COX14 | cytochrome c oxidase assembly factor COX14 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 414 | THCA | cuproptosis | COX23 | coiled-coil-helix-coiled-coil-helix domain containing 7 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 415 | THCA | cuproptosis | COX4I1 | cytochrome c oxidase subunit 4I1 | 12.54 | 12.65 | 0.01 | 0.07 | 0.09 | 0.0 | Detail |
| 416 | THCA | cuproptosis | COX7A1 | cytochrome c oxidase subunit 7A1 | 8.18 | 7.78 | -0.07 | 0.12 | 0.15 | 0.0 | Detail |
| 417 | THCA | cuproptosis | COX7B | cytochrome c oxidase subunit 7B | 10.49 | 10.36 | -0.02 | 0.08 | 0.11 | 0.0 | Detail |
| 418 | THCA | cuproptosis | COX7C | cytochrome c oxidase subunit 7C | 12.05 | 12.15 | 0.01 | 0.08 | 0.11 | 0.0 | Detail |
| 419 | THCA | cuproptosis | CSTF3 | cleavage stimulation factor subunit 3 | 9.28 | 9.29 | 0.0 | 0.96 | 0.96 | 0.0 | Detail |
| 420 | THCA | cuproptosis | CTR1 | solute carrier family 31 member 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 421 | THCA | cuproptosis | CTR2 | solute carrier family 11 member 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 422 | THCA | cuproptosis | CYC1 | cytochrome c1 | 10.69 | 10.66 | -0.0 | 0.31 | 0.35 | 0.0 | Detail |
| 423 | THCA | cuproptosis | CYP2D6 | cytochrome P450 family 2 subfamily D member 6 | 3.74 | 3.96 | 0.08 | 0.27 | 0.31 | 0.0 | Detail |
| 424 | THCA | cuproptosis | DARS1 | aspartyl-tRNA synthetase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 425 | THCA | cuproptosis | DCYTB | cytochrome b reductase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 426 | THCA | cuproptosis | DLG3 | discs large MAGUK scaffold protein 3 | 10.54 | 10.66 | 0.02 | 0.13 | 0.16 | 0.0 | Detail |
| 427 | THCA | cuproptosis | DMT1 | solute carrier family 11 member 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 428 | THCA | cuproptosis | DNA2 | DNA replication helicase/nuclease 2 | 4.76 | 4.76 | -0.0 | 0.66 | 0.69 | 0.0 | Detail |
| 429 | THCA | cuproptosis | DPH1 | diphthamide biosynthesis 1 | 9.4 | 9.43 | 0.0 | 0.46 | 0.5 | 0.0 | Detail |
| 430 | THCA | cuproptosis | EARS2 | glutamyl-tRNA synthetase 2, mitochondrial | 9.21 | 9.26 | 0.01 | 0.63 | 0.66 | 0.0 | Detail |
| 431 | THCA | cuproptosis | EFCC1 | EF-hand and coiled-coil domain containing 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 432 | THCA | cuproptosis | EGFR | epidermal growth factor receptor | 9.46 | 9.54 | 0.01 | 0.84 | 0.85 | 0.0 | Detail |
| 433 | THCA | cuproptosis | EIF4G1 | eukaryotic translation initiation factor 4 gamma 1 | 12.68 | 12.74 | 0.01 | 0.19 | 0.22 | 0.0 | Detail |
| 434 | THCA | cuproptosis | ELOB | elongin B | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 435 | THCA | cuproptosis | EMC6 | ER membrane protein complex subunit 6 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 436 | THCA | cuproptosis | ENTPD3 | ectonucleoside triphosphate diphosphohydrolase 3 | 7.07 | 7.11 | 0.01 | 0.14 | 0.17 | 0.0 | Detail |
| 437 | THCA | cuproptosis | ERK1 | mitogen-activated protein kinase 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 438 | THCA | cuproptosis | EXOC3 | exocyst complex component 3 | 10.33 | 10.27 | -0.01 | 0.07 | 0.09 | 0.0 | Detail |
| 439 | THCA | cuproptosis | FAK | protein tyrosine kinase 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 440 | THCA | cuproptosis | FDX2 | ferredoxin 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 441 | THCA | cuproptosis | FOXD2-AS1 | FOXD2 adjacent opposite strand RNA 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 442 | THCA | cuproptosis | GATB | glutamyl-tRNA amidotransferase subunit B | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 443 | THCA | cuproptosis | GLS40 | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 444 | THCA | cuproptosis | glycerophospholipid | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 445 | THCA | cuproptosis | GNL3 | G protein nucleolar 3 | 9.79 | 9.81 | 0.0 | 0.48 | 0.52 | 0.0 | Detail |
| 446 | THCA | cuproptosis | GRX1 | glutaredoxin | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 447 | THCA | cuproptosis | GSH | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 448 | THCA | cuproptosis | GTF2B | general transcription factor IIB | 9.45 | 9.33 | -0.02 | 0.16 | 0.2 | 0.0 | Detail |
| 449 | THCA | cuproptosis | GTPBP6 | GTP binding protein 6 (putative) | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 450 | THCA | cuproptosis | H1-10-AS1 | H1-10 antisense RNA 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 451 | THCA | cuproptosis | H2AC6 | H2A clustered histone 6 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 452 | THCA | cuproptosis | H2BC6-AS1 | H2BC6 antisense RNA 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 453 | THCA | cuproptosis | H3C1 | H3 clustered histone 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 454 | THCA | cuproptosis | HIST1H3A | H3 clustered histone 1 | 1.07 | 0.83 | -0.38 | 0.04 | 0.05 | 0.0 | Detail |
| 455 | THCA | cuproptosis | HMBS | hydroxymethylbilane synthase | 8.31 | 8.5 | 0.03 | 0.05 | 0.07 | 0.0 | Detail |
| 456 | THCA | cuproptosis | HSCB | HscB mitochondrial iron-sulfur cluster cochaperone | 7.94 | 8.0 | 0.01 | 0.09 | 0.11 | 0.0 | Detail |
| 457 | THCA | cuproptosis | HSP70 | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 458 | THCA | cuproptosis | HSPD1 | heat shock protein family D (Hsp60) member 1 | 11.99 | 11.96 | -0.0 | 0.59 | 0.63 | 0.0 | Detail |
| 459 | THCA | cuproptosis | HSPH1 | heat shock protein family H (Hsp110) member 1 | 11.06 | 11.02 | -0.01 | 0.3 | 0.35 | 0.0 | Detail |
| 460 | THCA | cuproptosis | IBA57 | iron-sulfur cluster assembly factor IBA57 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 461 | THCA | cuproptosis | IFT81 | intraflagellar transport 81 | 8.84 | 8.76 | -0.01 | 0.07 | 0.09 | 0.0 | Detail |
| 462 | THCA | cuproptosis | IL-8 | C-X-C motif chemokine ligand 8 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 463 | THCA | cuproptosis | INKA1 | inka box actin regulator 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 464 | THCA | cuproptosis | INSL4 | insulin like 4 | 0.02 | 0.01 | -0.41 | 0.39 | 0.44 | 0.0 | Detail |
| 465 | THCA | cuproptosis | INTS6 | integrator complex subunit 6 | 9.73 | 9.58 | -0.02 | 0.19 | 0.22 | 0.0 | Detail |
| 466 | THCA | cuproptosis | IOP1 | cytosolic iron-sulfur assembly component 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 467 | THCA | cuproptosis | ISCU2 | iron-sulfur cluster assembly enzyme | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 468 | THCA | cuproptosis | ISD11 | LYR motif containing 4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 469 | THCA | cuproptosis | KAT8 | lysine acetyltransferase 8 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 470 | THCA | cuproptosis | KIDINS220 | kinase D interacting substrate 220 | 11.04 | 11.07 | 0.0 | 0.94 | 0.94 | 0.0 | Detail |
| 471 | THCA | cuproptosis | KLF5 | KLF transcription factor 5 | 10.8 | 10.68 | -0.02 | 0.66 | 0.69 | 0.0 | Detail |
| 472 | THCA | cuproptosis | KMT2A | lysine methyltransferase 2A | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 473 | THCA | cuproptosis | LASTR | lncRNA associated with SART3 regulation of splicing | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 474 | THCA | cuproptosis | LINC02038 | long intergenic non-protein coding RNA 2038 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 475 | THCA | cuproptosis | LINC02154 | long intergenic non-protein coding RNA 2154 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 476 | THCA | cuproptosis | LINC03085 | long intergenic non-protein coding RNA 3085 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 477 | THCA | cuproptosis | LOX | lysyl oxidase | 7.66 | 7.87 | 0.04 | 0.34 | 0.38 | 0.0 | Detail |
| 478 | THCA | cuproptosis | MARCHF5 | membrane associated ring-CH-type finger 5 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 479 | THCA | cuproptosis | MAU2 | MAU2 sister chromatid cohesion factor | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 480 | THCA | cuproptosis | MCAT | malonyl-CoA-acyl carrier protein transacylase | 8.42 | 8.44 | 0.0 | 0.52 | 0.56 | 0.0 | Detail |
| 481 | THCA | cuproptosis | MCMBP | minichromosome maintenance complex binding protein | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 482 | THCA | cuproptosis | MCUR1 | mitochondrial calcium uniporter regulator 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 483 | THCA | cuproptosis | MDH2 | malate dehydrogenase 2 | 11.34 | 11.37 | 0.0 | 0.82 | 0.84 | 0.0 | Detail |
| 484 | THCA | cuproptosis | MEK1 | mitogen-activated protein kinase kinase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 485 | THCA | cuproptosis | MEK2 | mitogen-activated protein kinase kinase 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 486 | THCA | cuproptosis | METTL17 | methyltransferase like 17 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 487 | THCA | cuproptosis | MFF | mitochondrial fission factor | 10.1 | 10.1 | -0.0 | 0.95 | 0.96 | 0.0 | Detail |
| 488 | THCA | cuproptosis | MFN2 | mitofusin 2 | 12.04 | 11.96 | -0.01 | 0.04 | 0.06 | 0.0 | Detail |
| 489 | THCA | cuproptosis | MPC1 | mitochondrial pyruvate carrier 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 490 | THCA | cuproptosis | MPZL3 | myelin protein zero like 3 | 5.77 | 5.83 | 0.01 | 0.36 | 0.4 | 0.0 | Detail |
| 491 | THCA | cuproptosis | MRM2 | mitochondrial rRNA methyltransferase 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 492 | THCA | cuproptosis | MRPL13 | mitochondrial ribosomal protein L13 | 8.77 | 8.76 | -0.0 | 0.87 | 0.89 | 0.0 | Detail |
| 493 | THCA | cuproptosis | MRPL16 | mitochondrial ribosomal protein L16 | 8.66 | 8.53 | -0.02 | 0.12 | 0.15 | 0.0 | Detail |
| 494 | THCA | cuproptosis | MRPL21 | mitochondrial ribosomal protein L21 | 9.16 | 9.3 | 0.02 | 0.06 | 0.08 | 0.0 | Detail |
| 495 | THCA | cuproptosis | MRPL2 | mitochondrial ribosomal protein L2 | 9.14 | 9.18 | 0.01 | 0.36 | 0.4 | 0.0 | Detail |
| 496 | THCA | cuproptosis | MRPL34 | mitochondrial ribosomal protein L34 | 9.59 | 9.65 | 0.01 | 0.6 | 0.64 | 0.0 | Detail |
| 497 | THCA | cuproptosis | MRPL40 | mitochondrial ribosomal protein L40 | 10.34 | 10.44 | 0.01 | 0.26 | 0.31 | 0.0 | Detail |
| 498 | THCA | cuproptosis | MRPL44 | mitochondrial ribosomal protein L44 | 9.03 | 8.94 | -0.01 | 0.04 | 0.06 | 0.0 | Detail |
| 499 | THCA | cuproptosis | MRPL53 | mitochondrial ribosomal protein L53 | 9.55 | 9.61 | 0.01 | 0.15 | 0.18 | 0.0 | Detail |
| 500 | THCA | cuproptosis | MRPL57 | mitochondrial ribosomal protein L57 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 501 | THCA | cuproptosis | MRPS18B | mitochondrial ribosomal protein S18B | 10.53 | 10.55 | 0.0 | 0.74 | 0.77 | 0.0 | Detail |
| 502 | THCA | cuproptosis | MRPS18C | mitochondrial ribosomal protein S18C | 8.41 | 8.28 | -0.02 | 0.04 | 0.05 | 0.0 | Detail |
| 503 | THCA | cuproptosis | MRPS2 | mitochondrial ribosomal protein S2 | 9.47 | 9.39 | -0.01 | 0.12 | 0.15 | 0.0 | Detail |
| 504 | THCA | cuproptosis | MRPS5 | mitochondrial ribosomal protein S5 | 9.81 | 9.86 | 0.01 | 0.12 | 0.15 | 0.0 | Detail |
| 505 | THCA | cuproptosis | MRPS7 | mitochondrial ribosomal protein S7 | 10.02 | 9.94 | -0.01 | 0.07 | 0.1 | 0.0 | Detail |
| 506 | THCA | cuproptosis | MRPS9 | mitochondrial ribosomal protein S9 | 9.07 | 9.03 | -0.01 | 0.3 | 0.34 | 0.0 | Detail |
| 507 | THCA | cuproptosis | MT1 | metallothionein 1A | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 508 | THCA | cuproptosis | MT2 | metallothionein 1B | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 509 | THCA | cuproptosis | MT3A | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 510 | THCA | cuproptosis | MTBP | MDM2 binding protein | 5.06 | 4.85 | -0.06 | 0.1 | 0.13 | 0.0 | Detail |
| 511 | THCA | cuproptosis | MT-CO1 | mitochondrially encoded cytochrome c oxidase I | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 512 | THCA | cuproptosis | MT-CO2 | mitochondrially encoded cytochrome c oxidase II | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 513 | THCA | cuproptosis | MTCO2P12 | MT-CO2 pseudogene 12 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 514 | THCA | cuproptosis | MT-CO3 | mitochondrially encoded cytochrome c oxidase III | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 515 | THCA | cuproptosis | MT | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 516 | THCA | cuproptosis | MTERF1 | mitochondrial transcription termination factor 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 517 | THCA | cuproptosis | MTFR1L | mitochondrial fission regulator 1 like | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 518 | THCA | cuproptosis | MTG1 | mitochondrial ribosome associated GTPase 1 | 8.99 | 9.03 | 0.01 | 0.2 | 0.24 | 0.0 | Detail |
| 519 | THCA | cuproptosis | MTO1 | mitochondrial tRNA translation optimization 1 | 8.94 | 8.84 | -0.02 | 0.05 | 0.07 | 0.0 | Detail |
| 520 | THCA | cuproptosis | mTOR | mechanistic target of rapamycin kinase | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 521 | THCA | cuproptosis | NAPA | NSF attachment protein alpha | 11.56 | 11.52 | -0.0 | 0.48 | 0.52 | 0.0 | Detail |
| 522 | THCA | cuproptosis | NARS2 | asparaginyl-tRNA synthetase 2, mitochondrial | 8.67 | 8.71 | 0.01 | 0.32 | 0.37 | 0.0 | Detail |
| 523 | THCA | cuproptosis | NDUFA6 | NADH:ubiquinone oxidoreductase subunit A6 | 10.09 | 9.97 | -0.02 | 0.1 | 0.13 | 0.0 | Detail |
| 524 | THCA | cuproptosis | NDUFA8 | NADH:ubiquinone oxidoreductase subunit A8 | 9.64 | 9.64 | 0.0 | 0.91 | 0.93 | 0.0 | Detail |
| 525 | THCA | cuproptosis | NDUFA9 | NADH:ubiquinone oxidoreductase subunit A9 | 9.98 | 10.09 | 0.01 | 0.09 | 0.12 | 0.0 | Detail |
| 526 | THCA | cuproptosis | NDUFAF6 | NADH:ubiquinone oxidoreductase complex assembly factor 6 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 527 | THCA | cuproptosis | NDUFB11 | NADH:ubiquinone oxidoreductase subunit B11 | 10.98 | 11.03 | 0.01 | 0.34 | 0.39 | 0.0 | Detail |
| 528 | THCA | cuproptosis | NDUFB2 | NADH:ubiquinone oxidoreductase subunit B2 | 11.67 | 11.52 | -0.02 | 0.16 | 0.2 | 0.0 | Detail |
| 529 | THCA | cuproptosis | NDUFB4 | NADH:ubiquinone oxidoreductase subunit B4 | 10.55 | 10.66 | 0.01 | 0.05 | 0.07 | 0.0 | Detail |
| 530 | THCA | cuproptosis | NDUFB5 | NADH:ubiquinone oxidoreductase subunit B5 | 10.32 | 10.23 | -0.01 | 0.05 | 0.07 | 0.0 | Detail |
| 531 | THCA | cuproptosis | NDUFB7 | NADH:ubiquinone oxidoreductase subunit B7 | 10.64 | 10.62 | -0.0 | 0.71 | 0.73 | 0.0 | Detail |
| 532 | THCA | cuproptosis | NDUFC1 | NADH:ubiquinone oxidoreductase subunit C1 | 10.09 | 10.11 | 0.0 | 0.83 | 0.85 | 0.0 | Detail |
| 533 | THCA | cuproptosis | NDUFC2 | NADH:ubiquinone oxidoreductase subunit C2 | 11.83 | 11.94 | 0.01 | 0.13 | 0.16 | 0.0 | Detail |
| 534 | THCA | cuproptosis | NDUFS2 | NADH:ubiquinone oxidoreductase core subunit S2 | 10.62 | 10.59 | -0.0 | 0.05 | 0.07 | 0.0 | Detail |
| 535 | THCA | cuproptosis | NDUFS3 | NADH:ubiquinone oxidoreductase core subunit S3 | 10.3 | 10.41 | 0.02 | 0.05 | 0.07 | 0.0 | Detail |
| 536 | THCA | cuproptosis | NDUFS6 | NADH:ubiquinone oxidoreductase subunit S6 | 9.76 | 9.85 | 0.01 | 0.13 | 0.17 | 0.0 | Detail |
| 537 | THCA | cuproptosis | NDUFV2 | NADH:ubiquinone oxidoreductase core subunit V2 | 10.28 | 10.26 | -0.0 | 0.76 | 0.78 | 0.0 | Detail |
| 538 | THCA | cuproptosis | NFU1 | NFU1 iron-sulfur cluster scaffold | 9.33 | 9.34 | 0.0 | 0.6 | 0.64 | 0.0 | Detail |
| 539 | THCA | cuproptosis | NLRP3 | NLR family pyrin domain containing 3 | 5.96 | 5.77 | -0.05 | 0.78 | 0.81 | 0.0 | Detail |
| 540 | THCA | cuproptosis | NPL4 | NPL4 homolog, ubiquitin recognition factor | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 541 | THCA | cuproptosis | NSF | N-ethylmaleimide sensitive factor, vesicle fusing ATPase | 9.96 | 9.92 | -0.0 | 0.49 | 0.53 | 0.0 | Detail |
| 542 | THCA | cuproptosis | NTHL1 | nth like DNA glycosylase 1 | 7.95 | 7.86 | -0.02 | 0.27 | 0.31 | 0.0 | Detail |
| 543 | THCA | cuproptosis | NUBP1 | NUBP iron-sulfur cluster assembly factor 1, cytosolic | 9.28 | 9.28 | -0.0 | 0.82 | 0.85 | 0.0 | Detail |
| 544 | THCA | cuproptosis | P3H3 | prolyl 3-hydroxylase 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 545 | THCA | cuproptosis | p53 | tumor protein p53 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 546 | THCA | cuproptosis | P53 | tumor protein p53 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 547 | THCA | cuproptosis | p97 | eukaryotic translation initiation factor 4 gamma 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 548 | THCA | cuproptosis | PARS2 | prolyl-tRNA synthetase 2, mitochondrial | 6.71 | 6.95 | 0.05 | 0.06 | 0.09 | 0.0 | Detail |
| 549 | THCA | cuproptosis | PCGF6 | polycomb group ring finger 6 | 6.8 | 6.76 | -0.01 | 0.63 | 0.67 | 0.0 | Detail |
| 550 | THCA | cuproptosis | PDHA | pyruvate dehydrogenase E1 subunit alpha 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 551 | THCA | cuproptosis | PDH | pyruvate dehydrogenase phosphatase catalytic subunit 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 552 | THCA | cuproptosis | PDHX | pyruvate dehydrogenase complex component X | 8.78 | 8.72 | -0.01 | 0.06 | 0.08 | 0.0 | Detail |
| 553 | THCA | cuproptosis | PET100 | PET100 cytochrome c oxidase chaperone | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 554 | THCA | cuproptosis | PET117 | PET117 cytochrome c oxidase chaperone | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 555 | THCA | cuproptosis | PGP | phosphoglycolate phosphatase | 8.79 | 9.0 | 0.03 | 0.05 | 0.06 | 0.0 | Detail |
| 556 | THCA | cuproptosis | PHOB | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 557 | THCA | cuproptosis | PI3K | phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 558 | THCA | cuproptosis | PMVK | phosphomevalonate kinase | 10.14 | 10.13 | -0.0 | 0.98 | 0.98 | 0.0 | Detail |
| 559 | THCA | cuproptosis | POLG | DNA polymerase gamma, catalytic subunit | 9.57 | 9.53 | -0.01 | 0.52 | 0.56 | 0.0 | Detail |
| 560 | THCA | cuproptosis | PPAT | phosphoribosyl pyrophosphate amidotransferase | 7.65 | 7.53 | -0.02 | 0.07 | 0.1 | 0.0 | Detail |
| 561 | THCA | cuproptosis | PSMD6 | proteasome 26S subunit, non-ATPase 6 | 10.01 | 10.02 | 0.0 | 0.59 | 0.63 | 0.0 | Detail |
| 562 | THCA | cuproptosis | RANTES | C-C motif chemokine ligand 5 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 563 | THCA | cuproptosis | RB1 | RB transcriptional corepressor 1 | 10.07 | 10.12 | 0.01 | 0.1 | 0.13 | 0.0 | Detail |
| 564 | THCA | cuproptosis | RCC1L | RCC1 like | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 565 | THCA | cuproptosis | RPAIN | RPA interacting protein | 9.08 | 9.12 | 0.01 | 0.3 | 0.34 | 0.0 | Detail |
| 566 | THCA | cuproptosis | RPS25 | ribosomal protein S25 | 13.2 | 13.18 | -0.0 | 0.66 | 0.69 | 0.0 | Detail |
| 567 | THCA | cuproptosis | RPUSD4 | RNA pseudouridine synthase D4 | 8.62 | 8.61 | -0.0 | 0.89 | 0.91 | 0.0 | Detail |
| 568 | THCA | cuproptosis | RRN3 | RRN3 homolog, RNA polymerase I transcription factor | 10.02 | 9.98 | -0.0 | 0.34 | 0.38 | 0.0 | Detail |
| 569 | THCA | cuproptosis | RSL1D1 | ribosomal L1 domain containing 1 | 11.43 | 11.52 | 0.01 | 0.05 | 0.06 | 0.0 | Detail |
| 570 | THCA | cuproptosis | S6K1 | ribosomal protein S6 kinase B1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 571 | THCA | cuproptosis | SDHAF2 | succinate dehydrogenase complex assembly factor 2 | 9.49 | 9.4 | -0.01 | 0.08 | 0.1 | 0.0 | Detail |
| 572 | THCA | cuproptosis | SDHB | succinate dehydrogenase complex iron sulfur subunit B | 10.21 | 10.17 | -0.01 | 0.35 | 0.39 | 0.0 | Detail |
| 573 | THCA | cuproptosis | SEPTIN1 | septin 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 574 | THCA | cuproptosis | SGF29 | SAGA complex associated factor 29 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 575 | THCA | cuproptosis | SLC25A51 | solute carrier family 25 member 51 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 576 | THCA | cuproptosis | SLC30A7 | solute carrier family 30 member 7 | 9.62 | 9.69 | 0.01 | 0.06 | 0.09 | 0.0 | Detail |
| 577 | THCA | cuproptosis | SLC52A2 | solute carrier family 52 member 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 578 | THCA | cuproptosis | SLC7A11 | solute carrier family 7 member 11 | 7.37 | 7.22 | -0.03 | 0.24 | 0.28 | 0.0 | Detail |
| 579 | THCA | cuproptosis | SMAD3-AS1 | SMAD3 antisense RNA 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 580 | THCA | cuproptosis | SOD1 | superoxide dismutase 1 | 11.95 | 11.9 | -0.01 | 0.4 | 0.44 | 0.0 | Detail |
| 581 | THCA | cuproptosis | SOD2 | superoxide dismutase 2 | 11.51 | 11.35 | -0.02 | 0.04 | 0.06 | 0.0 | Detail |
| 582 | THCA | cuproptosis | sp1 | Sp1 transcription factor | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 583 | THCA | cuproptosis | SP1 | Sp1 transcription factor | 11.02 | 10.98 | -0.01 | 0.29 | 0.33 | 0.0 | Detail |
| 584 | THCA | cuproptosis | STEAP4 | unknown | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 585 | THCA | cuproptosis | STEAP | STEAP family member 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 586 | THCA | cuproptosis | SURF1 | SURF1 cytochrome c oxidase assembly factor | 10.01 | 9.91 | -0.02 | 0.09 | 0.12 | 0.0 | Detail |
| 587 | THCA | cuproptosis | SURF6 | surfeit 6 | 9.72 | 9.77 | 0.01 | 0.29 | 0.33 | 0.0 | Detail |
| 588 | THCA | cuproptosis | TAF4 | TATA-box binding protein associated factor 4 | 8.19 | 8.14 | -0.01 | 0.33 | 0.37 | 0.0 | Detail |
| 589 | THCA | cuproptosis | TAF6L | TATA-box binding protein associated factor 6 like | 8.63 | 8.53 | -0.02 | 0.03 | 0.05 | 0.0 | Detail |
| 590 | THCA | cuproptosis | TARS2 | threonyl-tRNA synthetase 2, mitochondrial | 9.0 | 9.0 | 0.0 | 0.45 | 0.49 | 0.0 | Detail |
| 591 | THCA | cuproptosis | TEDC1 | tubulin epsilon and delta complex 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 592 | THCA | cuproptosis | TEFM | transcription elongation factor, mitochondrial | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 593 | THCA | cuproptosis | TIMM9 | translocase of inner mitochondrial membrane 9 | 8.82 | 8.87 | 0.01 | 0.29 | 0.33 | 0.0 | Detail |
| 594 | THCA | cuproptosis | TIMMDC1 | translocase of inner mitochondrial membrane domain containing 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 595 | THCA | cuproptosis | TMEM191B | transmembrane protein 191B | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 596 | THCA | cuproptosis | TMEM199 | transmembrane protein 199 | 8.78 | 8.78 | -0.0 | 0.58 | 0.62 | 0.0 | Detail |
| 597 | THCA | cuproptosis | TOP1MT | DNA topoisomerase I mitochondrial | 9.49 | 9.42 | -0.01 | 0.24 | 0.28 | 0.0 | Detail |
| 598 | THCA | cuproptosis | TPI1 | triosephosphate isomerase 1 | 12.33 | 12.4 | 0.01 | 0.11 | 0.14 | 0.0 | Detail |
| 599 | THCA | cuproptosis | TRAIP | TRAF interacting protein | 5.68 | 5.58 | -0.03 | 0.41 | 0.45 | 0.0 | Detail |
| 600 | THCA | cuproptosis | TRAPPC11 | trafficking protein particle complex subunit 11 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 601 | THCA | cuproptosis | TRIM32 | tripartite motif containing 32 | 8.33 | 8.37 | 0.01 | 0.26 | 0.31 | 0.0 | Detail |
| 602 | THCA | cuproptosis | TRRAP | transformation/transcription domain associated protein | 11.03 | 10.97 | -0.01 | 0.12 | 0.15 | 0.0 | Detail |
| 603 | THCA | cuproptosis | TWNK | twinkle mtDNA helicase | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 604 | THCA | cuproptosis | TYR | tyrosinase | 0.1 | 0.16 | 0.65 | 0.13 | 0.17 | 0.0 | Detail |
| 605 | THCA | cuproptosis | U2SURP | U2 snRNP associated SURP domain containing | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 606 | THCA | cuproptosis | UBE2D1 | ubiquitin conjugating enzyme E2 D1 | 8.54 | 8.5 | -0.01 | 0.75 | 0.77 | 0.0 | Detail |
| 607 | THCA | cuproptosis | UlK1 | unc-51 like autophagy activating kinase 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 608 | THCA | cuproptosis | ULK1 | unc-51 like autophagy activating kinase 1 | 10.1 | 10.12 | 0.0 | 0.67 | 0.7 | 0.0 | Detail |
| 609 | THCA | cuproptosis | UQCC1 | ubiquinol-cytochrome c reductase complex assembly factor 1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 610 | THCA | cuproptosis | UQCC2 | ubiquinol-cytochrome c reductase complex assembly factor 2 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 611 | THCA | cuproptosis | UQCR10 | ubiquinol-cytochrome c reductase, complex III subunit X | 10.52 | 10.42 | -0.01 | 0.15 | 0.18 | 0.0 | Detail |
| 612 | THCA | cuproptosis | UTP25 | UTP25 small subunit processome component | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 613 | THCA | cuproptosis | VARS2 | valyl-tRNA synthetase 2, mitochondrial | 9.4 | 9.5 | 0.02 | 0.2 | 0.23 | 0.0 | Detail |
| 614 | THCA | cuproptosis | VPS45 | vacuolar protein sorting 45 homolog | 9.8 | 9.75 | -0.01 | 0.07 | 0.09 | 0.0 | Detail |
| 615 | THCA | cuproptosis | WARS2 | tryptophanyl tRNA synthetase 2, mitochondrial | 8.39 | 8.29 | -0.02 | 0.08 | 0.11 | 0.0 | Detail |
| 616 | THCA | cuproptosis | WDR12 | WD repeat domain 12 | 8.07 | 8.02 | -0.01 | 0.39 | 0.43 | 0.0 | Detail |
| 617 | THCA | cuproptosis | xCT | solute carrier family 7 member 11 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 618 | THCA | cuproptosis | YAE1 | YAE1 maturation factor of ABCE1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | Detail |
| 619 | THCA | cuproptosis | ZBP1 | Z-DNA binding protein 1 | 4.23 | 3.47 | -0.29 | 0.05 | 0.07 | 0.0 | Detail |
| 620 | THCA | cuproptosis | ZNF365 | zinc finger protein 365 | 4.56 | 4.73 | 0.05 | 0.05 | 0.07 | 0.0 | Detail |
Ensemble learning visualization:
Ensemble learning for THCA based on specific cuproptosis gene regulaors.